Dino Sejdinovic
A Perspective on Gaussian Processes for Earth Observation
Jul 02, 2020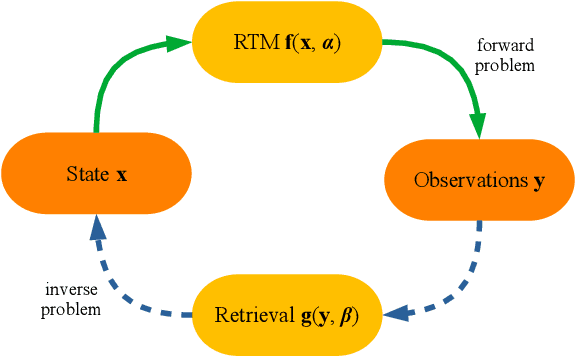
Abstract:Earth observation (EO) by airborne and satellite remote sensing and in-situ observations play a fundamental role in monitoring our planet. In the last decade, machine learning and Gaussian processes (GPs) in particular has attained outstanding results in the estimation of bio-geo-physical variables from the acquired images at local and global scales in a time-resolved manner. GPs provide not only accurate estimates but also principled uncertainty estimates for the predictions, can easily accommodate multimodal data coming from different sensors and from multitemporal acquisitions, allow the introduction of physical knowledge, and a formal treatment of uncertainty quantification and error propagation. Despite great advances in forward and inverse modelling, GP models still have to face important challenges that are revised in this perspective paper. GP models should evolve towards data-driven physics-aware models that respect signal characteristics, be consistent with elementary laws of physics, and move from pure regression to observational causal inference.
* 1 figure
Learning Inconsistent Preferences with Kernel Methods
Jun 06, 2020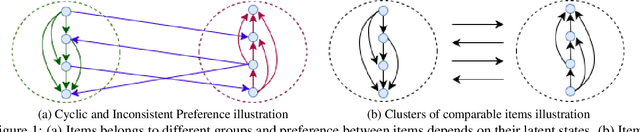
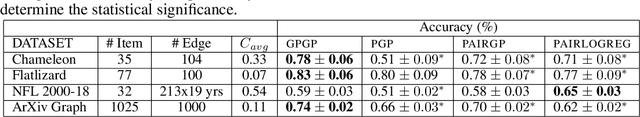
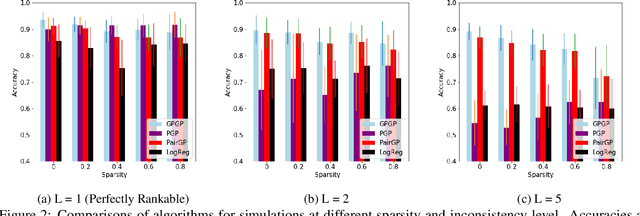
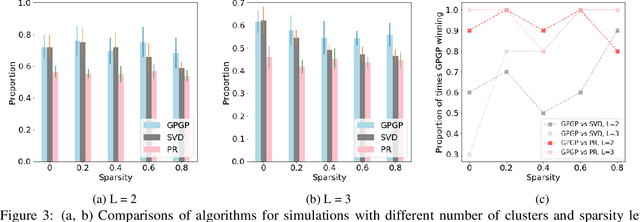
Abstract:We propose a probabilistic kernel approach for preferential learning from pairwise duelling data using Gaussian Processes. Different from previous methods, we do not impose a total order on the item space, hence can capture more expressive latent preferential structures such as inconsistent preferences and clusters of comparable items. Furthermore, we prove the universality of the proposed kernels, i.e. that the corresponding reproducing kernel Hilbert Space (RKHS) is dense in the space of skew-symmetric preference functions. To conclude the paper, we provide an extensive set of numerical experiments on simulated and real-world datasets showcasing the competitiveness of our proposed method with state-of-the-art.
Spectral Ranking with Covariates
May 13, 2020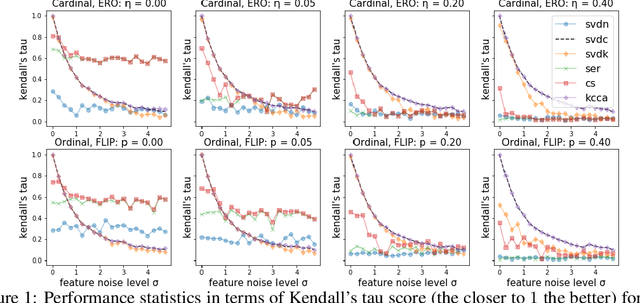
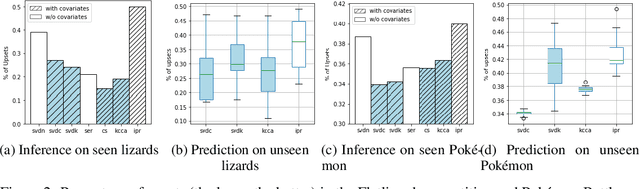
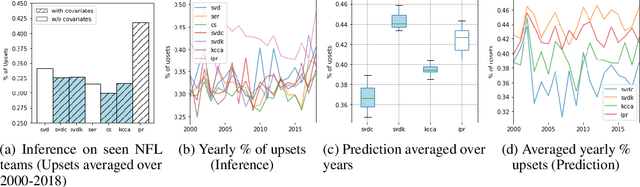
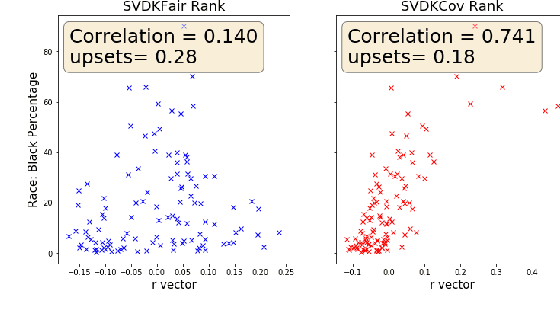
Abstract:We consider approaches to the classical problem of establishing a statistical ranking on a given set of items from incomplete and noisy pairwise comparisons, and propose spectral algorithms able to leverage available covariate information about the items. We give a comprehensive study of several ways such side information can be useful in spectral ranking. We establish connections of the resulting algorithms to reproducing kernel Hilbert spaces and associated dependence measures, along with an extension to fair ranking using statistical parity. We present an extensive set of numerical experiments showcasing the competitiveness of the proposed algorithms with state-of-the-art methods.
Large Scale Tensor Regression using Kernels and Variational Inference
Feb 11, 2020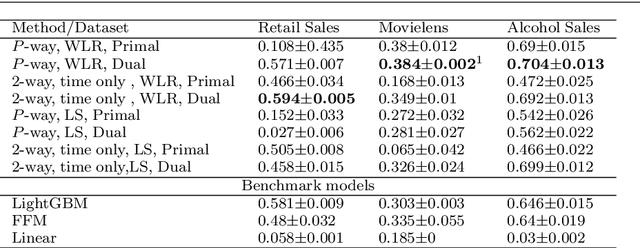
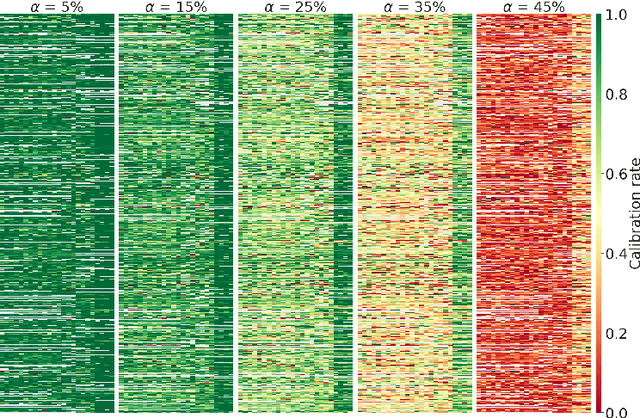
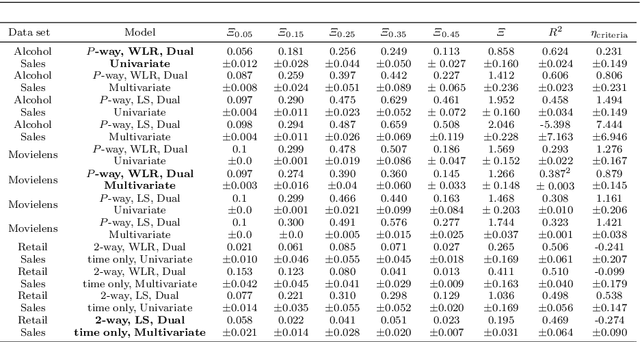
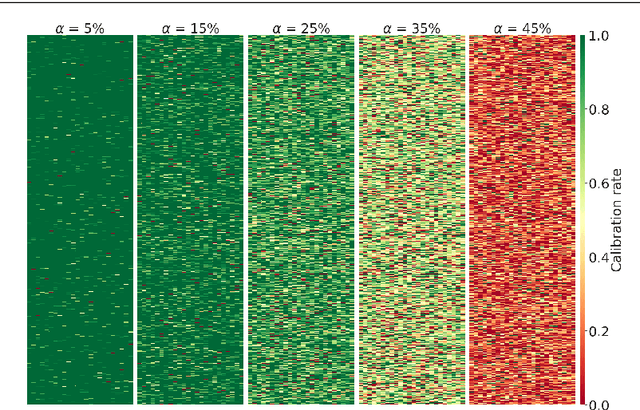
Abstract:We outline an inherent weakness of tensor factorization models when latent factors are expressed as a function of side information and propose a novel method to mitigate this weakness. We coin our method \textit{Kernel Fried Tensor}(KFT) and present it as a large scale forecasting tool for high dimensional data. Our results show superior performance against \textit{LightGBM} and \textit{Field Aware Factorization Machines}(FFM), two algorithms with proven track records widely used in industrial forecasting. We also develop a variational inference framework for KFT and associate our forecasts with calibrated uncertainty estimates on three large scale datasets. Furthermore, KFT is empirically shown to be robust against uninformative side information in terms of constants and Gaussian noise.
A kernel log-rank test of independence for right-censored data
Dec 08, 2019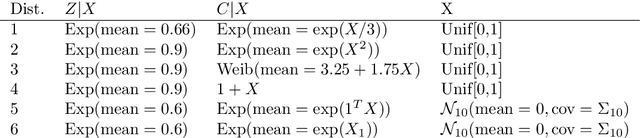
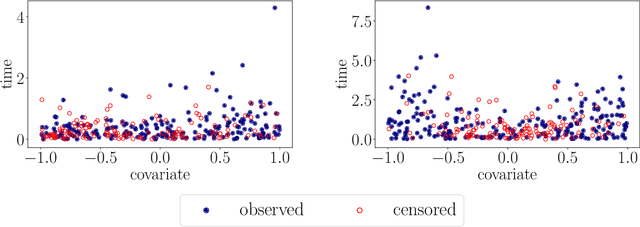
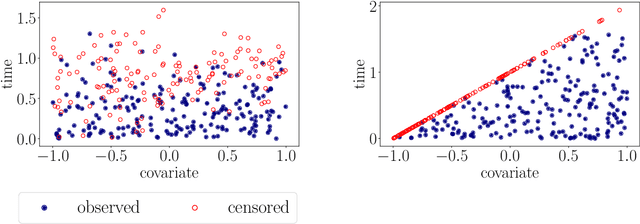
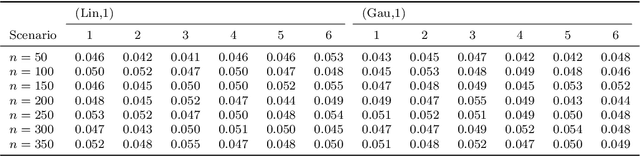
Abstract:With the incorporation of new data gathering methods in clinical research, it becomes fundamental for survival analysis techniques to deal with high-dimensional or/and non-standard covariates. In this paper we introduce a general non-parametric independence test between right-censored survival times and covariates taking values on a general (not necessarily Euclidean) space $\mathcal{X}$. We show that our test statistic has a dual interpretation, first in terms of the supremum of a potentially infinite collection of weight-indexed log-rank tests, with weight functions belonging to a reproducing kernel Hilbert space (RKHS) of functions; and second, as the norm of the difference of embeddings of certain finite measures into the RKHS, similar to the Hilbert-Schmidt Independence Criterion (HSIC) test-statistic. We study the asymptotic properties of the test, finding sufficient conditions to ensure that our test is omnibus. The test statistic can be computed straightforwardly, and the rejection threshold is obtained via an asymptotically consistent Wild-Bootstrap procedure. We perform extensive simulations demonstrating that our testing procedure generally performs better than competing approaches in detecting complex nonlinear dependence.
Detecting anthropogenic cloud perturbations with deep learning
Nov 29, 2019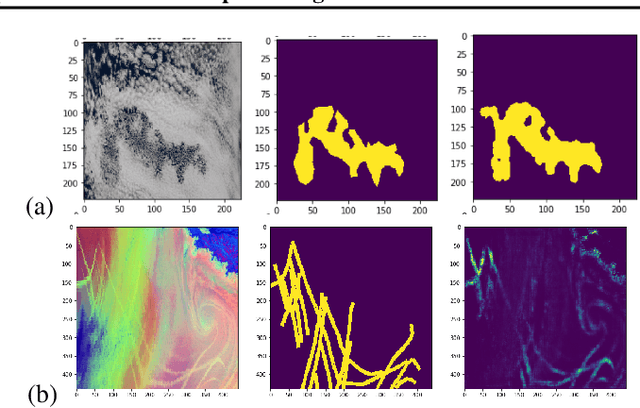
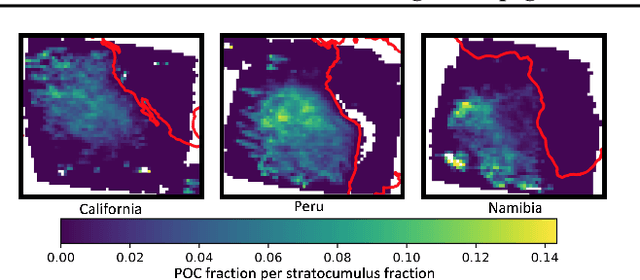
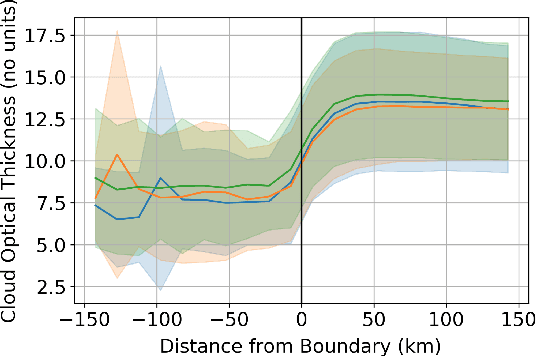
Abstract:One of the most pressing questions in climate science is that of the effect of anthropogenic aerosol on the Earth's energy balance. Aerosols provide the `seeds' on which cloud droplets form, and changes in the amount of aerosol available to a cloud can change its brightness and other physical properties such as optical thickness and spatial extent. Clouds play a critical role in moderating global temperatures and small perturbations can lead to significant amounts of cooling or warming. Uncertainty in this effect is so large it is not currently known if it is negligible, or provides a large enough cooling to largely negate present-day warming by CO2. This work uses deep convolutional neural networks to look for two particular perturbations in clouds due to anthropogenic aerosol and assess their properties and prevalence, providing valuable insights into their climatic effects.
Kernel Dependence Regularizers and Gaussian Processes with Applications to Algorithmic Fairness
Nov 11, 2019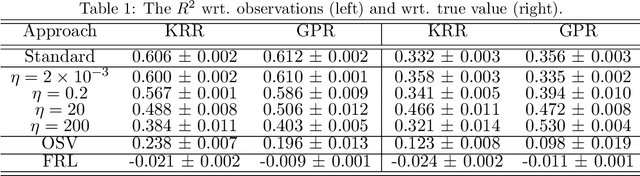
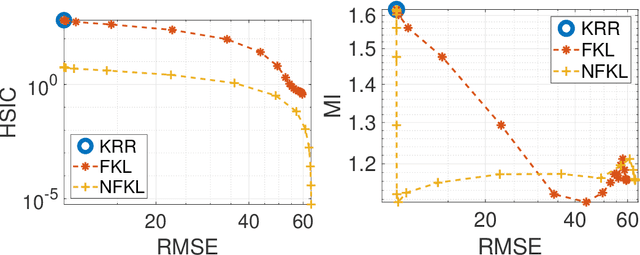
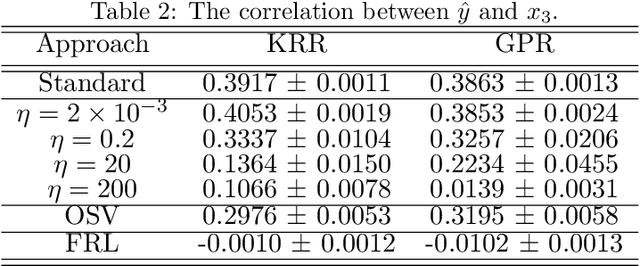
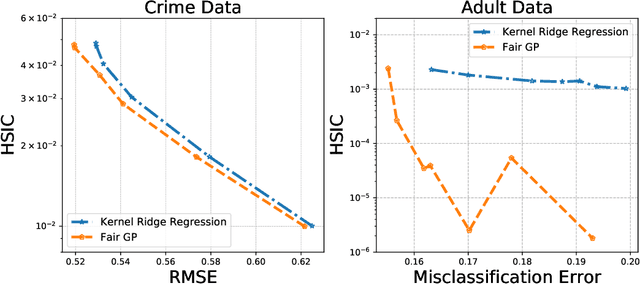
Abstract:Current adoption of machine learning in industrial, societal and economical activities has raised concerns about the fairness, equity and ethics of automated decisions. Predictive models are often developed using biased datasets and thus retain or even exacerbate biases in their decisions and recommendations. Removing the sensitive covariates, such as gender or race, is insufficient to remedy this issue since the biases may be retained due to other related covariates. We present a regularization approach to this problem that trades off predictive accuracy of the learned models (with respect to biased labels) for the fairness in terms of statistical parity, i.e. independence of the decisions from the sensitive covariates. In particular, we consider a general framework of regularized empirical risk minimization over reproducing kernel Hilbert spaces and impose an additional regularizer of dependence between predictors and sensitive covariates using kernel-based measures of dependence, namely the Hilbert-Schmidt Independence Criterion (HSIC) and its normalized version. This approach leads to a closed-form solution in the case of squared loss, i.e. ridge regression. Moreover, we show that the dependence regularizer has an interpretation as modifying the corresponding Gaussian process (GP) prior. As a consequence, a GP model with a prior that encourages fairness to sensitive variables can be derived, allowing principled hyperparameter selection and studying of the relative relevance of covariates under fairness constraints. Experimental results in synthetic examples and in real problems of income and crime prediction illustrate the potential of the approach to improve fairness of automated decisions.
Nonparametric Independence Testing for Right-Censored Data using Optimal Transport
Jun 10, 2019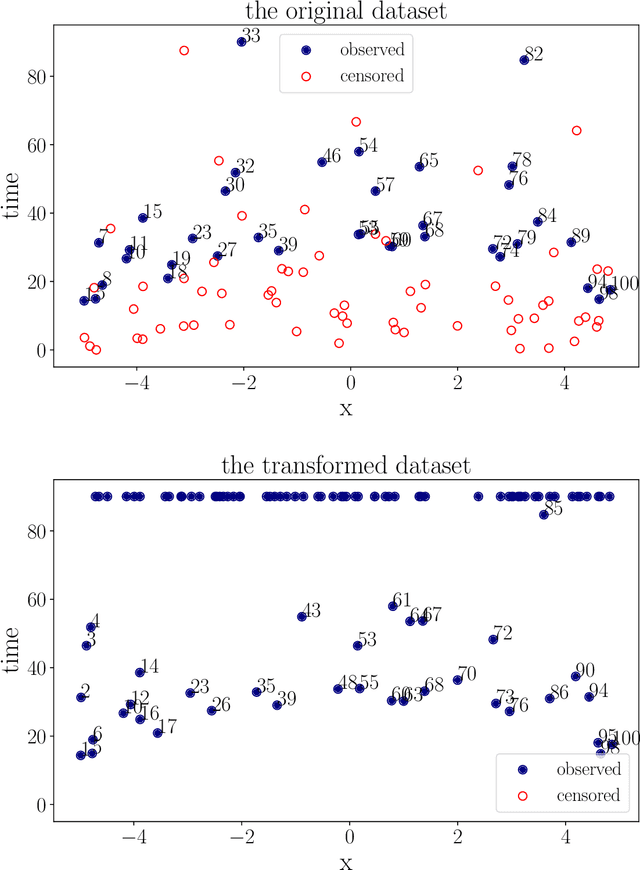
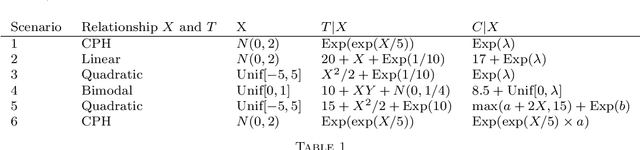
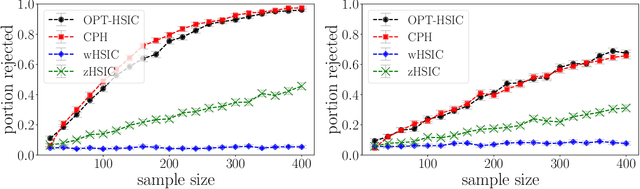
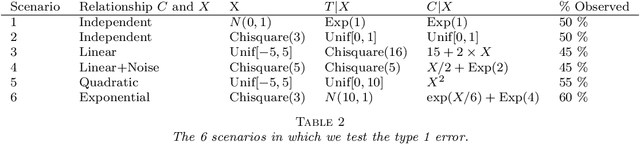
Abstract:We propose a nonparametric test of independence, termed OPT-HSIC, between a covariate and a right-censored lifetime. Because the presence of censoring creates a challenge in applying the standard permutation-based testing approaches, we use optimal transport to transform the censored dataset into an uncensored one, while preserving the relevant dependencies. We then apply a permutation test using the kernel-based dependence measure as a statistic to the transformed dataset. The type 1 error is proven to be correct in the case where censoring is independent of the covariate. Experiments indicate that OPT-HSIC has power against a much wider class of alternatives than Cox proportional hazards regression and that it has the correct type 1 control even in the challenging cases where censoring strongly depends on the covariate.
Noise Contrastive Meta-Learning for Conditional Density Estimation using Kernel Mean Embeddings
Jun 05, 2019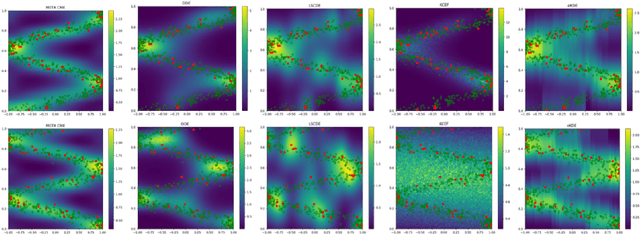
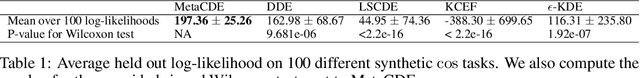
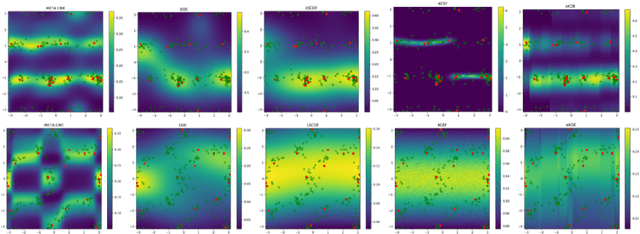

Abstract:Current meta-learning approaches focus on learning functional representations of relationships between variables, i.e. on estimating conditional expectations in regression. In many applications, however, we are faced with conditional distributions which cannot be meaningfully summarized using expectation only (due to e.g. multimodality). Hence, we consider the problem of conditional density estimation in the meta-learning setting. We introduce a novel technique for meta-learning which combines neural representation and noise-contrastive estimation with the established literature of conditional mean embeddings into reproducing kernel Hilbert spaces. The method is validated on synthetic and real-world problems, demonstrating the utility of sharing learned representations across multiple conditional density estimation tasks.
Rejoinder for "Probabilistic Integration: A Role in Statistical Computation?"
Nov 26, 2018Abstract:This article is the rejoinder for the paper "Probabilistic Integration: A Role in Statistical Computation?" to appear in Statistical Science with discussion. We would first like to thank the reviewers and many of our colleagues who helped shape this paper, the editor for selecting our paper for discussion, and of course all of the discussants for their thoughtful, insightful and constructive comments. In this rejoinder, we respond to some of the points raised by the discussants and comment further on the fundamental questions underlying the paper: (i) Should Bayesian ideas be used in numerical analysis?, and (ii) If so, what role should such approaches have in statistical computation?
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge