Defu Yang
Tau-BNO: Brain Neural Operator for Tau Transport Model
Mar 09, 2026Abstract:Mechanistic modeling provides a biophysically grounded framework for studying the spread of pathological tau protein in tauopathies like Alzheimer's disease. Existing approaches typically model tau propagation as a diffusive process on the brain's structural connectome, reproducing macroscopic patterns but neglecting microscale cellular transport and reaction mechanisms. The Network Transport Model (NTM) was introduced to fill this gap, explaining how region-level progression of tau emerges from microscale biophysical processes. However, the NTM faces a common challenge for complex models defined by large systems of partial differential equations: the inability to perform parameter inference and mechanistic discovery due to high computational burden and slow model simulations. To overcome this barrier, we propose Tau-BNO, a Brain Neural Operator surrogate framework for rapidly approximating NTM dynamics that captures both intra-regional reaction kinetics and inter-regional network transport. Tau-BNO combines a function operator that encodes kinetic parameters with a query operator that preserves initial state information, while approximating anisotropic transport through a spectral kernel that retains directionality. Empirical evaluations demonstrate high predictive accuracy ($R^2\approx$ 0.98) across diverse biophysical regimes and an 89\% performance improvement over state-of-the-art sequence models like Transformers and Mamba, which lack inherent structural priors. By reducing simulation time from hours to seconds, we show that the surrogate model is capable of producing new insights and generating new hypotheses. This framework is readily extensible to a broader class of connectome-based biophysical models, showcasing the transformative value of deep learning surrogates to accelerate analysis of large-scale, computationally intensive dynamical systems.
Interpretable deep learning illuminates multiple structures fluorescence imaging: a path toward trustworthy artificial intelligence in microscopy
Jan 09, 2025Abstract:Live-cell imaging of multiple subcellular structures is essential for understanding subcellular dynamics. However, the conventional multi-color sequential fluorescence microscopy suffers from significant imaging delays and limited number of subcellular structure separate labeling, resulting in substantial limitations for real-time live-cell research applications. Here, we present the Adaptive Explainable Multi-Structure Network (AEMS-Net), a deep-learning framework that enables simultaneous prediction of two subcellular structures from a single image. The model normalizes staining intensity and prioritizes critical image features by integrating attention mechanisms and brightness adaptation layers. Leveraging the Kolmogorov-Arnold representation theorem, our model decomposes learned features into interpretable univariate functions, enhancing the explainability of complex subcellular morphologies. We demonstrate that AEMS-Net allows real-time recording of interactions between mitochondria and microtubules, requiring only half the conventional sequential-channel imaging procedures. Notably, this approach achieves over 30% improvement in imaging quality compared to traditional deep learning methods, establishing a new paradigm for long-term, interpretable live-cell imaging that advances the ability to explore subcellular dynamics.
Pathology Steered Stratification Network for Subtype Identification in Alzheimer's Disease
Oct 12, 2022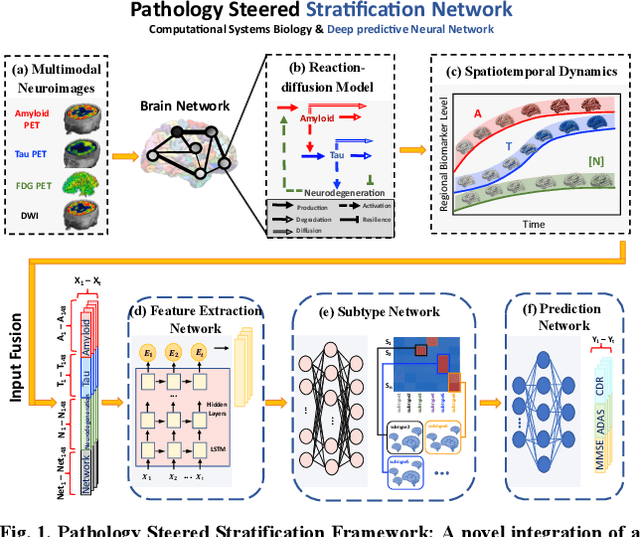
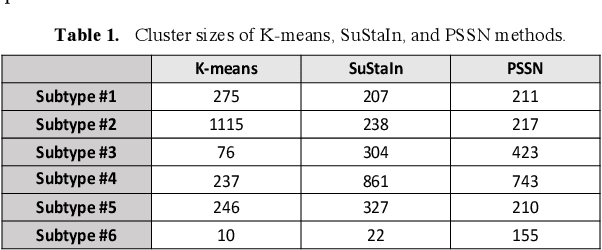
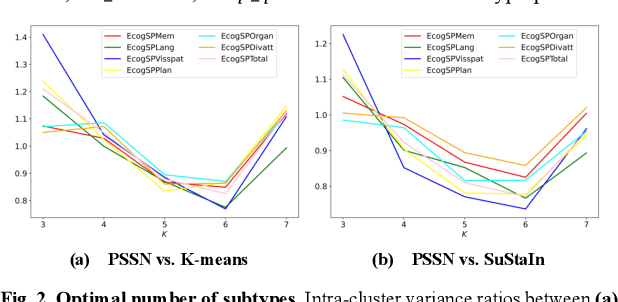
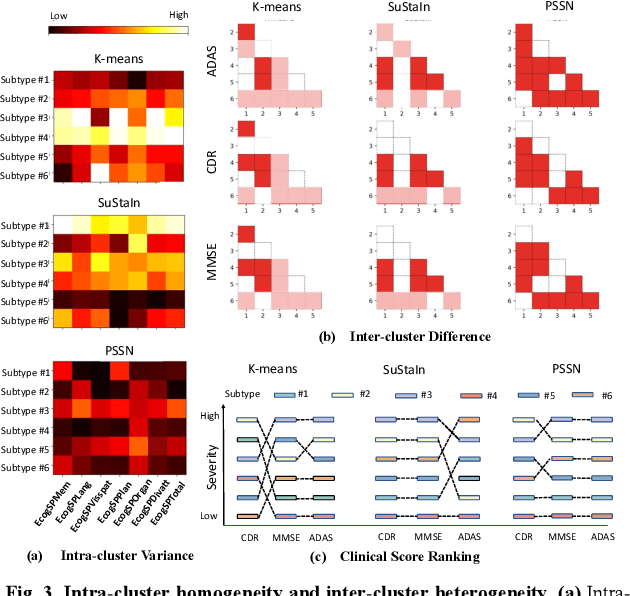
Abstract:Alzheimer's disease (AD) is a heterogeneous, multifactorial neurodegenerative disorder characterized by beta-amyloid, pathologic tau, and neurodegeneration. The massive heterogeneity between neurobiological examinations and clinical assessment is the current biggest challenge in the early diagnosis of Alzheimer's disease, urging for a comprehensive stratification of the aging population that is defined by reliable neurobiological biomarkers and closely associated with clinical outcomes. However, existing statistical inference approaches in neuroimaging studies of AD subtype identification fail to take into account the neuropathological domain knowledge, which could lead to ill-posed results that are sometimes inconsistent with neurological principles. To fill this knowledge gap, we propose a novel pathology steered stratification network (PSSN) that integrates mainstream AD pathology with multimodal longitudinal neuroimaging data to categorize the aging population. By combining theory-based biological modeling and data-driven deep learning, this cross-disciplinary approach can not only generate long-term biomarker prediction consistent with the end-state of individuals but also stratifies subjects into fine-grained subtypes with distinct neurological underpinnings, where ag-ing brains within the same subtype share com-mon biological behaviors that emerge as similar trajectories of cognitive decline. Our stratification outperforms K-means and SuStaIn in both inter-cluster heterogeneity and intra-cluster homogeneity of various clinical scores. Importantly, we identify six subtypes spanning AD spectrum, where each subtype exhibits a distinctive biomarker pattern that is consistent with its clinical outcome. A disease evolutionary graph is further provided by quantifying subtype transition probabilities, which may assist pre-symptomatic diagnosis and guide therapeutic treatments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge