Causal Effect Inference with Deep Latent-Variable Models
Nov 06, 2017Christos Louizos, Uri Shalit, Joris Mooij, David Sontag, Richard Zemel, Max Welling
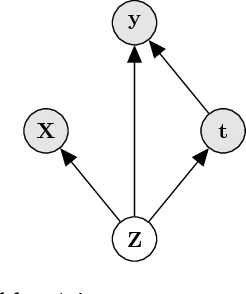
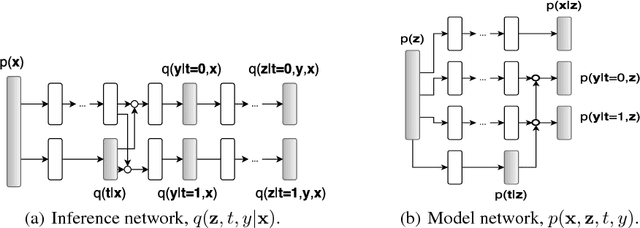
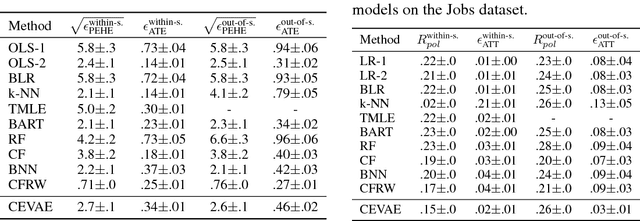
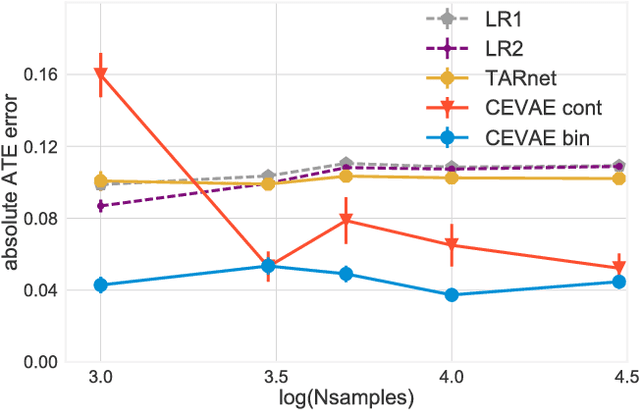
Learning individual-level causal effects from observational data, such as inferring the most effective medication for a specific patient, is a problem of growing importance for policy makers. The most important aspect of inferring causal effects from observational data is the handling of confounders, factors that affect both an intervention and its outcome. A carefully designed observational study attempts to measure all important confounders. However, even if one does not have direct access to all confounders, there may exist noisy and uncertain measurement of proxies for confounders. We build on recent advances in latent variable modeling to simultaneously estimate the unknown latent space summarizing the confounders and the causal effect. Our method is based on Variational Autoencoders (VAE) which follow the causal structure of inference with proxies. We show our method is significantly more robust than existing methods, and matches the state-of-the-art on previous benchmarks focused on individual treatment effects.
Grounded Recurrent Neural Networks
May 23, 2017Ankit Vani, Yacine Jernite, David Sontag
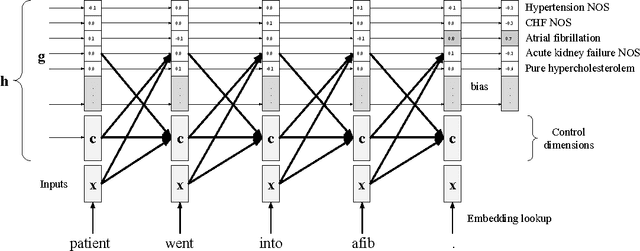
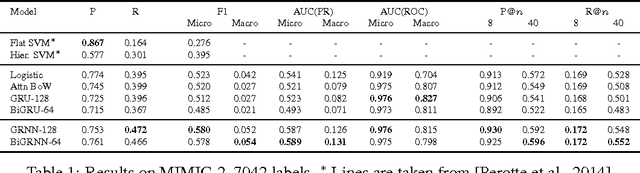
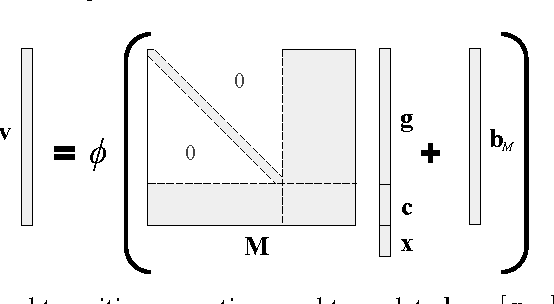
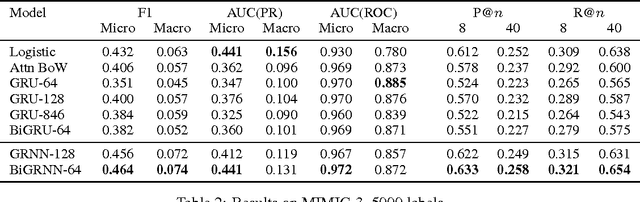
In this work, we present the Grounded Recurrent Neural Network (GRNN), a recurrent neural network architecture for multi-label prediction which explicitly ties labels to specific dimensions of the recurrent hidden state (we call this process "grounding"). The approach is particularly well-suited for extracting large numbers of concepts from text. We apply the new model to address an important problem in healthcare of understanding what medical concepts are discussed in clinical text. Using a publicly available dataset derived from Intensive Care Units, we learn to label a patient's diagnoses and procedures from their discharge summary. Our evaluation shows a clear advantage to using our proposed architecture over a variety of strong baselines.
Estimating individual treatment effect: generalization bounds and algorithms
May 16, 2017Uri Shalit, Fredrik D. Johansson, David Sontag
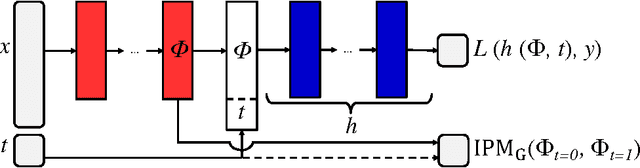
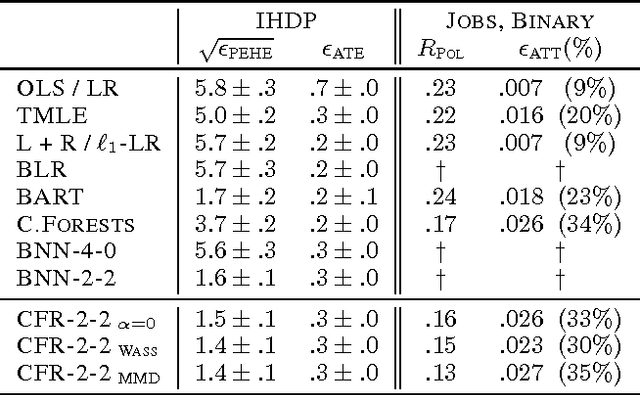
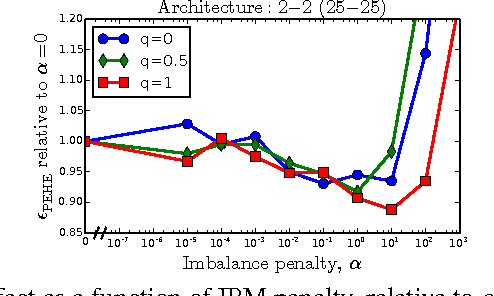
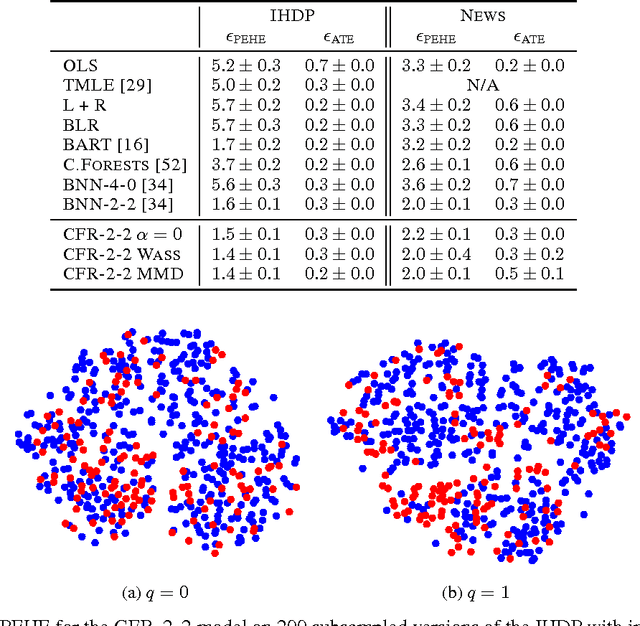
There is intense interest in applying machine learning to problems of causal inference in fields such as healthcare, economics and education. In particular, individual-level causal inference has important applications such as precision medicine. We give a new theoretical analysis and family of algorithms for predicting individual treatment effect (ITE) from observational data, under the assumption known as strong ignorability. The algorithms learn a "balanced" representation such that the induced treated and control distributions look similar. We give a novel, simple and intuitive generalization-error bound showing that the expected ITE estimation error of a representation is bounded by a sum of the standard generalization-error of that representation and the distance between the treated and control distributions induced by the representation. We use Integral Probability Metrics to measure distances between distributions, deriving explicit bounds for the Wasserstein and Maximum Mean Discrepancy (MMD) distances. Experiments on real and simulated data show the new algorithms match or outperform the state-of-the-art.
Discourse-Based Objectives for Fast Unsupervised Sentence Representation Learning
Apr 23, 2017Yacine Jernite, Samuel R. Bowman, David Sontag
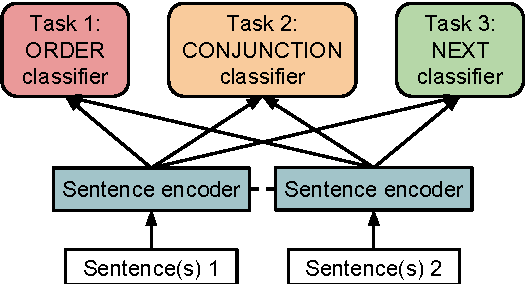
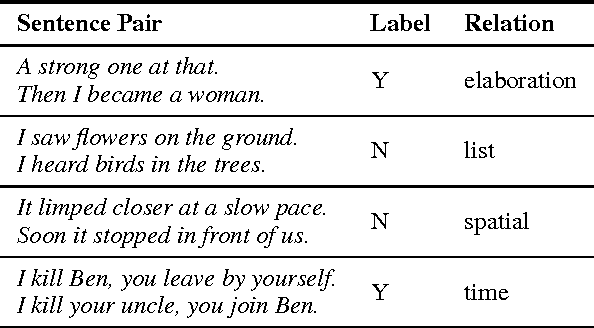

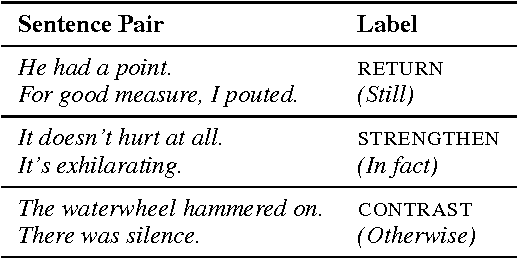
This work presents a novel objective function for the unsupervised training of neural network sentence encoders. It exploits signals from paragraph-level discourse coherence to train these models to understand text. Our objective is purely discriminative, allowing us to train models many times faster than was possible under prior methods, and it yields models which perform well in extrinsic evaluations.
Simultaneous Learning of Trees and Representations for Extreme Classification and Density Estimation
Mar 02, 2017Yacine Jernite, Anna Choromanska, David Sontag
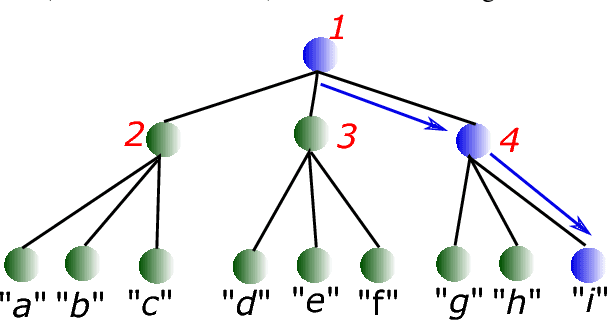
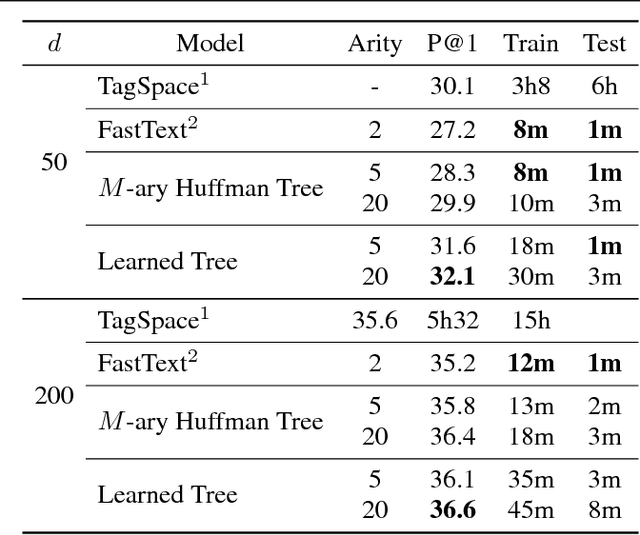
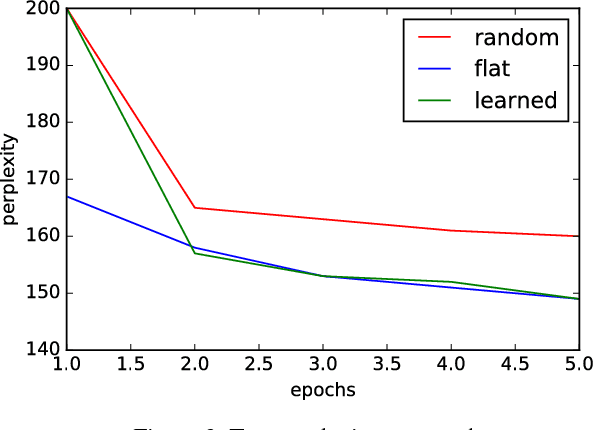
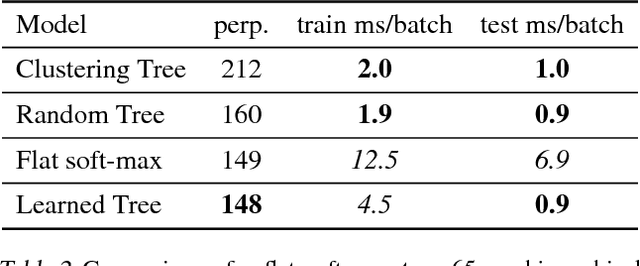
We consider multi-class classification where the predictor has a hierarchical structure that allows for a very large number of labels both at train and test time. The predictive power of such models can heavily depend on the structure of the tree, and although past work showed how to learn the tree structure, it expected that the feature vectors remained static. We provide a novel algorithm to simultaneously perform representation learning for the input data and learning of the hierarchi- cal predictor. Our approach optimizes an objec- tive function which favors balanced and easily- separable multi-way node partitions. We theoret- ically analyze this objective, showing that it gives rise to a boosting style property and a bound on classification error. We next show how to extend the algorithm to conditional density estimation. We empirically validate both variants of the al- gorithm on text classification and language mod- eling, respectively, and show that they compare favorably to common baselines in terms of accu- racy and running time.
Structured Inference Networks for Nonlinear State Space Models
Dec 05, 2016Rahul G. Krishnan, Uri Shalit, David Sontag
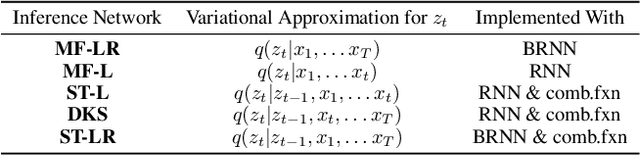
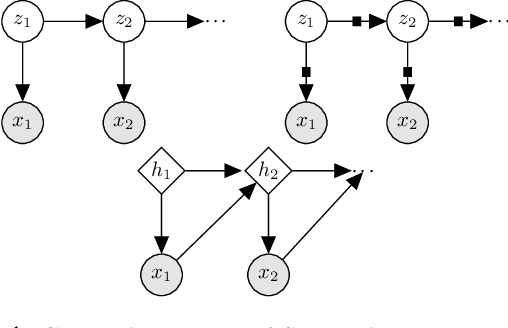

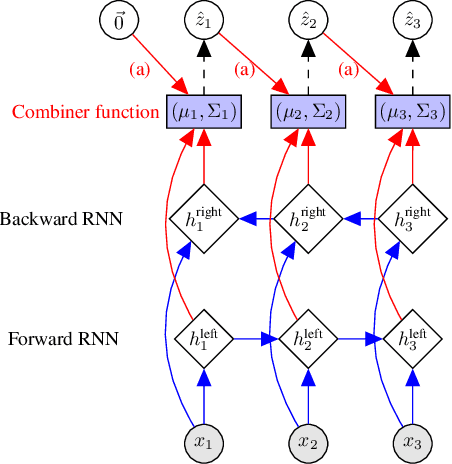
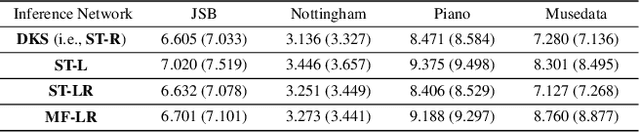
Gaussian state space models have been used for decades as generative models of sequential data. They admit an intuitive probabilistic interpretation, have a simple functional form, and enjoy widespread adoption. We introduce a unified algorithm to efficiently learn a broad class of linear and non-linear state space models, including variants where the emission and transition distributions are modeled by deep neural networks. Our learning algorithm simultaneously learns a compiled inference network and the generative model, leveraging a structured variational approximation parameterized by recurrent neural networks to mimic the posterior distribution. We apply the learning algorithm to both synthetic and real-world datasets, demonstrating its scalability and versatility. We find that using the structured approximation to the posterior results in models with significantly higher held-out likelihood.
Recurrent Neural Networks for Multivariate Time Series with Missing Values
Nov 07, 2016Zhengping Che, Sanjay Purushotham, Kyunghyun Cho, David Sontag, Yan Liu
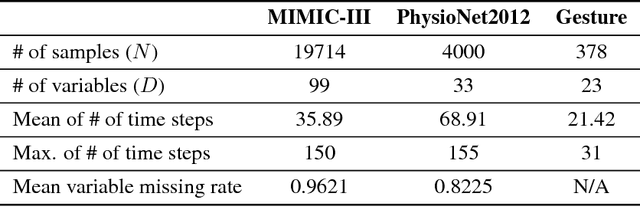
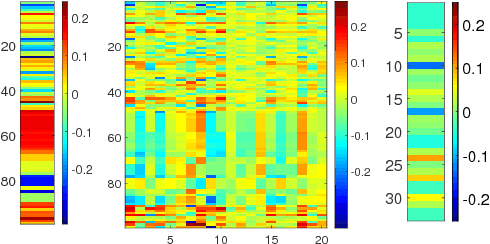
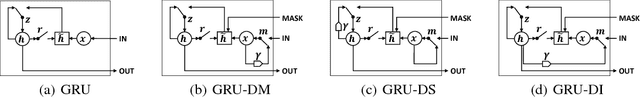
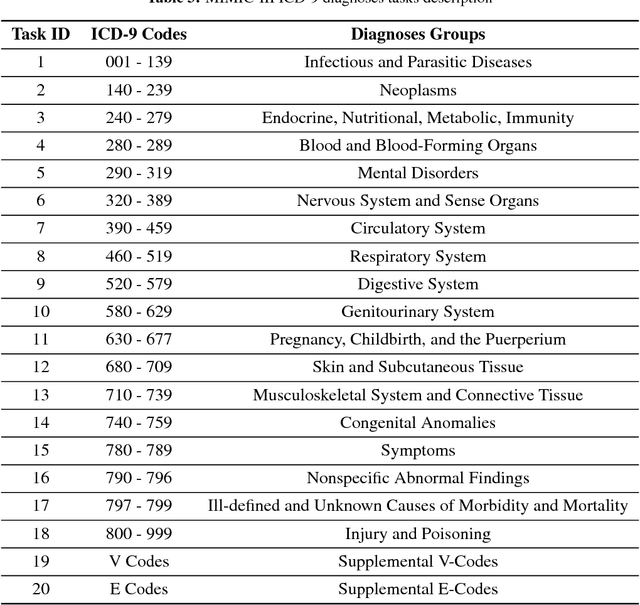
Multivariate time series data in practical applications, such as health care, geoscience, and biology, are characterized by a variety of missing values. In time series prediction and other related tasks, it has been noted that missing values and their missing patterns are often correlated with the target labels, a.k.a., informative missingness. There is very limited work on exploiting the missing patterns for effective imputation and improving prediction performance. In this paper, we develop novel deep learning models, namely GRU-D, as one of the early attempts. GRU-D is based on Gated Recurrent Unit (GRU), a state-of-the-art recurrent neural network. It takes two representations of missing patterns, i.e., masking and time interval, and effectively incorporates them into a deep model architecture so that it not only captures the long-term temporal dependencies in time series, but also utilizes the missing patterns to achieve better prediction results. Experiments of time series classification tasks on real-world clinical datasets (MIMIC-III, PhysioNet) and synthetic datasets demonstrate that our models achieve state-of-the-art performance and provides useful insights for better understanding and utilization of missing values in time series analysis.
Clinical Tagging with Joint Probabilistic Models
Sep 22, 2016Yoni Halpern, Steven Horng, David Sontag
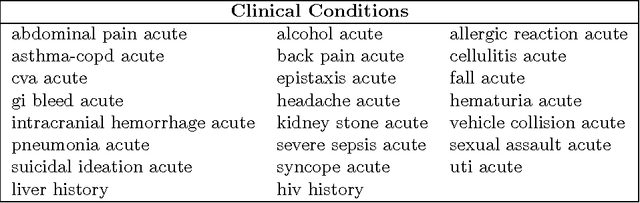
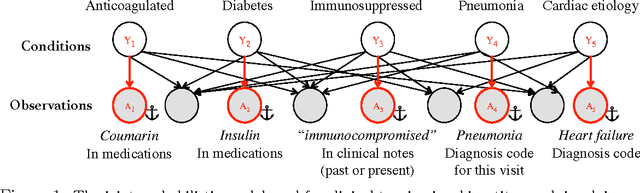
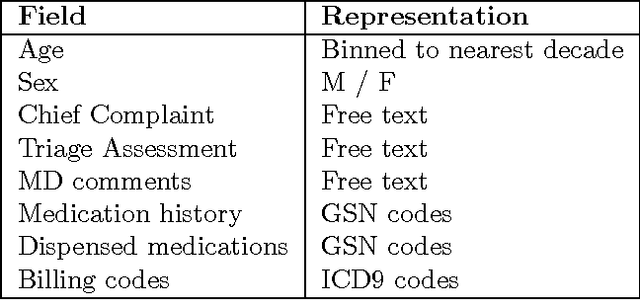

We describe a method for parameter estimation in bipartite probabilistic graphical models for joint prediction of clinical conditions from the electronic medical record. The method does not rely on the availability of gold-standard labels, but rather uses noisy labels, called anchors, for learning. We provide a likelihood-based objective and a moments-based initialization that are effective at learning the model parameters. The learned model is evaluated in a task of assigning a heldout clinical condition to patients based on retrospective analysis of the records, and outperforms baselines which do not account for the noisiness in the labels or do not model the conditions jointly.
Multi-task Prediction of Disease Onsets from Longitudinal Lab Tests
Sep 20, 2016Narges Razavian, Jake Marcus, David Sontag
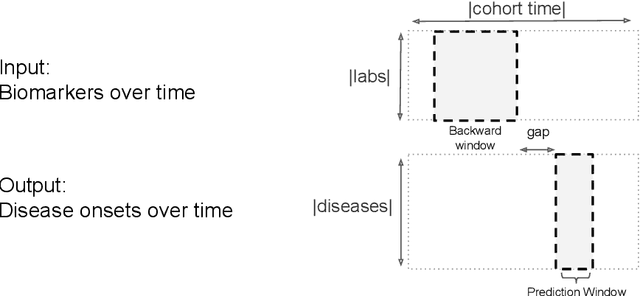
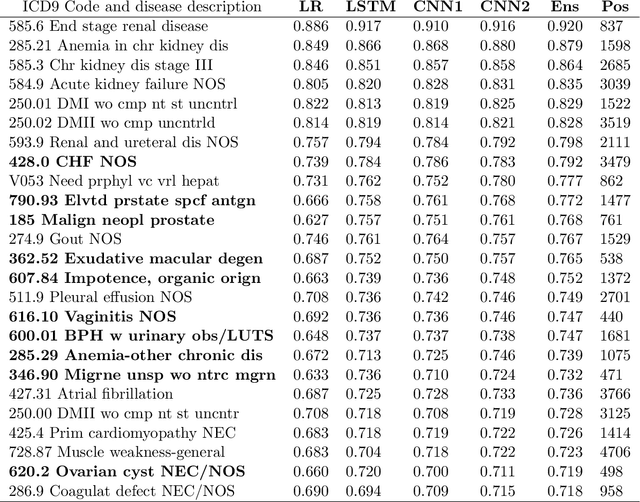
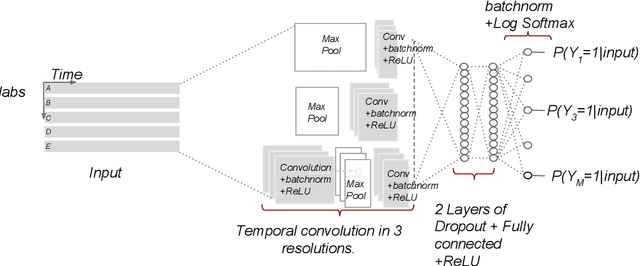
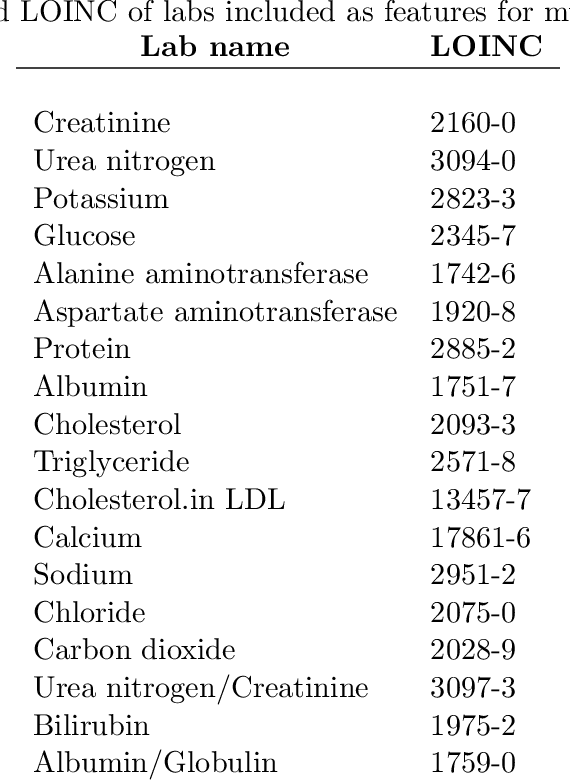
Disparate areas of machine learning have benefited from models that can take raw data with little preprocessing as input and learn rich representations of that raw data in order to perform well on a given prediction task. We evaluate this approach in healthcare by using longitudinal measurements of lab tests, one of the more raw signals of a patient's health state widely available in clinical data, to predict disease onsets. In particular, we train a Long Short-Term Memory (LSTM) recurrent neural network and two novel convolutional neural networks for multi-task prediction of disease onset for 133 conditions based on 18 common lab tests measured over time in a cohort of 298K patients derived from 8 years of administrative claims data. We compare the neural networks to a logistic regression with several hand-engineered, clinically relevant features. We find that the representation-based learning approaches significantly outperform this baseline. We believe that our work suggests a new avenue for patient risk stratification based solely on lab results.
Identifiable Phenotyping using Constrained Non-Negative Matrix Factorization
Sep 20, 2016Shalmali Joshi, Suriya Gunasekar, David Sontag, Joydeep Ghosh
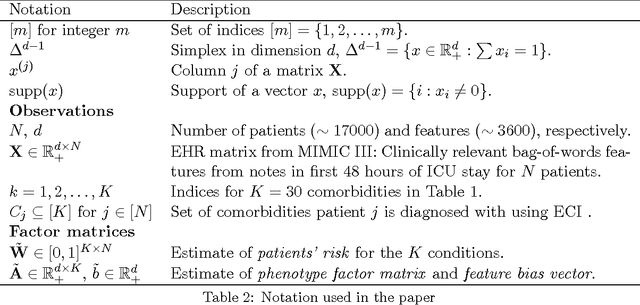

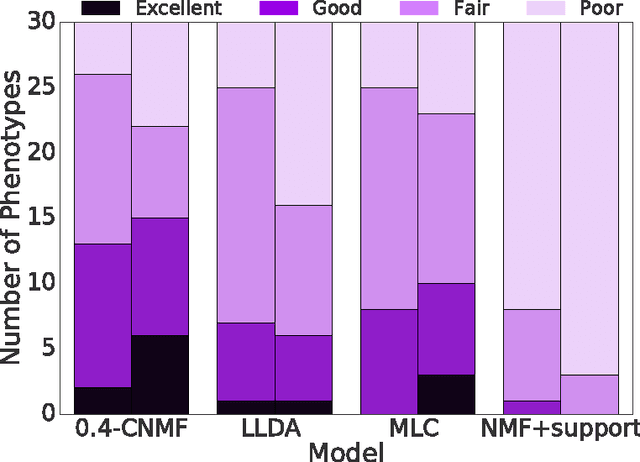

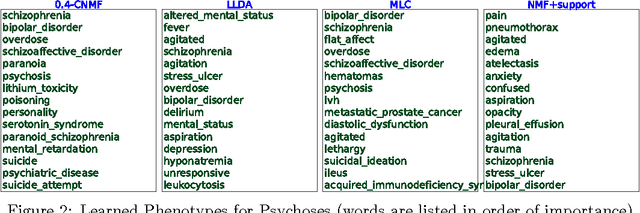
This work proposes a new algorithm for automated and simultaneous phenotyping of multiple co-occurring medical conditions, also referred as comorbidities, using clinical notes from the electronic health records (EHRs). A basic latent factor estimation technique of non-negative matrix factorization (NMF) is augmented with domain specific constraints to obtain sparse latent factors that are anchored to a fixed set of chronic conditions. The proposed anchoring mechanism ensures a one-to-one identifiable and interpretable mapping between the latent factors and the target comorbidities. Qualitative assessment of the empirical results by clinical experts suggests that the proposed model learns clinically interpretable phenotypes while being predictive of 30 day mortality. The proposed method can be readily adapted to any non-negative EHR data across various healthcare institutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge