Christoph Walz
Beyond Attention Heatmaps: How to Get Better Explanations for Multiple Instance Learning Models in Histopathology
Mar 09, 2026Abstract:Multiple instance learning (MIL) has enabled substantial progress in computational histopathology, where a large amount of patches from gigapixel whole slide images are aggregated into slide-level predictions. Heatmaps are widely used to validate MIL models and to discover tissue biomarkers. Yet, the validity of these heatmaps has barely been investigated. In this work, we introduce a general framework for evaluating the quality of MIL heatmaps without requiring additional labels. We conduct a large-scale benchmark experiment to assess six explanation methods across histopathology task types (classification, regression, survival), MIL model architectures (Attention-, Transformer-, Mamba-based), and patch encoder backbones (UNI2, Virchow2). Our results show that explanation quality mostly depends on MIL model architecture and task type, with perturbation ("Single"), layer-wise relevance propagation (LRP), and integrated gradients (IG) consistently outperforming attention-based and gradient-based saliency heatmaps, which often fail to reflect model decision mechanisms. We further demonstrate the advanced capabilities of the best-performing explanation methods: (i) We provide a proof-of-concept that MIL heatmaps of a bulk gene expression prediction model can be correlated with spatial transcriptomics for biological validation, and (ii) showcase the discovery of distinct model strategies for predicting human papillomavirus (HPV) infection from head and neck cancer slides. Our work highlights the importance of validating MIL heatmaps and establishes that improved explainability can enable more reliable model validation and yield biological insights, making a case for a broader adoption of explainable AI in digital pathology. Our code is provided in a public GitHub repository: https://github.com/bifold-pathomics/xMIL/tree/xmil-journal
BACH: Grand Challenge on Breast Cancer Histology Images
Aug 13, 2018
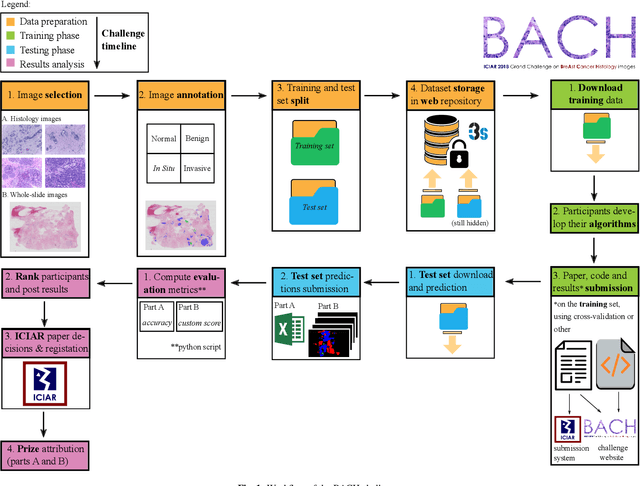
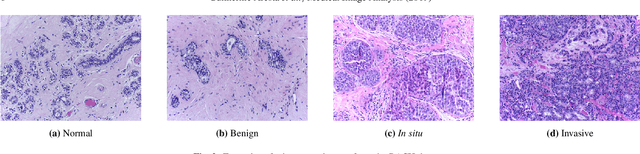
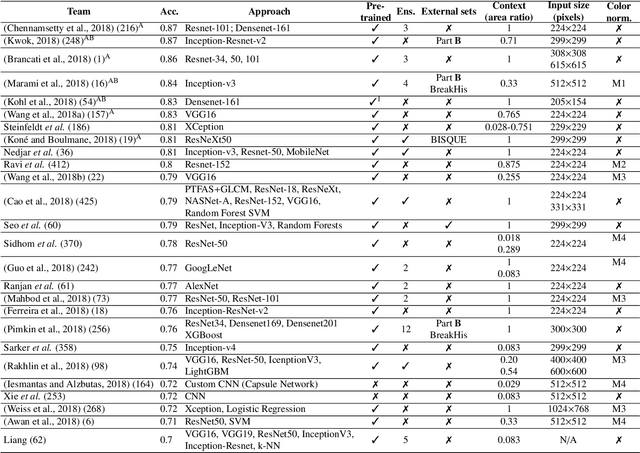
Abstract:Breast cancer is the most common invasive cancer in women, affecting more than 10% of women worldwide. Microscopic analysis of a biopsy remains one of the most important methods to diagnose the type of breast cancer. This requires specialized analysis by pathologists, in a task that i) is highly time- and cost-consuming and ii) often leads to nonconsensual results. The relevance and potential of automatic classification algorithms using hematoxylin-eosin stained histopathological images has already been demonstrated, but the reported results are still sub-optimal for clinical use. With the goal of advancing the state-of-the-art in automatic classification, the Grand Challenge on BreAst Cancer Histology images (BACH) was organized in conjunction with the 15th International Conference on Image Analysis and Recognition (ICIAR 2018). A large annotated dataset, composed of both microscopy and whole-slide images, was specifically compiled and made publicly available for the BACH challenge. Following a positive response from the scientific community, a total of 64 submissions, out of 677 registrations, effectively entered the competition. From the submitted algorithms it was possible to push forward the state-of-the-art in terms of accuracy (87%) in automatic classification of breast cancer with histopathological images. Convolutional neuronal networks were the most successful methodology in the BACH challenge. Detailed analysis of the collective results allowed the identification of remaining challenges in the field and recommendations for future developments. The BACH dataset remains publically available as to promote further improvements to the field of automatic classification in digital pathology.
Assessment of Breast Cancer Histology using Densely Connected Convolutional Networks
Apr 09, 2018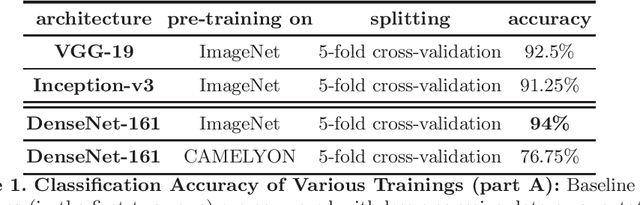

Abstract:Breast cancer is the most frequently diagnosed cancer and leading cause of cancer-related death among females worldwide. In this article, we investigate the applicability of densely connected convolutional neural networks to the problems of histology image classification and whole slide image segmentation in the area of computer-aided diagnoses for breast cancer. To this end, we study various approaches for transfer learning and apply them to the data set from the 2018 grand challenge on breast cancer histology images (BACH).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge