Christian Igel
Energy Consumption-Aware Tabular Benchmarks for Neural Architecture Search
Oct 12, 2022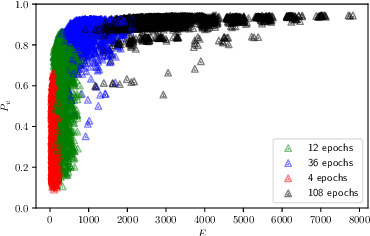
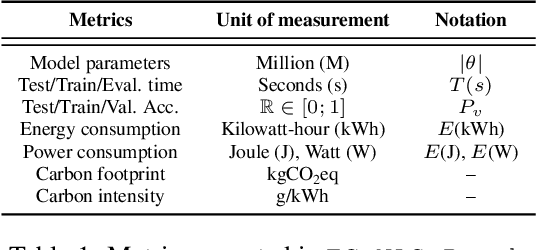

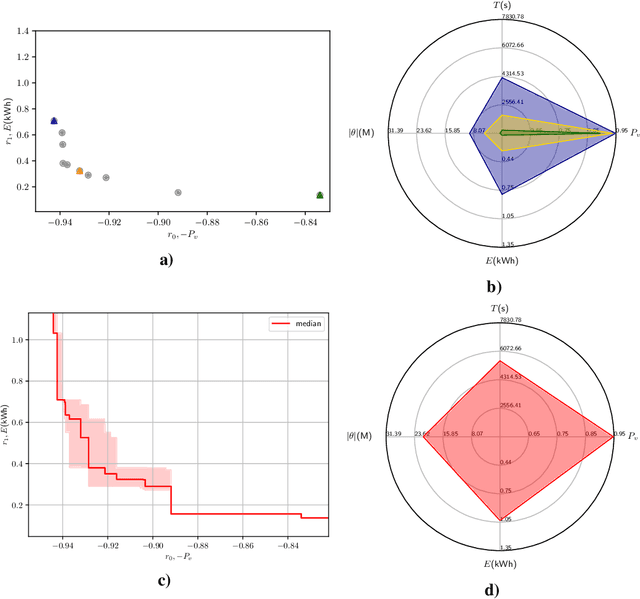
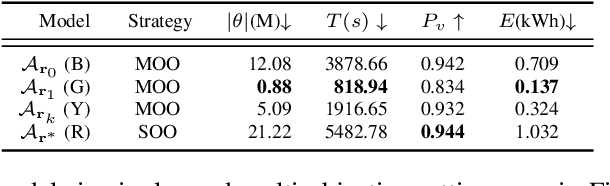
Abstract:The demand for large-scale computational resources for Neural Architecture Search (NAS) has been lessened by tabular benchmarks for NAS. Evaluating NAS strategies is now possible on extensive search spaces and at a moderate computational cost. But so far, NAS has mainly focused on maximising performance on some hold-out validation/test set. However, energy consumption is a partially conflicting objective that should not be neglected. We hypothesise that constraining NAS to include the energy consumption of training the models could reveal a sub-space of undiscovered architectures that are more computationally efficient with a smaller carbon footprint. To support the hypothesis, an existing tabular benchmark for NAS is augmented with the energy consumption of each architecture. We then perform multi-objective optimisation that includes energy consumption as an additional objective. We demonstrate the usefulness of multi-objective NAS for uncovering the trade-off between performance and energy consumption as well as for finding more energy-efficient architectures. The updated tabular benchmark, EC-NAS-Bench, is open-sourced to encourage the further exploration of energy consumption-aware NAS.
Self-Supervised Speech Representation Learning: A Review
May 21, 2022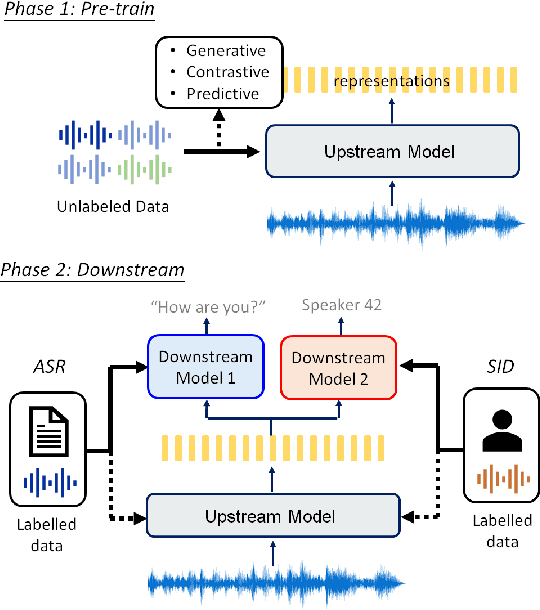
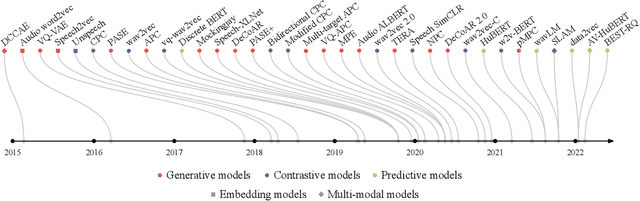
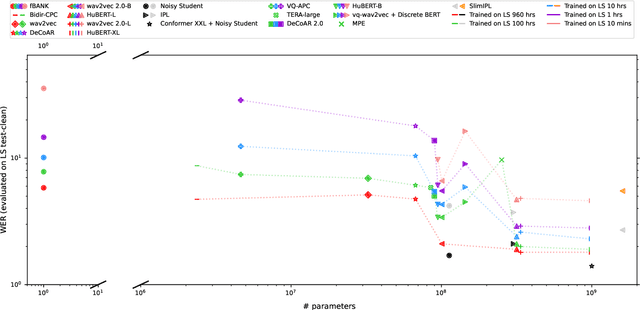
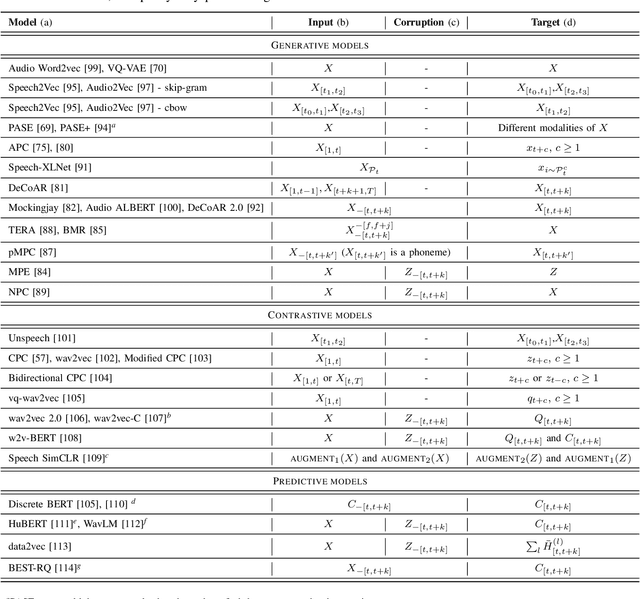
Abstract:Although supervised deep learning has revolutionized speech and audio processing, it has necessitated the building of specialist models for individual tasks and application scenarios. It is likewise difficult to apply this to dialects and languages for which only limited labeled data is available. Self-supervised representation learning methods promise a single universal model that would benefit a wide variety of tasks and domains. Such methods have shown success in natural language processing and computer vision domains, achieving new levels of performance while reducing the number of labels required for many downstream scenarios. Speech representation learning is experiencing similar progress in three main categories: generative, contrastive, and predictive methods. Other approaches rely on multi-modal data for pre-training, mixing text or visual data streams with speech. Although self-supervised speech representation is still a nascent research area, it is closely related to acoustic word embedding and learning with zero lexical resources, both of which have seen active research for many years. This review presents approaches for self-supervised speech representation learning and their connection to other research areas. Since many current methods focus solely on automatic speech recognition as a downstream task, we review recent efforts on benchmarking learned representations to extend the application beyond speech recognition.
Remember to correct the bias when using deep learning for regression!
Mar 30, 2022
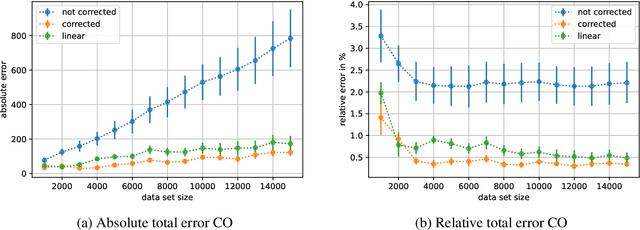

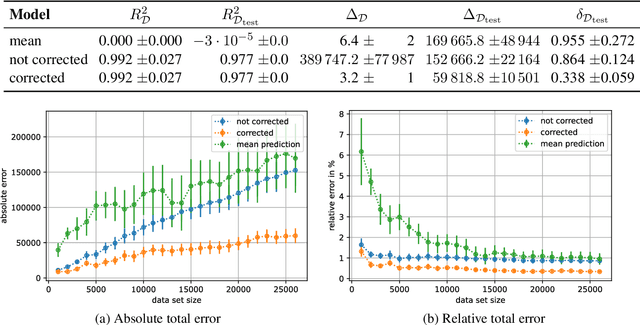
Abstract:When training deep learning models for least-squares regression, we cannot expect that the training error residuals of the final model, selected after a fixed training time or based on performance on a hold-out data set, sum to zero. This can introduce a systematic error that accumulates if we are interested in the total aggregated performance over many data points. We suggest to adjust the bias of the machine learning model after training as a default postprocessing step, which efficiently solves the problem. The severeness of the error accumulation and the effectiveness of the bias correction is demonstrated in exemplary experiments.
A Brief Overview of Unsupervised Neural Speech Representation Learning
Mar 01, 2022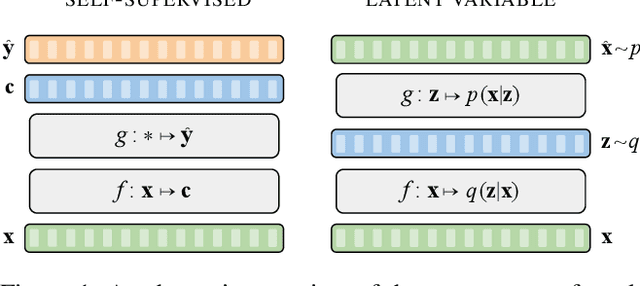
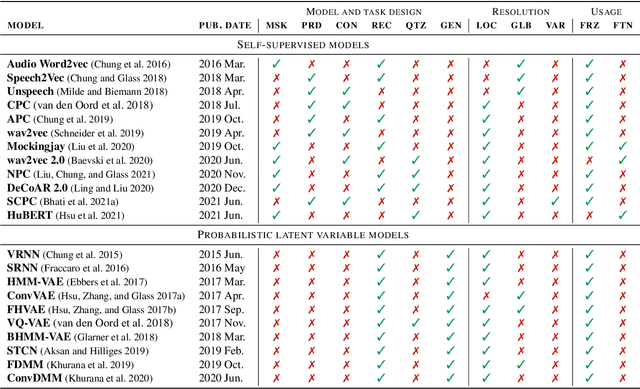
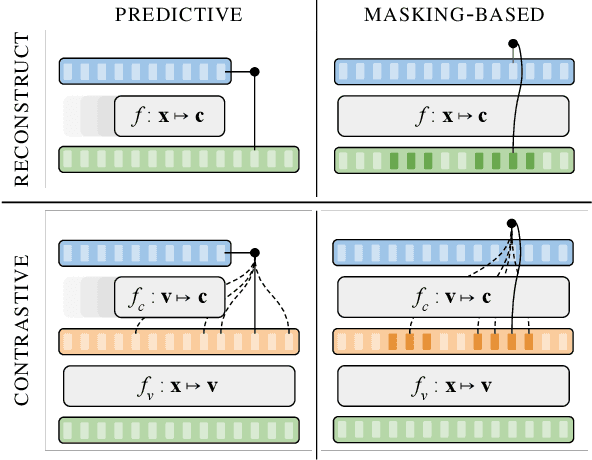
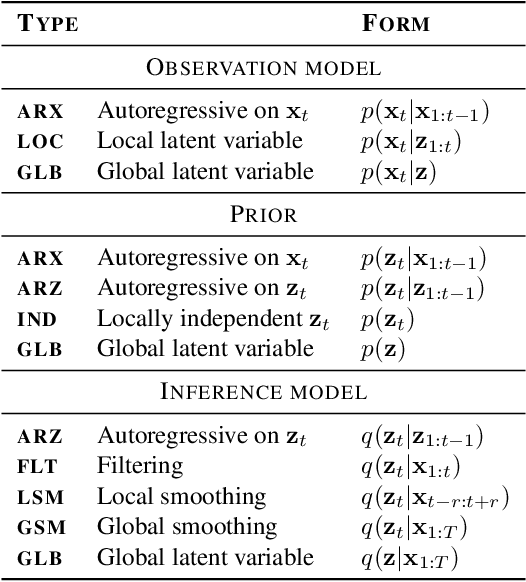
Abstract:Unsupervised representation learning for speech processing has matured greatly in the last few years. Work in computer vision and natural language processing has paved the way, but speech data offers unique challenges. As a result, methods from other domains rarely translate directly. We review the development of unsupervised representation learning for speech over the last decade. We identify two primary model categories: self-supervised methods and probabilistic latent variable models. We describe the models and develop a comprehensive taxonomy. Finally, we discuss and compare models from the two categories.
Deep Learning Based 3D Point Cloud Regression for Estimating Forest Biomass
Dec 22, 2021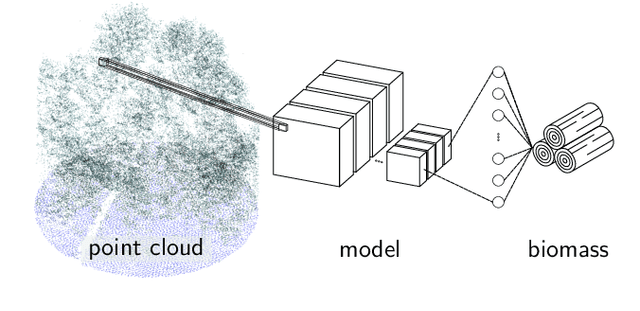
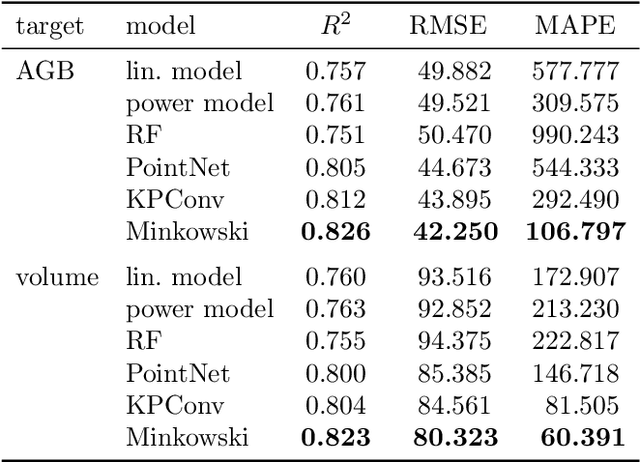
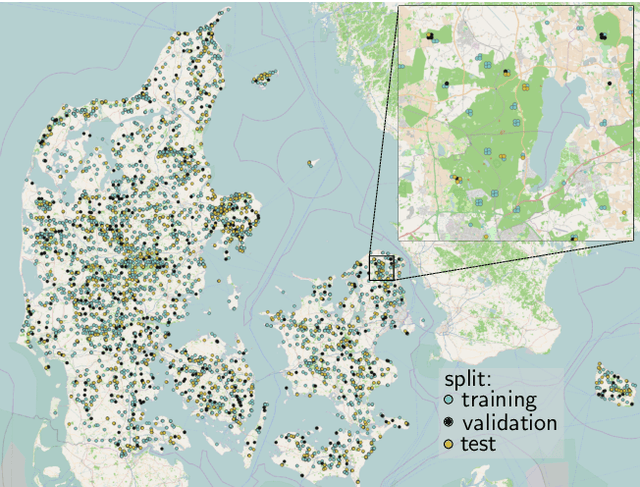
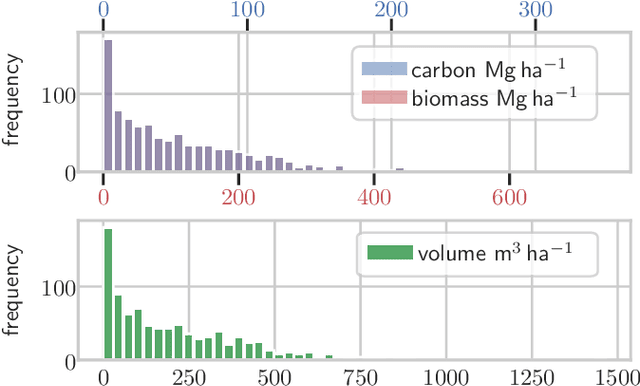
Abstract:Knowledge of forest biomass stocks and their development is important for implementing effective climate change mitigation measures. It is needed for studying the processes driving af-, re-, and deforestation and is a prerequisite for carbon-accounting. Remote sensing using airborne LiDAR can be used to measure vegetation biomass at large scale. We present deep learning systems for predicting wood volume, above-ground biomass (AGB), and subsequently carbon directly from 3D LiDAR point cloud data. We devise different neural network architectures for point cloud regression and evaluate them on remote sensing data of areas for which AGB estimates have been obtained from field measurements in a national forest inventory. Our adaptation of Minkowski convolutional neural networks for regression gave the best results. The deep neural networks produced significantly more accurate wood volume, AGB, and carbon estimates compared to state-of-the-art approaches operating on basic statistics of the point clouds, and we expect this finding to have a strong impact on LiDAR-based analyses of terrestrial ecosystem dynamics.
Do We Still Need Automatic Speech Recognition for Spoken Language Understanding?
Nov 29, 2021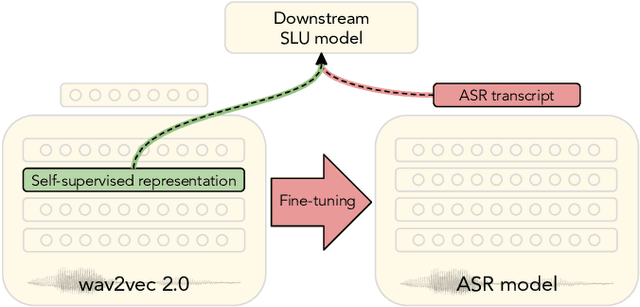

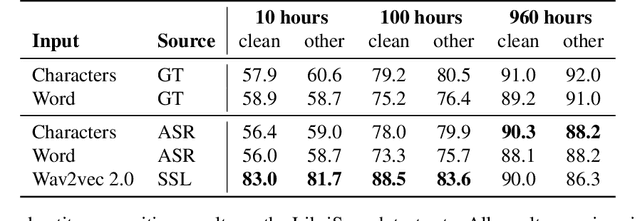

Abstract:Spoken language understanding (SLU) tasks are usually solved by first transcribing an utterance with automatic speech recognition (ASR) and then feeding the output to a text-based model. Recent advances in self-supervised representation learning for speech data have focused on improving the ASR component. We investigate whether representation learning for speech has matured enough to replace ASR in SLU. We compare learned speech features from wav2vec 2.0, state-of-the-art ASR transcripts, and the ground truth text as input for a novel speech-based named entity recognition task, a cardiac arrest detection task on real-world emergency calls and two existing SLU benchmarks. We show that learned speech features are superior to ASR transcripts on three classification tasks. For machine translation, ASR transcripts are still the better choice. We highlight the intrinsic robustness of wav2vec 2.0 representations to out-of-vocabulary words as key to better performance.
Chebyshev-Cantelli PAC-Bayes-Bennett Inequality for the Weighted Majority Vote
Jun 25, 2021
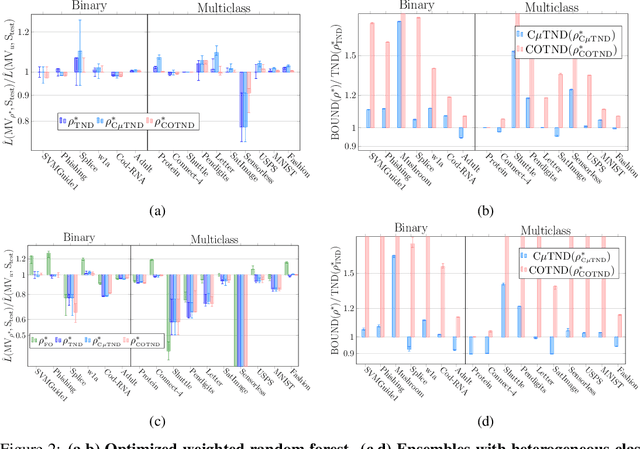
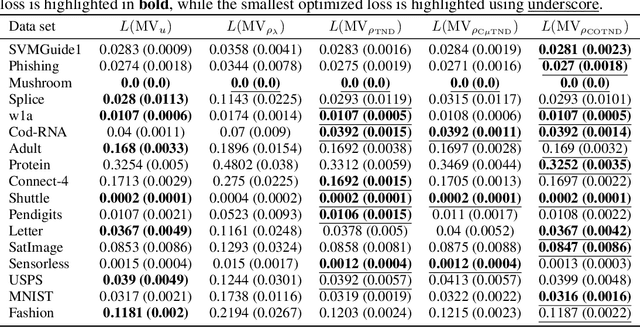
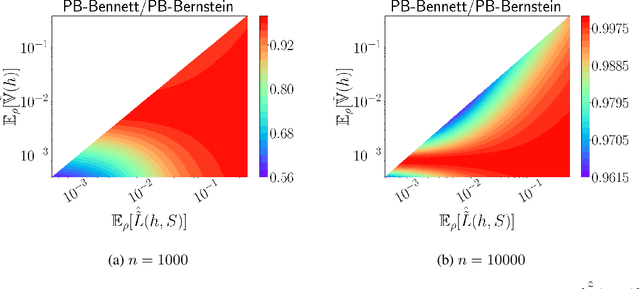
Abstract:We present a new second-order oracle bound for the expected risk of a weighted majority vote. The bound is based on a novel parametric form of the Chebyshev-Cantelli inequality (a.k.a.\ one-sided Chebyshev's), which is amenable to efficient minimization. The new form resolves the optimization challenge faced by prior oracle bounds based on the Chebyshev-Cantelli inequality, the C-bounds [Germain et al., 2015], and, at the same time, it improves on the oracle bound based on second order Markov's inequality introduced by Masegosa et al. [2020]. We also derive the PAC-Bayes-Bennett inequality, which we use for empirical estimation of the oracle bound. The PAC-Bayes-Bennett inequality improves on the PAC-Bayes-Bernstein inequality by Seldin et al. [2012]. We provide an empirical evaluation demonstrating that the new bounds can improve on the work by Masegosa et al. [2020]. Both the parametric form of the Chebyshev-Cantelli inequality and the PAC-Bayes-Bennett inequality may be of independent interest for the study of concentration of measure in other domains.
Information Bottleneck: Exact Analysis of (Quantized) Neural Networks
Jun 24, 2021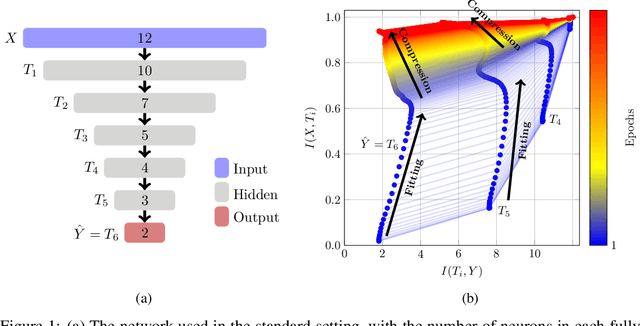
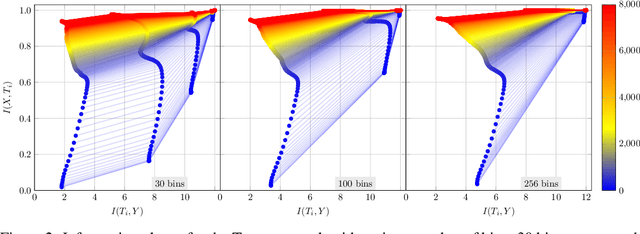
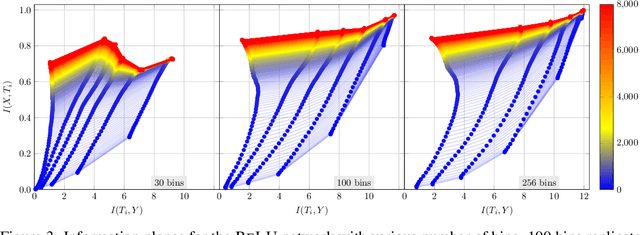
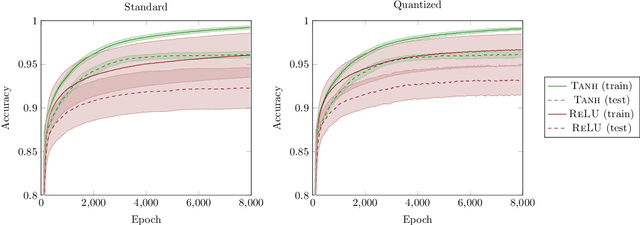
Abstract:The information bottleneck (IB) principle has been suggested as a way to analyze deep neural networks. The learning dynamics are studied by inspecting the mutual information (MI) between the hidden layers and the input and output. Notably, separate fitting and compression phases during training have been reported. This led to some controversy including claims that the observations are not reproducible and strongly dependent on the type of activation function used as well as on the way the MI is estimated. Our study confirms that different ways of binning when computing the MI lead to qualitatively different results, either supporting or refusing IB conjectures. To resolve the controversy, we study the IB principle in settings where MI is non-trivial and can be computed exactly. We monitor the dynamics of quantized neural networks, that is, we discretize the whole deep learning system so that no approximation is required when computing the MI. This allows us to quantify the information flow without measurement errors. In this setting, we observed a fitting phase for all layers and a compression phase for the output layer in all experiments; the compression in the hidden layers was dependent on the type of activation function. Our study shows that the initial IB results were not artifacts of binning when computing the MI. However, the critical claim that the compression phase may not be observed for some networks also holds true.
Do End-to-End Speech Recognition Models Care About Context?
Feb 17, 2021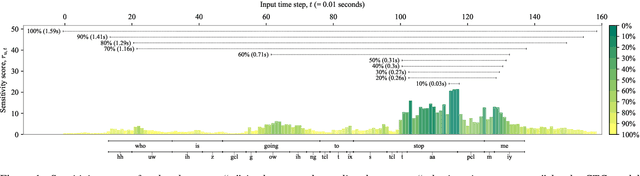
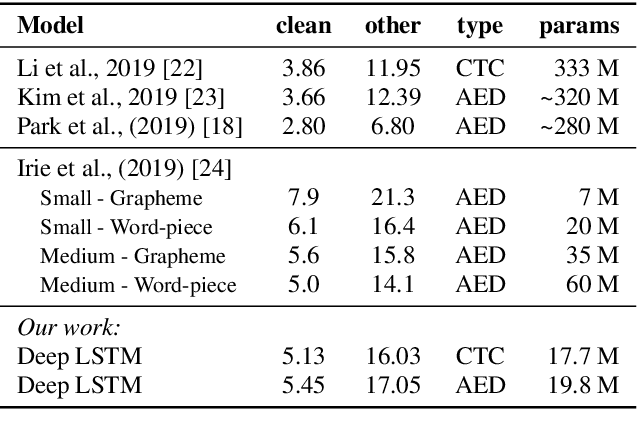
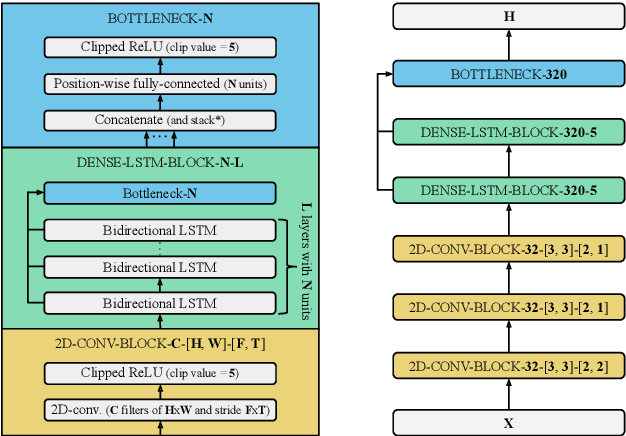
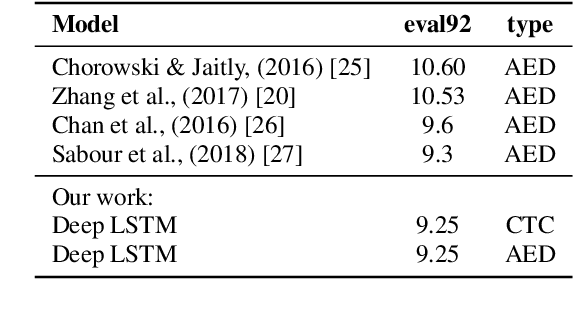
Abstract:The two most common paradigms for end-to-end speech recognition are connectionist temporal classification (CTC) and attention-based encoder-decoder (AED) models. It has been argued that the latter is better suited for learning an implicit language model. We test this hypothesis by measuring temporal context sensitivity and evaluate how the models perform when we constrain the amount of contextual information in the audio input. We find that the AED model is indeed more context sensitive, but that the gap can be closed by adding self-attention to the CTC model. Furthermore, the two models perform similarly when contextual information is constrained. Finally, in contrast to previous research, our results show that the CTC model is highly competitive on WSJ and LibriSpeech without the help of an external language model.
On Scaling Contrastive Representations for Low-Resource Speech Recognition
Feb 01, 2021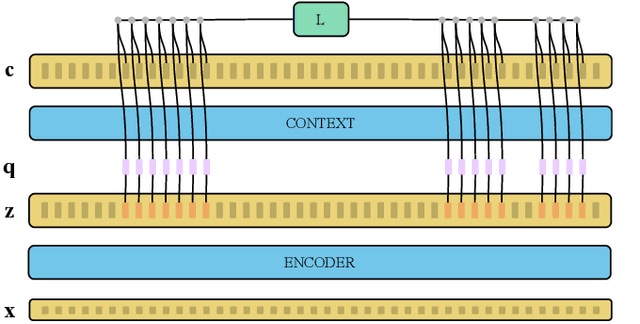
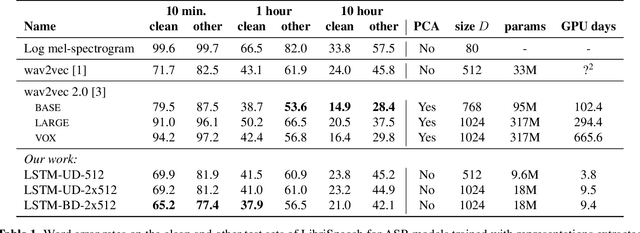
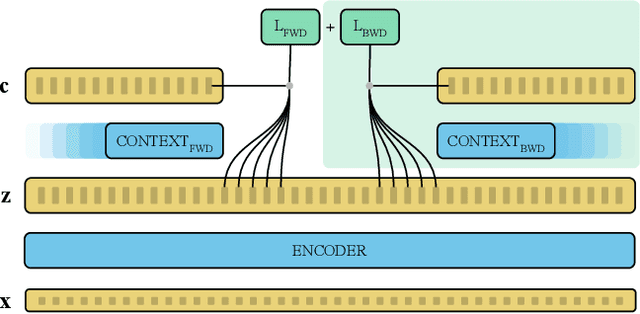
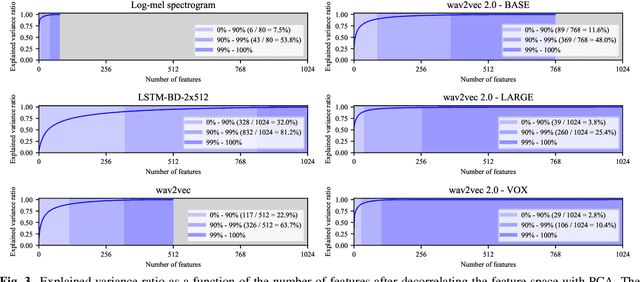
Abstract:Recent advances in self-supervised learning through contrastive training have shown that it is possible to learn a competitive speech recognition system with as little as 10 minutes of labeled data. However, these systems are computationally expensive since they require pre-training followed by fine-tuning in a large parameter space. We explore the performance of such systems without fine-tuning by training a state-of-the-art speech recognizer on the fixed representations from the computationally demanding wav2vec 2.0 framework. We find performance to decrease without fine-tuning and, in the extreme low-resource setting, wav2vec 2.0 is inferior to its predecessor. In addition, we find that wav2vec 2.0 representations live in a low dimensional subspace and that decorrelating the features of the representations can stabilize training of the automatic speech recognizer. Finally, we propose a bidirectional extension to the original wav2vec framework that consistently improves performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge