Bin Chong
Mitigating Homophily Disparity in Graph Anomaly Detection: A Scalable and Adaptive Approach
Mar 09, 2026Abstract:Graph anomaly detection (GAD) aims to identify nodes that deviate from normal patterns in structure or features. While recent GNN-based approaches have advanced this task, they struggle with two major challenges: 1) homophily disparity, where nodes exhibit varying homophily at both class and node levels; and 2) limited scalability, as many methods rely on costly whole-graph operations. To address them, we propose SAGAD, a Scalable and Adaptive framework for GAD. SAGAD precomputes multi-hop embeddings and applies reparameterized Chebyshev filters to extract low- and high-frequency information, enabling efficient training and capturing both homophilic and heterophilic patterns. To mitigate node-level homophily disparity, we introduce an Anomaly Context-Aware Adaptive Fusion, which adaptively fuses low- and high-pass embeddings using fusion coefficients conditioned on Rayleigh Quotient-guided anomalous subgraph structures for each node. To alleviate class-level disparity, we design a Frequency Preference Guidance Loss, which encourages anomalies to preserve more high-frequency information than normal nodes. SAGAD supports mini-batch training, achieves linear time and space complexity, and drastically reduces memory usage on large-scale graphs. Theoretically, SAGAD ensures asymptotic linear separability between normal and abnormal nodes under mild conditions. Extensive experiments on 10 benchmarks confirm SAGAD's superior accuracy and scalability over state-of-the-art methods.
Reversible Upper Confidence Bound Algorithm to Generate Diverse Optimized Candidates
Dec 30, 2021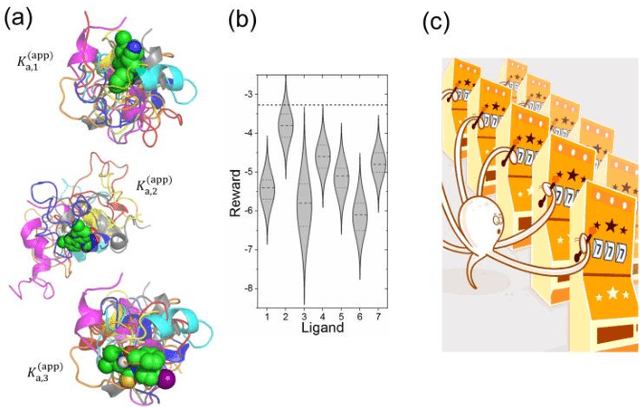
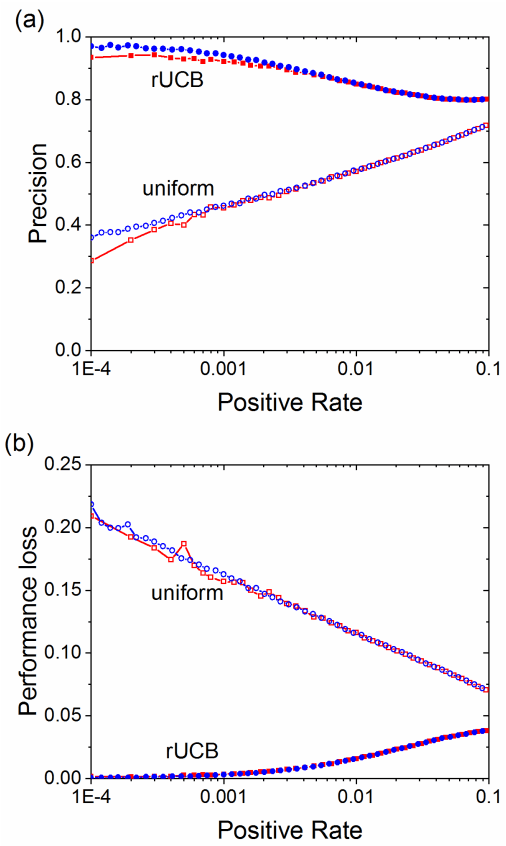
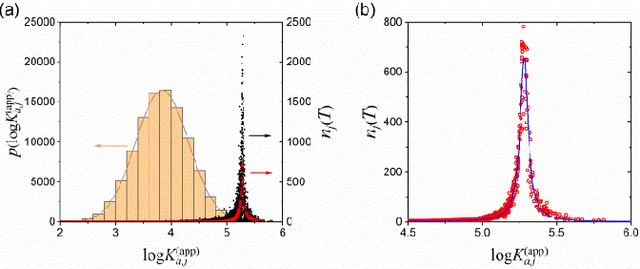
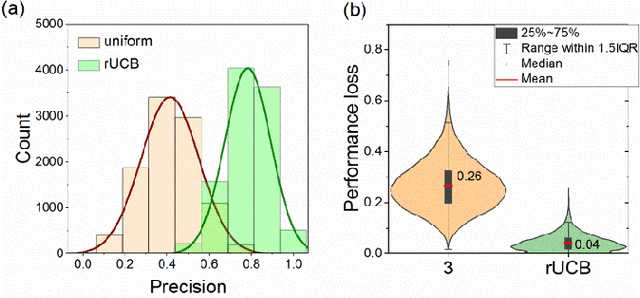
Abstract:Most algorithms for the multi-armed bandit problem in reinforcement learning aimed to maximize the expected reward, which are thus useful in searching the optimized candidate with the highest reward (function value) for diverse applications (e.g., AlphaGo). However, in some typical application scenaios such as drug discovery, the aim is to search a diverse set of candidates with high reward. Here we propose a reversible upper confidence bound (rUCB) algorithm for such a purpose, and demonstrate its application in virtual screening upon intrinsically disordered proteins (IDPs). It is shown that rUCB greatly reduces the query times while achieving both high accuracy and low performance loss.The rUCB may have potential application in multipoint optimization and other reinforcement-learning cases.
* 10 pages, 10 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge