Avi Z. Rosenberg
Johns Hopkins University
A Comprehensive Benchmark of Histopathology Foundation Models for Kidney Histopathology
Mar 16, 2026Abstract:Histopathology foundation models (HFMs), pretrained on large-scale cancer datasets, have advanced computational pathology. However, their applicability to non-cancerous chronic kidney disease remains underexplored, despite coexistence of renal pathology with malignancies such as renal cell and urothelial carcinoma. We systematically evaluate 11 publicly available HFMs across 11 kidney-specific downstream tasks spanning multiple stains (PAS, H&E, PASM, and IHC), spatial scales (tile and slide-level), task types (classification, regression, and copy detection), and clinical objectives, including detection, diagnosis, and prognosis. Tile-level performance is assessed using repeated stratified group cross-validation, while slide-level tasks are evaluated using repeated nested stratified cross-validation. Statistical significance is examined using Friedman test followed by pairwise Wilcoxon signed-rank testing with Holm-Bonferroni correction and compact letter display visualization. To promote reproducibility, we release an open-source Python package, kidney-hfm-eval, available at https://pypi.org/project/kidney-hfm-eval/ , that reproduces the evaluation pipelines. Results show moderate to strong performance on tasks driven by coarse meso-scale renal morphology, including diagnostic classification and detection of prominent structural alterations. In contrast, performance consistently declines for tasks requiring fine-grained microstructural discrimination, complex biological phenotypes, or slide-level prognostic inference, largely independent of stain type. Overall, current HFMs appear to encode predominantly static meso-scale representations and may have limited capacity to capture subtle renal pathology or prognosis-related signals. Our results highlight the need for kidney-specific, multi-stain, and multimodal foundation models to support clinically reliable decision-making in nephrology.
HistoWAS: A Pathomics Framework for Large-Scale Feature-Wide Association Studies of Tissue Topology and Patient Outcomes
Dec 23, 2025Abstract:High-throughput "pathomic" analysis of Whole Slide Images (WSIs) offers new opportunities to study tissue characteristics and for biomarker discovery. However, the clinical relevance of the tissue characteristics at the micro- and macro-environment level is limited by the lack of tools that facilitate the measurement of the spatial interaction of individual structure characteristics and their association with clinical parameters. To address these challenges, we introduce HistoWAS (Histology-Wide Association Study), a computational framework designed to link tissue spatial organization to clinical outcomes. Specifically, HistoWAS implements (1) a feature space that augments conventional metrics with 30 topological and spatial features, adapted from Geographic Information Systems (GIS) point pattern analysis, to quantify tissue micro-architecture; and (2) an association study engine, inspired by Phenome-Wide Association Studies (PheWAS), that performs mass univariate regression for each feature with statistical correction. As a proof of concept, we applied HistoWAS to analyze a total of 102 features (72 conventional object-level features and our 30 spatial features) using 385 PAS-stained WSIs from 206 participants in the Kidney Precision Medicine Project (KPMP). The code and data have been released to https://github.com/hrlblab/histoWAS.
Histo-fetch -- On-the-fly processing of gigapixel whole slide images simplifies and speeds neural network training
Mar 01, 2021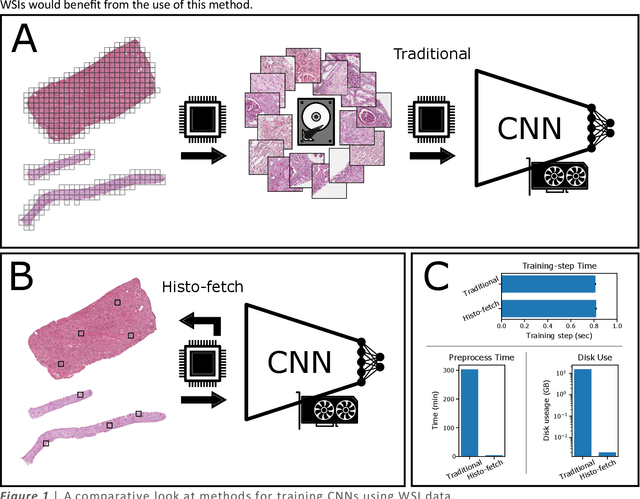
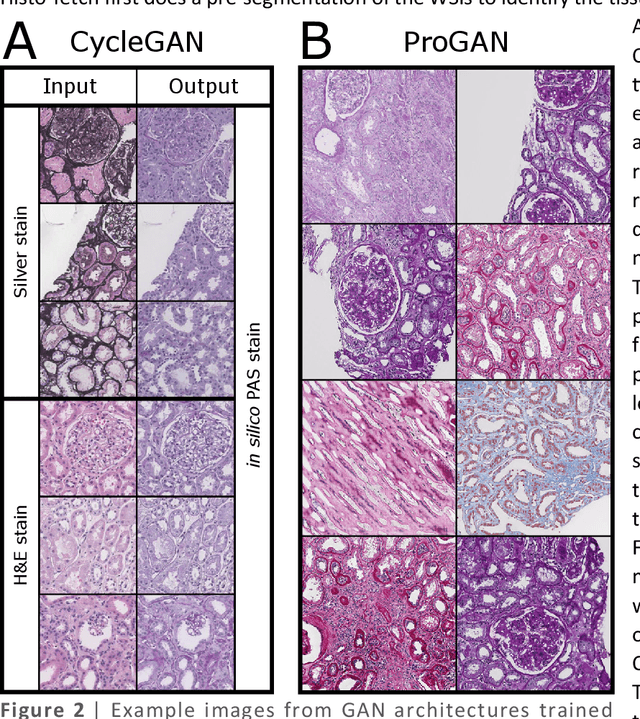
Abstract:We created a custom pipeline (histo-fetch) to efficiently extract random patches and labels from pathology whole slide images (WSIs) for input to a neural network on-the-fly. We prefetch these patches as needed during network training, avoiding the need for WSI preparation such as chopping/tiling. We demonstrate the utility of this pipeline to perform artificial stain transfer and image generation using the popular networks CycleGAN and ProGAN, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge