Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
Effect of Random Histogram Equalization on Breast Calcification Analysis Using Deep Learning
May 03, 2022
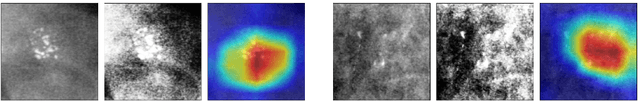
Abstract:Early detection and analysis of calcifications in mammogram images is crucial in a breast cancer diagnosis workflow. Management of calcifications that require immediate follow-up and further analyzing its benignancy or malignancy can result in a better prognosis. Recent studies have shown that deep learning-based algorithms can learn robust representations to analyze suspicious calcifications in mammography. In this work, we demonstrate that randomly equalizing the histograms of calcification patches as a data augmentation technique can significantly improve the classification performance for analyzing suspicious calcifications. We validate our approach by using the CBIS-DDSM dataset for two classification tasks. The results on both the tasks show that the proposed methodology gains more than 1% mean accuracy and F1-score when equalizing the data with a probability of 0.4 when compared to not using histogram equalization. This is further supported by the t-tests, where we obtain a p-value of p<0.0001, thus showing the statistical significance of our approach.
Continual Learning for Peer-to-Peer Federated Learning: A Study on Automated Brain Metastasis Identification
Apr 30, 2022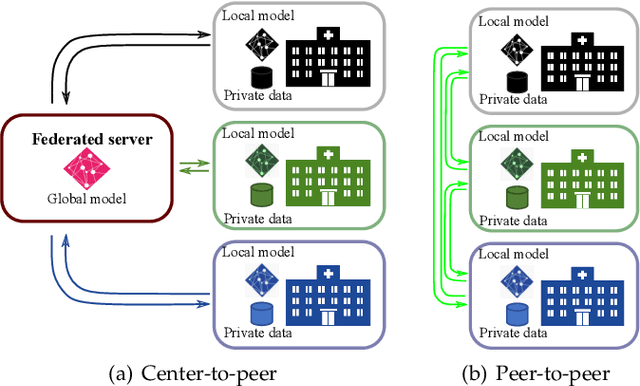
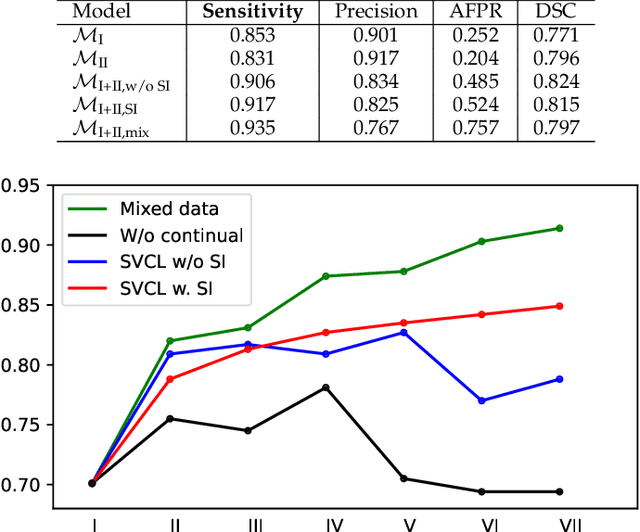
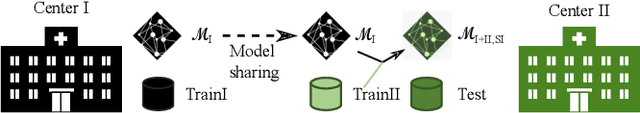
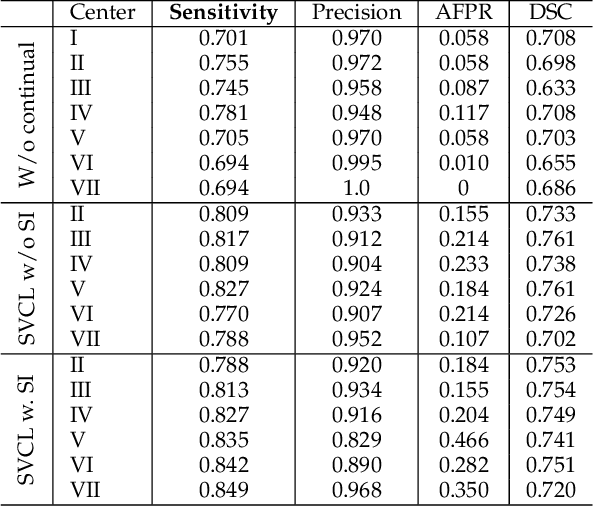
Abstract:Due to data privacy constraints, data sharing among multiple centers is restricted. Continual learning, as one approach to peer-to-peer federated learning, can promote multicenter collaboration on deep learning algorithm development by sharing intermediate models instead of training data. This work aims to investigate the feasibility of continual learning for multicenter collaboration on an exemplary application of brain metastasis identification using DeepMedic. 920 T1 MRI contrast enhanced volumes are split to simulate multicenter collaboration scenarios. A continual learning algorithm, synaptic intelligence (SI), is applied to preserve important model weights for training one center after another. In a bilateral collaboration scenario, continual learning with SI achieves a sensitivity of 0.917, and naive continual learning without SI achieves a sensitivity of 0.906, while two models trained on internal data solely without continual learning achieve sensitivity of 0.853 and 0.831 only. In a seven-center multilateral collaboration scenario, the models trained on internal datasets (100 volumes each center) without continual learning obtain a mean sensitivity value of 0.699. With single-visit continual learning (i.e., the shared model visits each center only once during training), the sensitivity is improved to 0.788 and 0.849 without SI and with SI, respectively. With iterative continual learning (i.e., the shared model revisits each center multiple times during training), the sensitivity is further improved to 0.914, which is identical to the sensitivity using mixed data for training. Our experiments demonstrate that continual learning can improve brain metastasis identification performance for centers with limited data. This study demonstrates the feasibility of applying continual learning for peer-to-peer federated learning in multicenter collaboration.
An Algorithm for the Labeling and Interactive Visualization of the Cerebrovascular System of Ischemic Strokes
Apr 26, 2022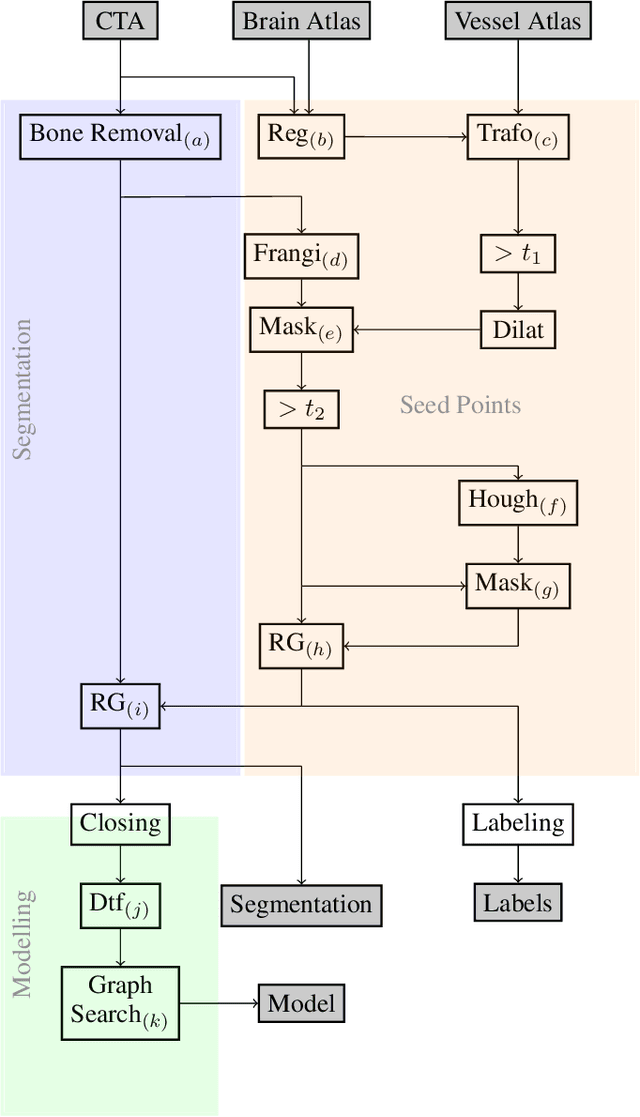
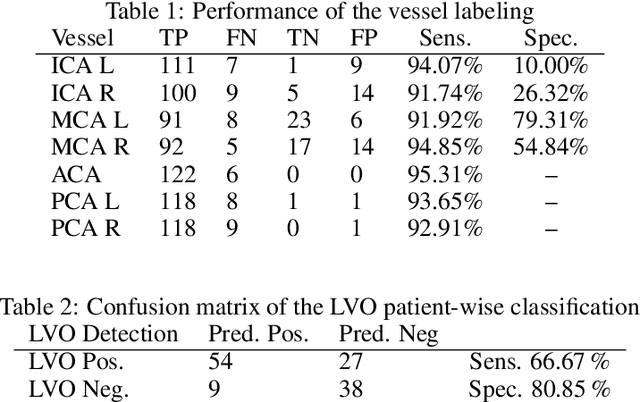
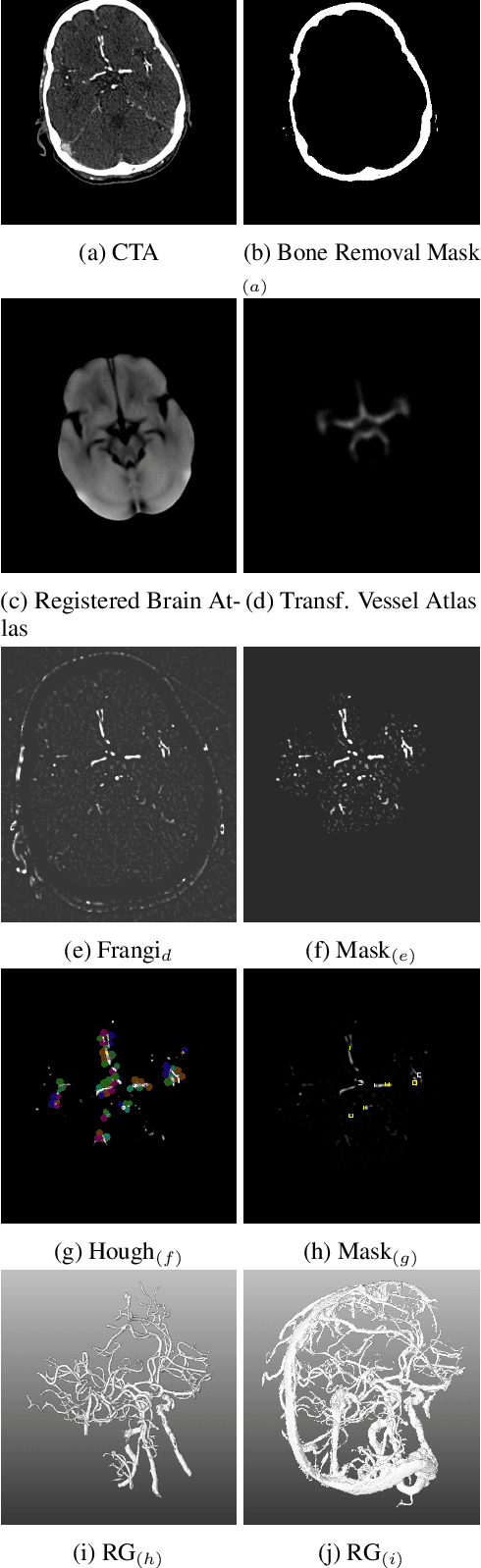
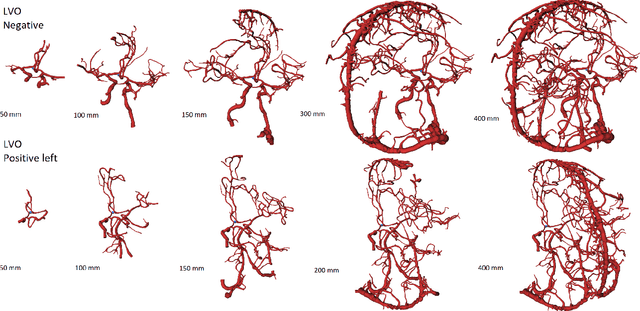
Abstract:During the diagnosis of ischemic strokes, the Circle of Willis and its surrounding vessels are the arteries of interest. Their visualization in case of an acute stroke is often enabled by Computed Tomography Angiography (CTA). Still, the identification and analysis of the cerebral arteries remain time consuming in such scans due to a large number of peripheral vessels which may disturb the visual impression. In previous work we proposed VirtualDSA++, an algorithm designed to segment and label the cerebrovascular tree on CTA scans. Especially with stroke patients, labeling is a delicate procedure, as in the worst case whole hemispheres may not be present due to impeded perfusion. Hence, we extended the labeling mechanism for the cerebral arteries to identify occluded vessels. In the work at hand, we place the algorithm in a clinical context by evaluating the labeling and occlusion detection on stroke patients, where we have achieved labeling sensitivities comparable to other works between 92\,\% and 95\,\%. To the best of our knowledge, ours is the first work to address labeling and occlusion detection at once, whereby a sensitivity of 67\,\% and a specificity of 81\,\% were obtained for the latter. VirtualDSA++ also automatically segments and models the intracranial system, which we further used in a deep learning driven follow up work. We present the generic concept of iterative systematic search for pathways on all nodes of said model, which enables new interactive features. Exemplary, we derive in detail, firstly, the interactive planning of vascular interventions like the mechanical thrombectomy and secondly, the interactive suppression of vessel structures that are not of interest in diagnosing strokes (like veins). We discuss both features as well as further possibilities emerging from the proposed concept.
Is Speech Pathology a Biomarker in Automatic Speaker Verification?
Apr 13, 2022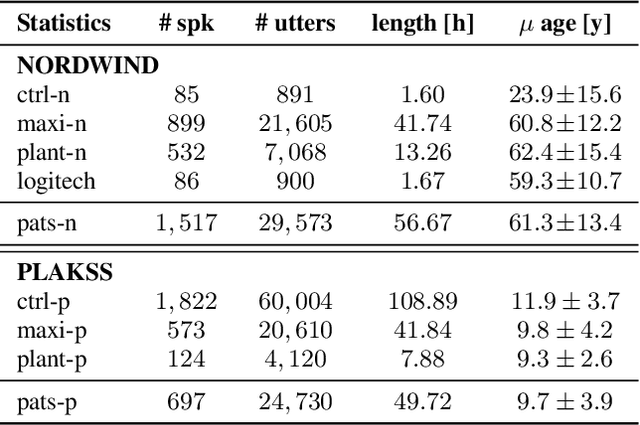
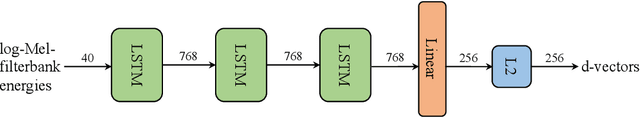
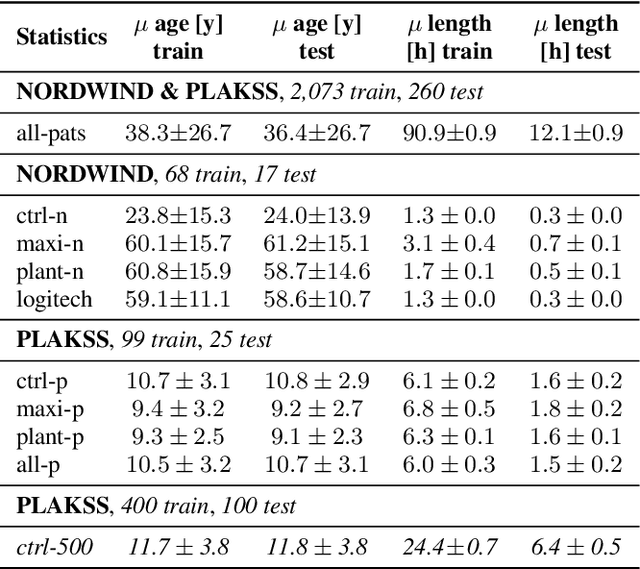
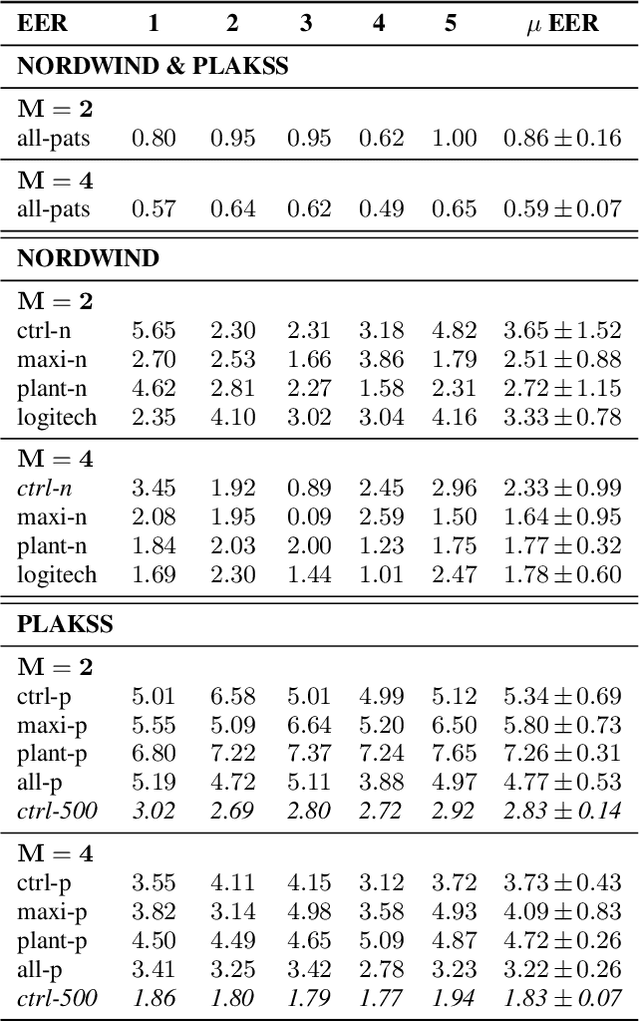
Abstract:With the advancements in deep learning (DL) and an increasing interest in data-driven speech processing methods, a major challenge for speech data scientists in the healthcare domain is the anonymization of pathological speech, which is a required step to be able to make them accessible as a public training resource. In this paper, we investigate pathological speech data and compare their speaker verifiability with that of healthy individuals. We utilize a large pathological speech corpus of more than 2,000 test subjects with various speech and voice disorders from different ages and apply DL-based automatic speaker verification (ASV) techniques. As a result, we obtained a mean equal error rate (EER) of 0.86% with a standard deviation of 0.16%, which is a factor of three lower than comparable healthy speech databases. We further perform detailed analyses of external influencing factors on ASV such as age, pathology, recording environment, and utterance length, to explore their respective effect. Our findings indicate that speech pathology is a potential biomarker in ASV. This is potentially of high interest for the anonymization of pathological speech data.
Disentangled Latent Speech Representation for Automatic Pathological Intelligibility Assessment
Apr 08, 2022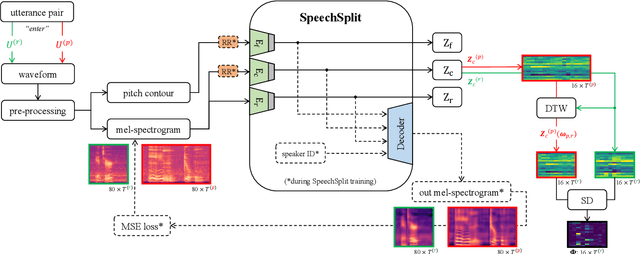
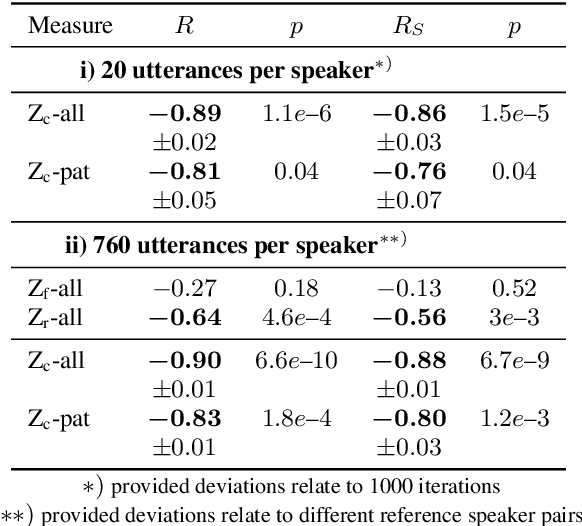
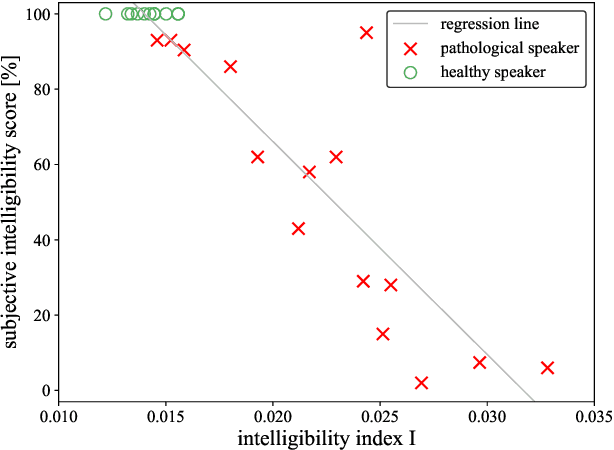
Abstract:Speech intelligibility assessment plays an important role in the therapy of patients suffering from pathological speech disorders. Automatic and objective measures are desirable to assist therapists in their traditionally subjective and labor-intensive assessments. In this work, we investigate a novel approach for obtaining such a measure using the divergence in disentangled latent speech representations of a parallel utterance pair, obtained from a healthy reference and a pathological speaker. Experiments on an English database of Cerebral Palsy patients, using all available utterances per speaker, show high and significant correlation values (R = -0.9) with subjective intelligibility measures, while having only minimal deviation (+-0.01) across four different reference speaker pairs. We also demonstrate the robustness of the proposed method (R = -0.89 deviating +-0.02 over 1000 iterations) by considering a significantly smaller amount of utterances per speaker. Our results are among the first to show that disentangled speech representations can be used for automatic pathological speech intelligibility assessment, resulting in a reference speaker pair invariant method, applicable in scenarios with only few utterances available.
Predictive Coding and Stochastic Resonance: Towards a Unified Theory of Auditory (Phantom) Perception
Apr 07, 2022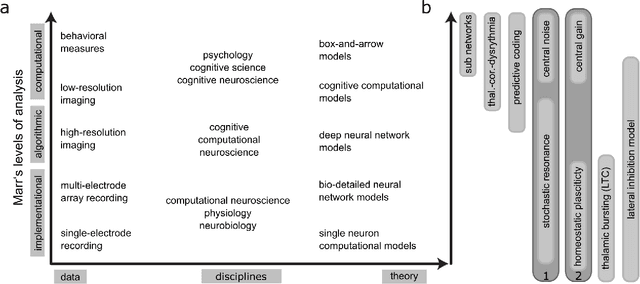
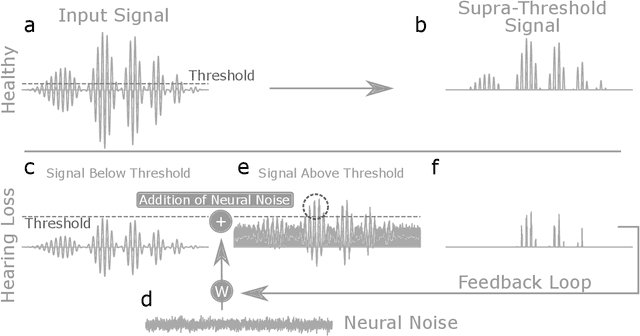
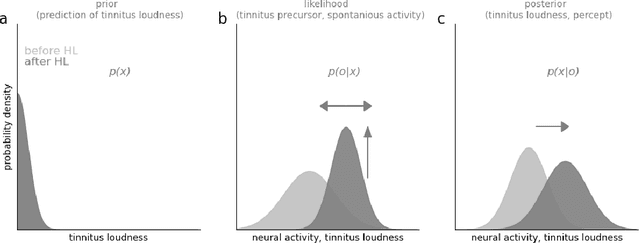
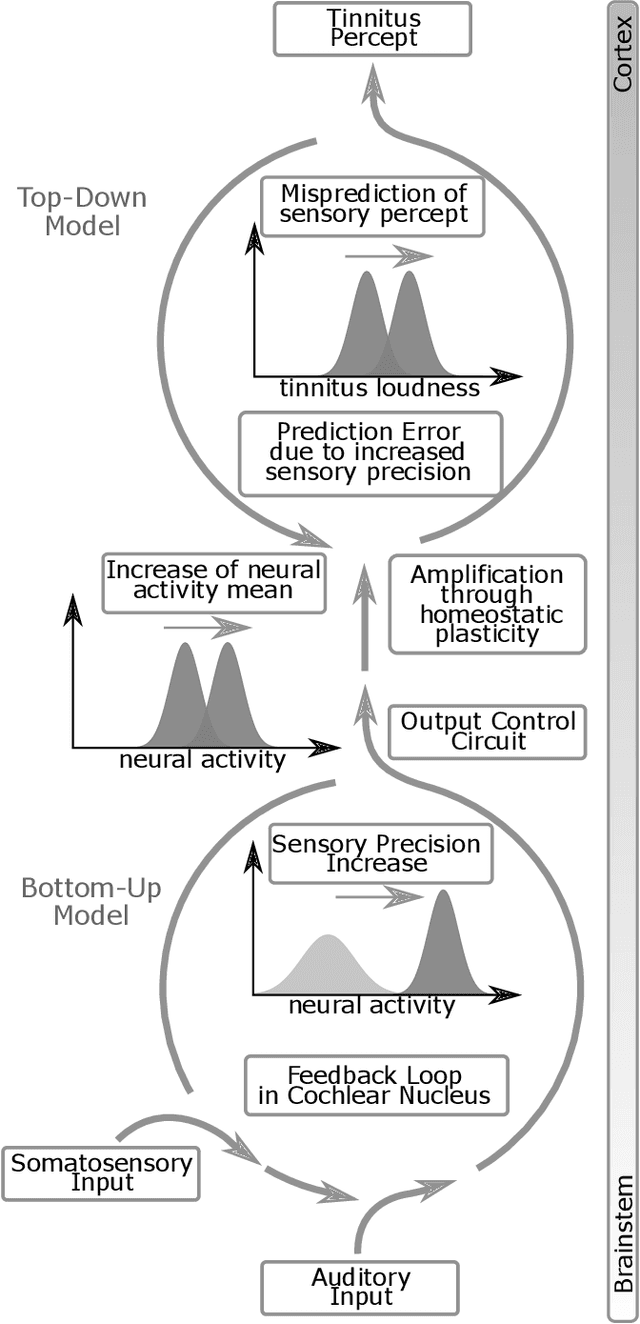
Abstract:Cognitive computational neuroscience (CCN) suggests that to gain a mechanistic understanding of brain function, hypothesis driven experiments should be accompanied by biologically plausible computational models. This novel research paradigm offers a way from alchemy to chemistry, in auditory neuroscience. With a special focus on tinnitus - as the prime example of auditory phantom perception - we review recent work at the intersection of artificial intelligence, psychology, and neuroscience, foregrounding the idea that experiments will yield mechanistic insight only when employed to test formal or computational models. This view challenges the popular notion that tinnitus research is primarily data limited, and that producing large, multi-modal, and complex data-sets, analyzed with advanced data analysis algorithms, will lead to fundamental insights into how tinnitus emerges. We conclude that two fundamental processing principles - being ubiquitous in the brain - best fit to a vast number of experimental results and therefore provide the most explanatory power: predictive coding as a top-down, and stochastic resonance as a complementary bottom-up mechanism. Furthermore, we argue that even though contemporary artificial intelligence and machine learning approaches largely lack biological plausibility, the models to be constructed will have to draw on concepts from these fields; since they provide a formal account of the requisite computations that underlie brain function. Nevertheless, biological fidelity will have to be addressed, allowing for testing possible treatment strategies in silico, before application in animal or patient studies. This iteration of computational and empirical studies may help to open the "black boxes" of both machine learning and the human brain.
Mitosis domain generalization in histopathology images -- The MIDOG challenge
Apr 06, 2022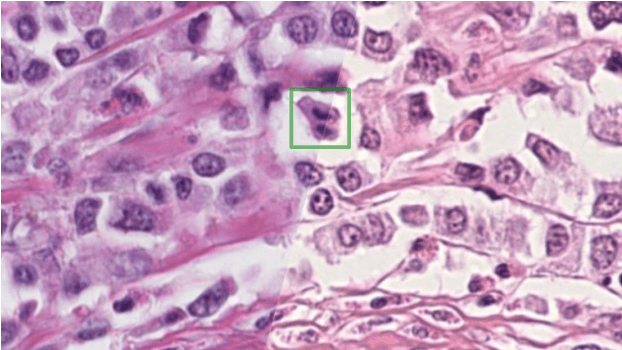
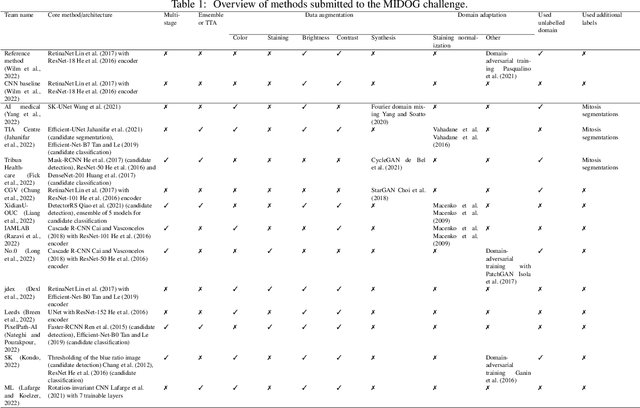
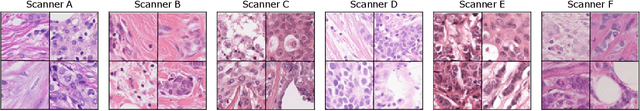
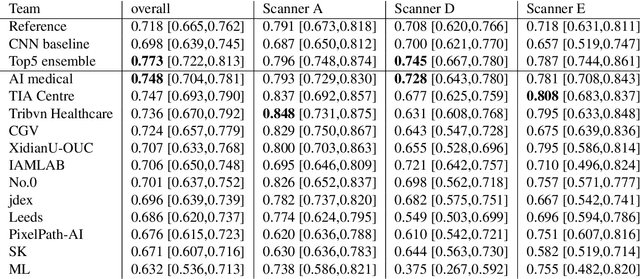
Abstract:The density of mitotic figures within tumor tissue is known to be highly correlated with tumor proliferation and thus is an important marker in tumor grading. Recognition of mitotic figures by pathologists is known to be subject to a strong inter-rater bias, which limits the prognostic value. State-of-the-art deep learning methods can support the expert in this assessment but are known to strongly deteriorate when applied in a different clinical environment than was used for training. One decisive component in the underlying domain shift has been identified as the variability caused by using different whole slide scanners. The goal of the MICCAI MIDOG 2021 challenge has been to propose and evaluate methods that counter this domain shift and derive scanner-agnostic mitosis detection algorithms. The challenge used a training set of 200 cases, split across four scanning systems. As a test set, an additional 100 cases split across four scanning systems, including two previously unseen scanners, were given. The best approaches performed on an expert level, with the winning algorithm yielding an F_1 score of 0.748 (CI95: 0.704-0.781). In this paper, we evaluate and compare the approaches that were submitted to the challenge and identify methodological factors contributing to better performance.
Self-Supervised Speech Representations Preserve Speech Characteristics while Anonymizing Voices
Apr 04, 2022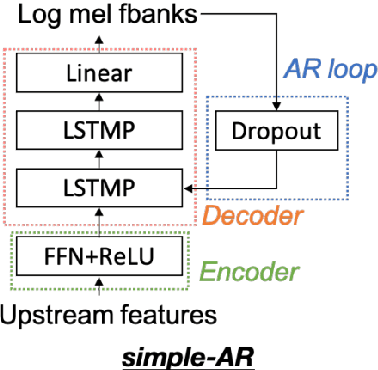
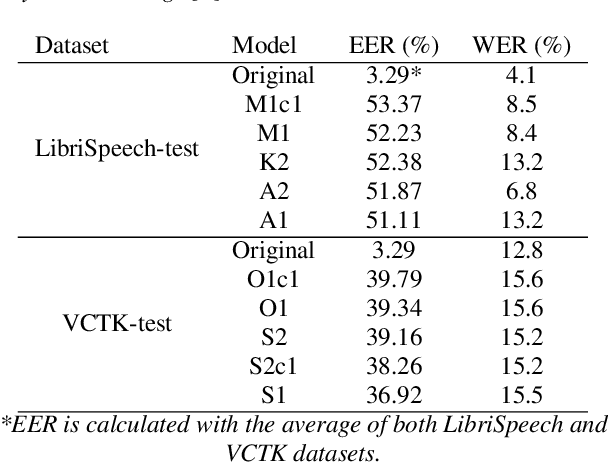

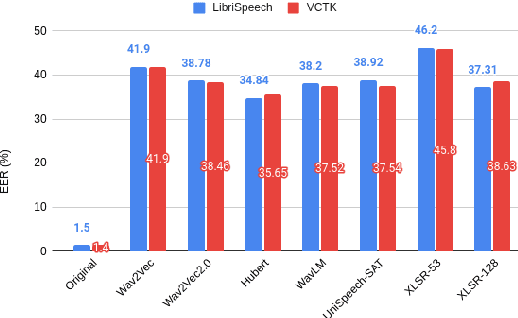
Abstract:Collecting speech data is an important step in training speech recognition systems and other speech-based machine learning models. However, the issue of privacy protection is an increasing concern that must be addressed. The current study investigates the use of voice conversion as a method for anonymizing voices. In particular, we train several voice conversion models using self-supervised speech representations including Wav2Vec2.0, Hubert and UniSpeech. Converted voices retain a low word error rate within 1% of the original voice. Equal error rate increases from 1.52% to 46.24% on the LibriSpeech test set and from 3.75% to 45.84% on speakers from the VCTK corpus which signifies degraded performance on speaker verification. Lastly, we conduct experiments on dysarthric speech data to show that speech features relevant to articulation, prosody, phonation and phonology can be extracted from anonymized voices for discriminating between healthy and pathological speech.
Cross-lingual Self-Supervised Speech Representations for Improved Dysarthric Speech Recognition
Apr 04, 2022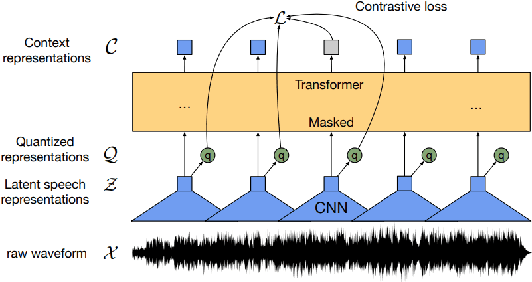

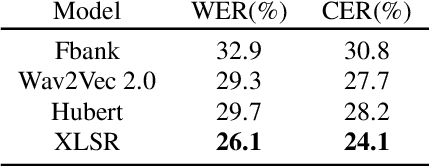
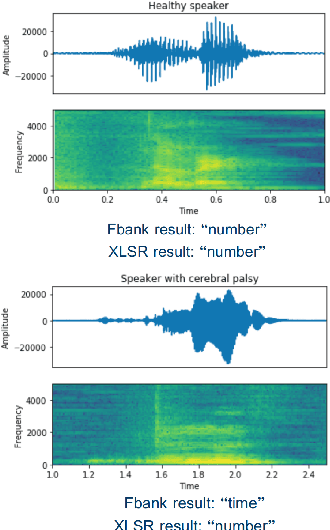
Abstract:State-of-the-art automatic speech recognition (ASR) systems perform well on healthy speech. However, the performance on impaired speech still remains an issue. The current study explores the usefulness of using Wav2Vec self-supervised speech representations as features for training an ASR system for dysarthric speech. Dysarthric speech recognition is particularly difficult as several aspects of speech such as articulation, prosody and phonation can be impaired. Specifically, we train an acoustic model with features extracted from Wav2Vec, Hubert, and the cross-lingual XLSR model. Results suggest that speech representations pretrained on large unlabelled data can improve word error rate (WER) performance. In particular, features from the multilingual model led to lower WERs than filterbanks (Fbank) or models trained on a single language. Improvements were observed in English speakers with cerebral palsy caused dysarthria (UASpeech corpus), Spanish speakers with Parkinsonian dysarthria (PC-GITA corpus) and Italian speakers with paralysis-based dysarthria (EasyCall corpus). Compared to using Fbank features, XLSR-based features reduced WERs by 6.8%, 22.0%, and 7.0% for the UASpeech, PC-GITA, and EasyCall corpus, respectively.
Limited Parameter Denoising for Low-dose X-ray Computed Tomography Using Deep Reinforcement Learning
Apr 01, 2022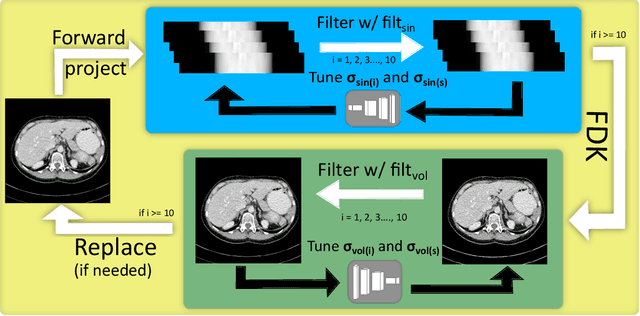
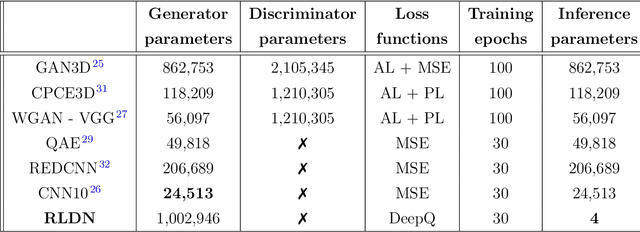
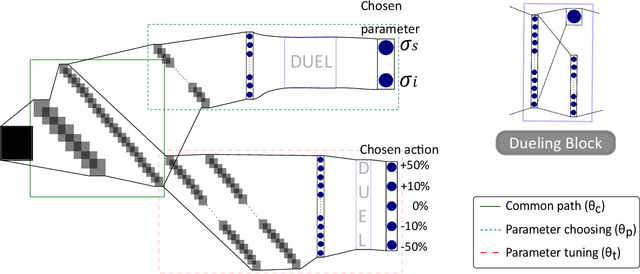
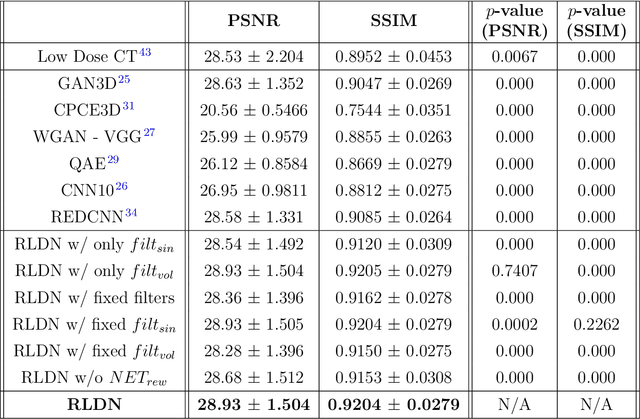
Abstract:The use of deep learning has successfully solved several problems in the field of medical imaging. Deep learning has been applied to the CT denoising problem successfully. However, the use of deep learning requires large amounts of data to train deep convolutional networks (CNNs). Moreover, due to large parameter count, such deep CNNs may cause unexpected results. In this study, we introduce a novel CT denoising framework, which has interpretable behaviour, and provides useful results with limited data. We employ bilateral filtering in both the projection and volume domains to remove noise. To account for non-stationary noise, we tune the $\sigma$ parameters of the volume for every projection view, and for every volume pixel. The tuning is carried out by two deep CNNs. Due to impracticality of labelling, the two deep CNNs are trained via a Deep-Q reinforcement learning task. The reward for the task is generated by using a custom reward function represented by a neural network. Our experiments were carried out on abdominal scans for the Mayo Clinic TCIA dataset, and the AAPM Low Dose CT Grand Challenge. Our denoising framework has excellent denoising performance increasing the PSNR from 28.53 to 28.93, and increasing the SSIM from 0.8952 to 0.9204. We outperform several state-of-the-art deep CNNs, which have several orders of magnitude higher number of parameters (p-value (PSNR) = 0.000, p-value (SSIM) = 0.000). Our method does not introduce any blurring, which is introduced by MSE loss based methods, or any deep learning artifacts, which are introduced by WGAN based models. Our ablation studies show that parameter tuning and using our reward network results in the best possible results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge