Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
A Spatiotemporal Model for Precise and Efficient Fully-automatic 3D Motion Correction in OCT
Sep 15, 2022Abstract:Optical coherence tomography (OCT) is a micrometer-scale, volumetric imaging modality that has become a clinical standard in ophthalmology. OCT instruments image by raster-scanning a focused light spot across the retina, acquiring sequential cross-sectional images to generate volumetric data. Patient eye motion during the acquisition poses unique challenges: Non-rigid, discontinuous distortions can occur, leading to gaps in data and distorted topographic measurements. We present a new distortion model and a corresponding fully-automatic, reference-free optimization strategy for computational motion correction in orthogonally raster-scanned, retinal OCT volumes. Using a novel, domain-specific spatiotemporal parametrization of forward-warping displacements, eye motion can be corrected continuously for the first time. Parameter estimation with temporal regularization improves robustness and accuracy over previous spatial approaches. We correct each A-scan individually in 3D in a single mapping, including repeated acquisitions used in OCT angiography protocols. Specialized 3D forward image warping reduces median runtime to < 9 s, fast enough for clinical use. We present a quantitative evaluation on 18 subjects with ocular pathology and demonstrate accurate correction during microsaccades. Transverse correction is limited only by ocular tremor, whereas submicron repeatability is achieved axially (0.51 um median of medians), representing a dramatic improvement over previous work. This allows assessing longitudinal changes in focal retinal pathologies as a marker of disease progression or treatment response, and promises to enable multiple new capabilities such as supersampled/super-resolution volume reconstruction and analysis of pathological eye motion occuring in neurological diseases.
Deep Learning for automatic head and neck lymph node level delineation
Aug 28, 2022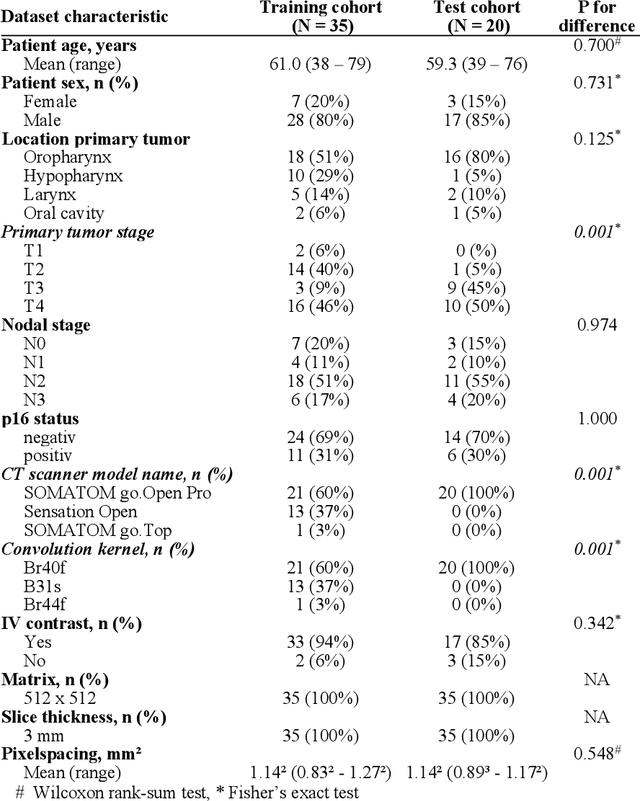
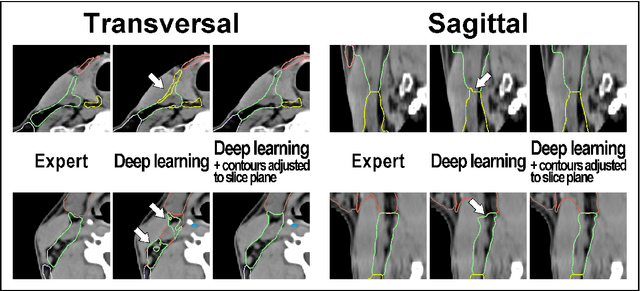
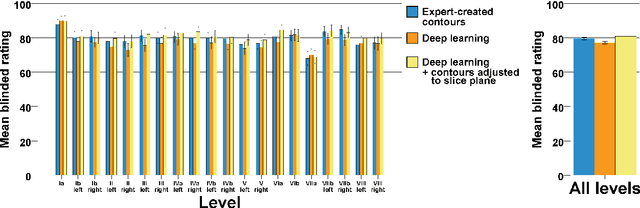
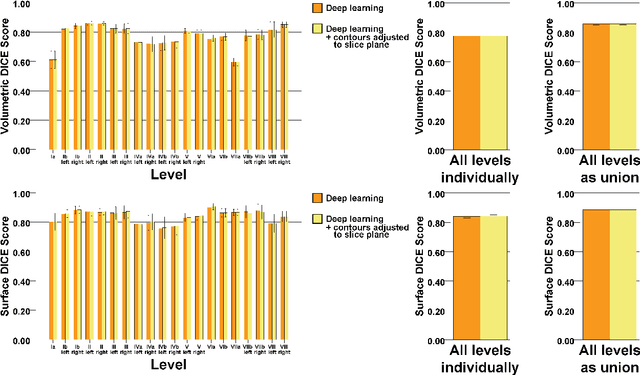
Abstract:Background: Deep learning-based head and neck lymph node level (HN_LNL) autodelineation is of high relevance to radiotherapy research and clinical treatment planning but still understudied in academic literature. Methods: An expert-delineated cohort of 35 planning CTs was used for training of an nnU-net 3D-fullres/2D-ensemble model for autosegmentation of 20 different HN_LNL. Validation was performed in an independent test set (n=20). In a completely blinded evaluation, 3 clinical experts rated the quality of deep learning autosegmentations in a head-to-head comparison with expert-created contours. For a subgroup of 10 cases, intraobserver variability was compared to deep learning autosegmentation performance. The effect of autocontour consistency with CT slice plane orientation on geometric accuracy and expert rating was investigated. Results: Mean blinded expert rating per level was significantly better for deep learning segmentations with CT slice plane adjustment than for expert-created contours (81.0 vs. 79.6, p<0.001), but deep learning segmentations without slice plane adjustment were rated significantly worse than expert-created contours (77.2 vs. 79.6, p<0.001). Geometric accuracy of deep learning segmentations was non-different from intraobserver variability (mean Dice per level, 0.78 vs. 0.77, p=0.064) with variance in accuracy between levels being improved (p<0.001). Clinical significance of contour consistency with CT slice plane orientation was not represented by geometric accuracy metrics (Dice, 0.78 vs. 0.78, p=0.572) Conclusions: We show that a nnU-net 3D-fullres/2D-ensemble model can be used for highly accurate autodelineation of HN_LNL using only a limited training dataset that is ideally suited for large-scale standardized autodelineation of HN_LNL in the research setting. Geometric accuracy metrics are only an imperfect surrogate for blinded expert rating.
A Multi-modal Registration and Visualization Software Tool for Artworks using CraquelureNet
Aug 18, 2022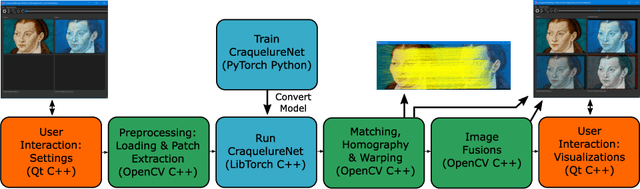
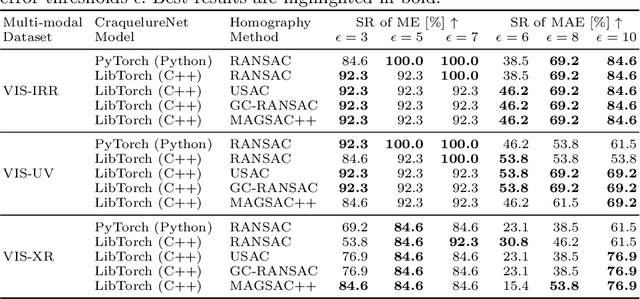
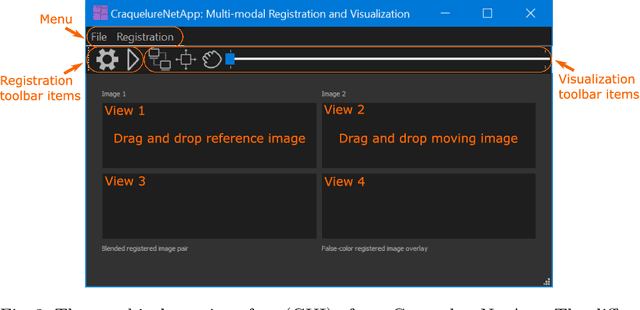
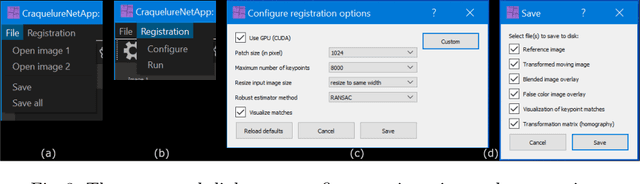
Abstract:For art investigations of paintings, multiple imaging technologies, such as visual light photography, infrared reflectography, ultraviolet fluorescence photography, and x-radiography are often used. For a pixel-wise comparison, the multi-modal images have to be registered. We present a registration and visualization software tool, that embeds a convolutional neural network to extract cross-modal features of the crack structures in historical paintings for automatic registration. The graphical user interface processes the user's input to configure the registration parameters and to interactively adapt the image views with the registered pair and image overlays, such as by individual or synchronized zoom or movements of the views. In the evaluation, we qualitatively and quantitatively show the effectiveness of our software tool in terms of registration performance and short inference time on multi-modal paintings and its transferability by applying our method to historical prints.
SYNTA: A novel approach for deep learning-based image analysis in muscle histopathology using photo-realistic synthetic data
Aug 03, 2022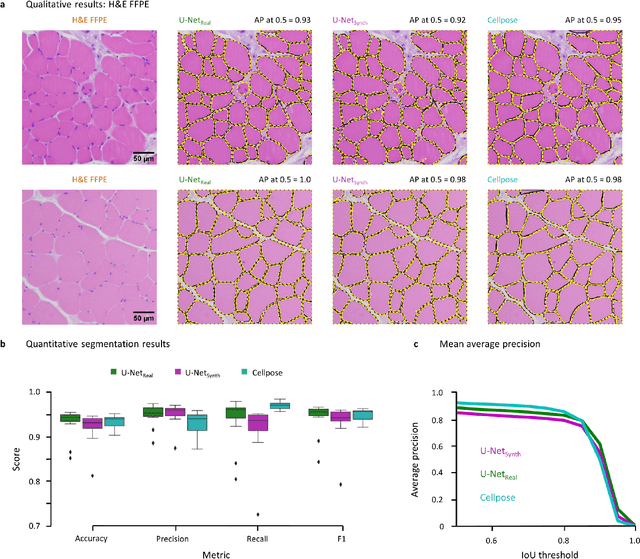
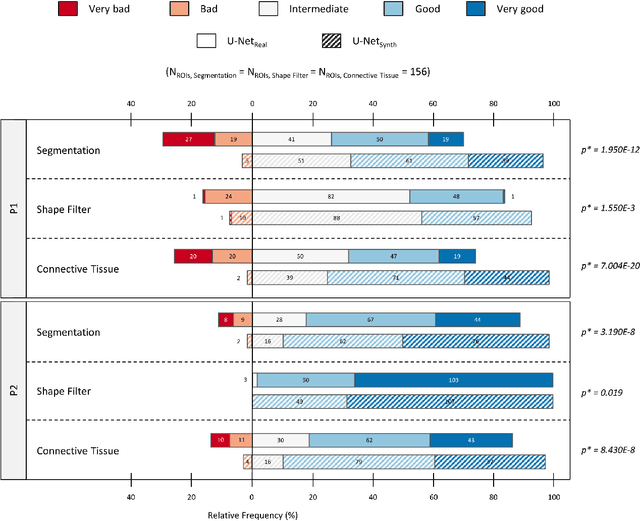
Abstract:Artificial intelligence (AI), machine learning, and deep learning (DL) methods are becoming increasingly important in the field of biomedical image analysis. However, to exploit the full potential of such methods, a representative number of experimentally acquired images containing a significant number of manually annotated objects is needed as training data. Here we introduce SYNTA (synthetic data) as a novel approach for the generation of synthetic, photo-realistic, and highly complex biomedical images as training data for DL systems. We show the versatility of our approach in the context of muscle fiber and connective tissue analysis in histological sections. We demonstrate that it is possible to perform robust and expert-level segmentation tasks on previously unseen real-world data, without the need for manual annotations using synthetic training data alone. Being a fully parametric technique, our approach poses an interpretable and controllable alternative to Generative Adversarial Networks (GANs) and has the potential to significantly accelerate quantitative image analysis in a variety of biomedical applications in microscopy and beyond.
Multi-modal Retinal Image Registration Using a Keypoint-Based Vessel Structure Aligning Network
Jul 21, 2022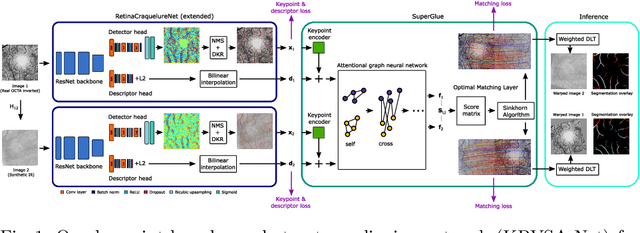
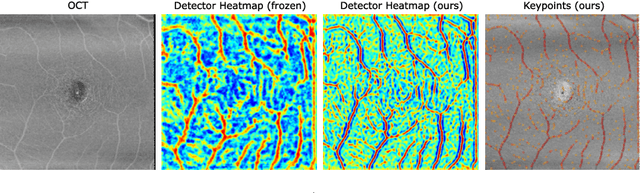
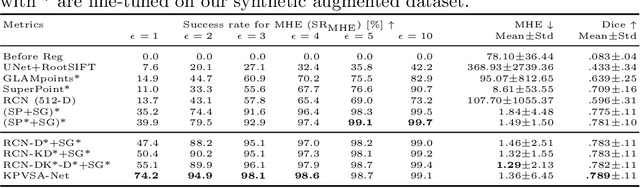

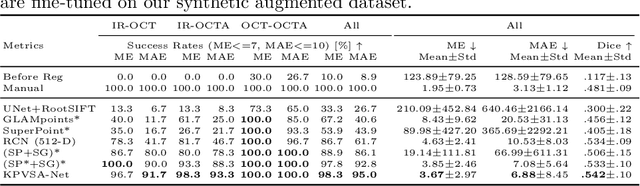
Abstract:In ophthalmological imaging, multiple imaging systems, such as color fundus, infrared, fluorescein angiography, optical coherence tomography (OCT) or OCT angiography, are often involved to make a diagnosis of retinal disease. Multi-modal retinal registration techniques can assist ophthalmologists by providing a pixel-based comparison of aligned vessel structures in images from different modalities or acquisition times. To this end, we propose an end-to-end trainable deep learning method for multi-modal retinal image registration. Our method extracts convolutional features from the vessel structure for keypoint detection and description and uses a graph neural network for feature matching. The keypoint detection and description network and graph neural network are jointly trained in a self-supervised manner using synthetic multi-modal image pairs and are guided by synthetically sampled ground truth homographies. Our method demonstrates higher registration accuracy as competing methods for our synthetic retinal dataset and generalizes well for our real macula dataset and a public fundus dataset.
Trainable Joint Bilateral Filters for Enhanced Prediction Stability in Low-dose CT
Jul 15, 2022
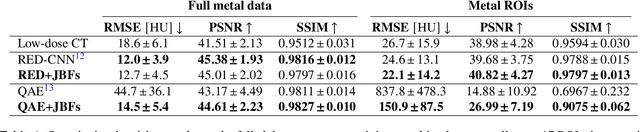
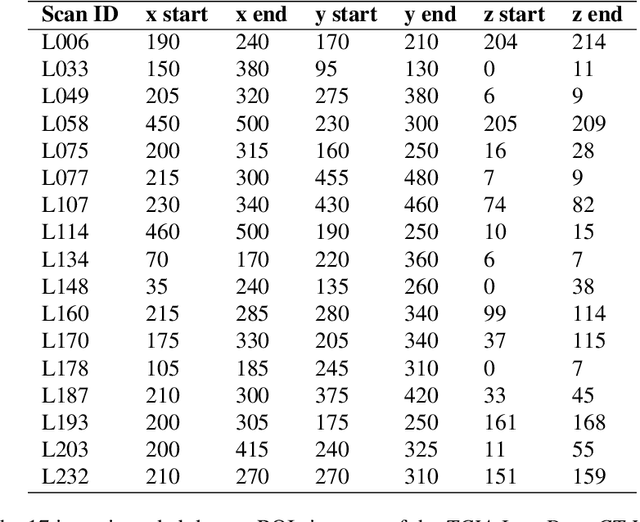
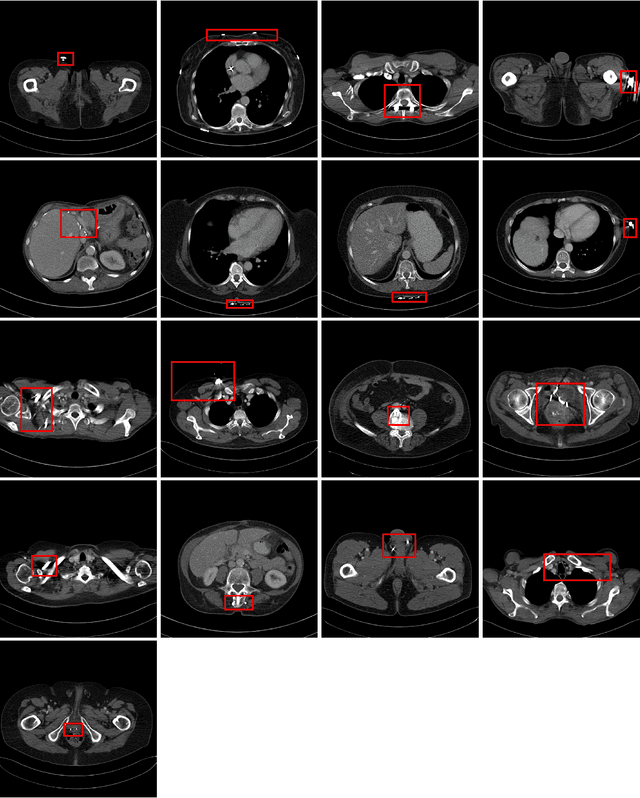
Abstract:Low-dose computed tomography (CT) denoising algorithms aim to enable reduced patient dose in routine CT acquisitions while maintaining high image quality. Recently, deep learning~(DL)-based methods were introduced, outperforming conventional denoising algorithms on this task due to their high model capacity. However, for the transition of DL-based denoising to clinical practice, these data-driven approaches must generalize robustly beyond the seen training data. We, therefore, propose a hybrid denoising approach consisting of a set of trainable joint bilateral filters (JBFs) combined with a convolutional DL-based denoising network to predict the guidance image. Our proposed denoising pipeline combines the high model capacity enabled by DL-based feature extraction with the reliability of the conventional JBF. The pipeline's ability to generalize is demonstrated by training on abdomen CT scans without metal implants and testing on abdomen scans with metal implants as well as on head CT data. When embedding two well-established DL-based denoisers (RED-CNN/QAE) in our pipeline, the denoising performance is improved by $10\,\%$/$82\,\%$ (RMSE) and $3\,\%$/$81\,\%$ (PSNR) in regions containing metal and by $6\,\%$/$78\,\%$ (RMSE) and $2\,\%$/$4\,\%$ (PSNR) on head CT data, compared to the respective vanilla model. Concluding, the proposed trainable JBFs limit the error bound of deep neural networks to facilitate the applicability of DL-based denoisers in low-dose CT pipelines.
AutoSpeed: A Linked Autoencoder Approach for Pulse-Echo Speed-of-Sound Imaging for Medical Ultrasound
Jul 04, 2022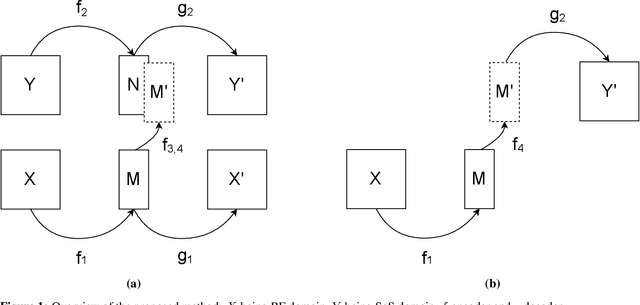
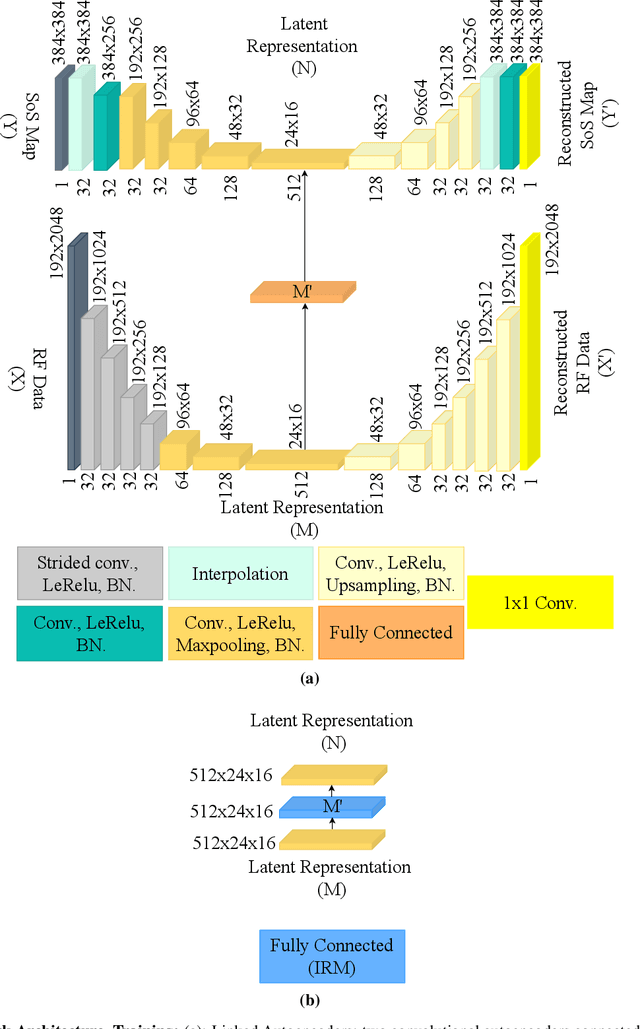
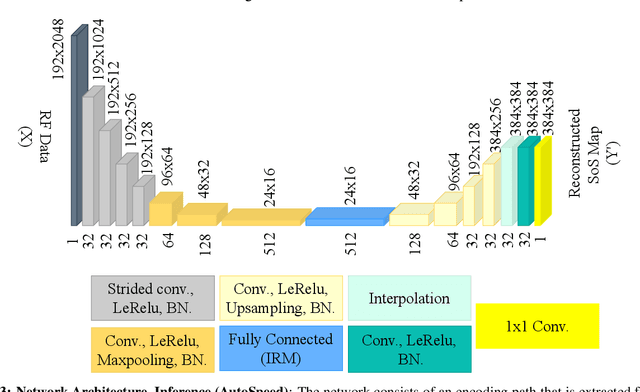
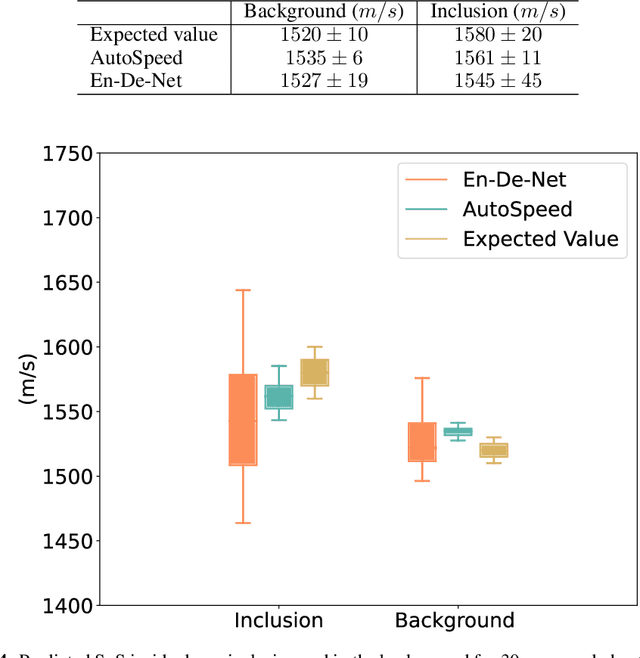
Abstract:Quantitative ultrasound, e.g., speed-of-sound (SoS) in tissues, provides information about tissue properties that have diagnostic value. Recent studies showed the possibility of extracting SoS information from pulse-echo ultrasound raw data (a.k.a. RF data) using deep neural networks that are fully trained on simulated data. These methods take sensor domain data, i.e., RF data, as input and train a network in an end-to-end fashion to learn the implicit mapping between the RF data domain and SoS domain. However, such networks are prone to overfitting to simulated data which results in poor performance and instability when tested on measured data. We propose a novel method for SoS mapping employing learned representations from two linked autoencoders. We test our approach on simulated and measured data acquired from human breast mimicking phantoms. We show that SoS mapping is possible using linked autoencoders. The proposed method has a Mean Absolute Percentage Error (MAPE) of 2.39% on the simulated data. On the measured data, the predictions of the proposed method are close to the expected values with MAPE of 1.1%. Compared to an end-to-end trained network, the proposed method shows higher stability and reproducibility.
PoCaP Corpus: A Multimodal Dataset for Smart Operating Room Speech Assistant using Interventional Radiology Workflow Analysis
Jun 24, 2022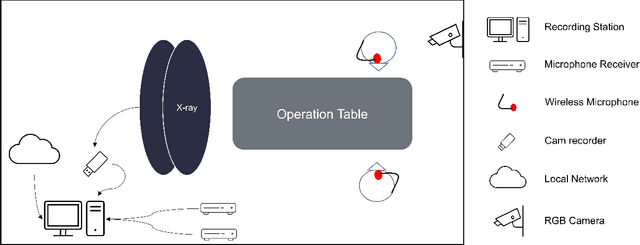
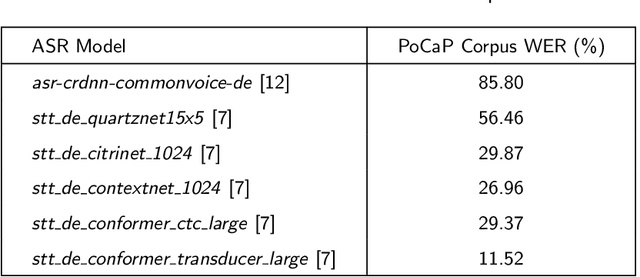
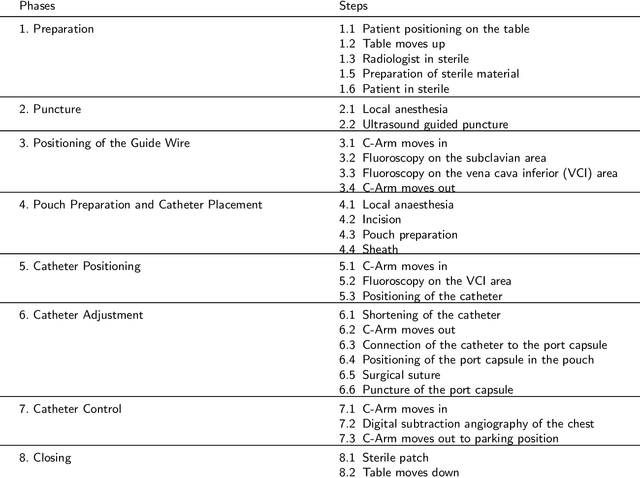
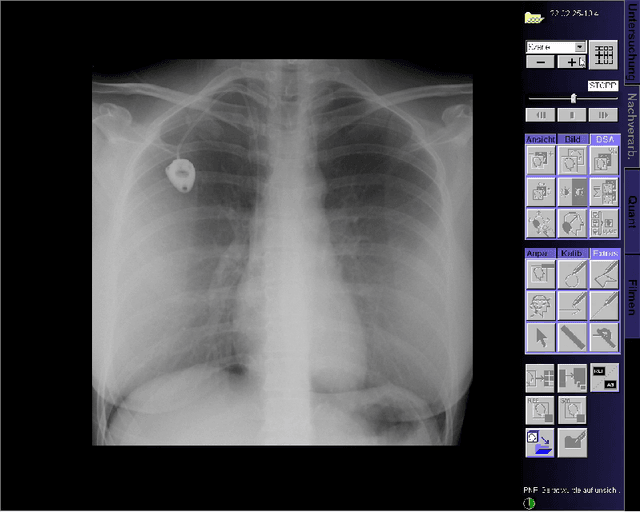

Abstract:This paper presents a new multimodal interventional radiology dataset, called PoCaP (Port Catheter Placement) Corpus. This corpus consists of speech and audio signals in German, X-ray images, and system commands collected from 31 PoCaP interventions by six surgeons with average duration of 81.4 $\pm$ 41.0 minutes. The corpus aims to provide a resource for developing a smart speech assistant in operating rooms. In particular, it may be used to develop a speech controlled system that enables surgeons to control the operation parameters such as C-arm movements and table positions. In order to record the dataset, we acquired consent by the institutional review board and workers council in the University Hospital Erlangen and by the patients for data privacy. We describe the recording set-up, data structure, workflow and preprocessing steps, and report the first PoCaP Corpus speech recognition analysis results with 11.52 $\%$ word error rate using pretrained models. The findings suggest that the data has the potential to build a robust command recognition system and will allow the development of a novel intervention support systems using speech and image processing in the medical domain.
ICC++: Explainable Image Retrieval for Art Historical Corpora using Image Composition Canvas
Jun 22, 2022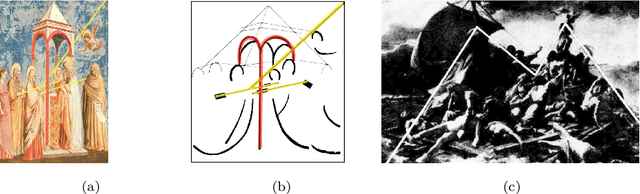
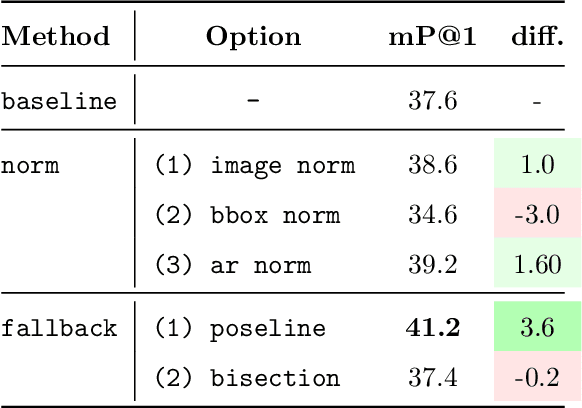
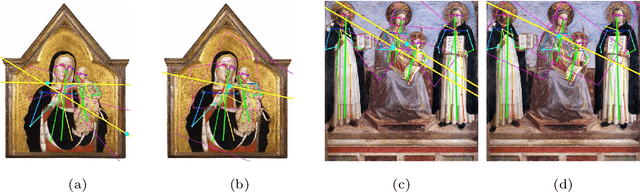
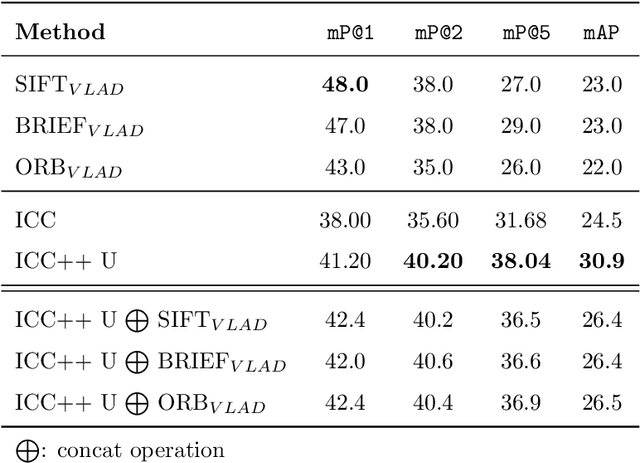
Abstract:Image compositions are helpful in the study of image structures and assist in discovering the semantics of the underlying scene portrayed across art forms and styles. With the digitization of artworks in recent years, thousands of images of a particular scene or narrative could potentially be linked together. However, manually linking this data with consistent objectiveness can be a highly challenging and time-consuming task. In this work, we present a novel approach called Image Composition Canvas (ICC++) to compare and retrieve images having similar compositional elements. ICC++ is an improvement over ICC specializing in generating low and high-level features (compositional elements) motivated by Max Imdahl's work. To this end, we present a rigorous quantitative and qualitative comparison of our approach with traditional and state-of-the-art (SOTA) methods showing that our proposed method outperforms all of them. In combination with deep features, our method outperforms the best deep learning-based method, opening the research direction for explainable machine learning for digital humanities. We will release the code and the data post-publication.
ConFUDA: Contrastive Fewshot Unsupervised Domain Adaptation for Medical Image Segmentation
Jun 08, 2022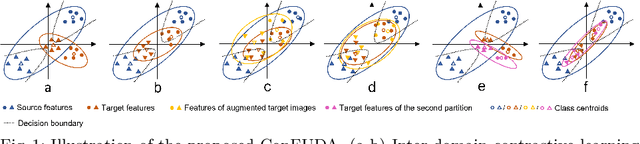
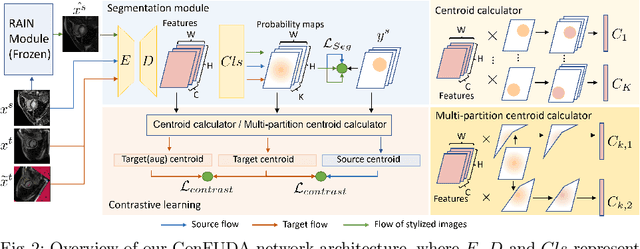
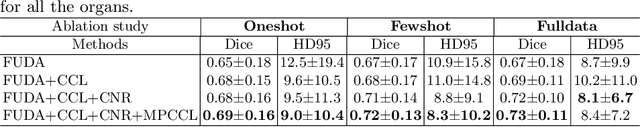
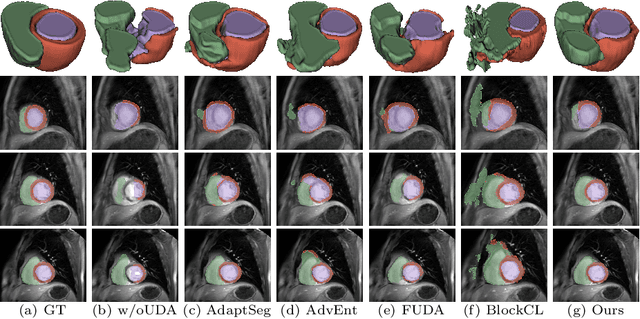
Abstract:Unsupervised domain adaptation (UDA) aims to transfer knowledge learned from a labeled source domain to an unlabeled target domain. Contrastive learning (CL) in the context of UDA can help to better separate classes in feature space. However, in image segmentation, the large memory footprint due to the computation of the pixel-wise contrastive loss makes it prohibitive to use. Furthermore, labeled target data is not easily available in medical imaging, and obtaining new samples is not economical. As a result, in this work, we tackle a more challenging UDA task when there are only a few (fewshot) or a single (oneshot) image available from the target domain. We apply a style transfer module to mitigate the scarcity of target samples. Then, to align the source and target features and tackle the memory issue of the traditional contrastive loss, we propose the centroid-based contrastive learning (CCL) and a centroid norm regularizer (CNR) to optimize the contrastive pairs in both direction and magnitude. In addition, we propose multi-partition centroid contrastive learning (MPCCL) to further reduce the variance in the target features. Fewshot evaluation on MS-CMRSeg dataset demonstrates that ConFUDA improves the segmentation performance by 0.34 of the Dice score on the target domain compared with the baseline, and 0.31 Dice score improvement in a more rigorous oneshot setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge