Stefan Fischer
phys-MCP: A Control Plane for Heterogeneous Physical Neural Networks
May 05, 2026Abstract:Physical neural networks (PNNs) embed computation directly in material dynamics, including molecular, chemical, biological, photonic, memristive, and mechanical substrates. They are attractive for edge computing, especially at the extreme edge, where computation can be placed at the interface to sensing, actuation, or the physical process itself. However, PNNs are difficult to integrate into edge-cloud software stacks because each substrate exposes distinct interfaces, timing behavior, observability limits, and lifecycle requirements. This paper argues that the missing systems component is a common control plane for heterogeneous PNNs. We present phys-MCP, a substrate-aware orchestration architecture that exposes physical neural substrates as discoverable and invocable resources for edge, fog, and cloud workflows, while preserving their possible placement at the extreme edge. phys-MCP defines a capability model, lifecycle semantics, telemetry interfaces, and digital-twin bindings that retain substrate-specific properties such as latency, resetability, plasticity, and I/O modality. We instantiate the architecture through a prototype with three representative backend classes, an HTTP-backed execution path, and an integrated Cortical Labs adapter exposing a wetware-facing API path through the same control model. The evaluation combines controlled experiments on representative backends with end-to-end validation of the Cortical Labs path. Results show descriptor-portable integration across heterogeneous backends, improved runtime-aware matching over simpler baselines, telemetry-aware recovery under representative faults, successful execution against the API-backed wetware path, and small local control-path overhead. Overall, results provide prototype-level evidence that substrate-aware control can span heterogeneous physical AI resources, twin-backed backends, and a wetware-facing API path.
Beyond Silicon: Materials, Mechanisms, and Methods for Physical Neural Computing
Apr 10, 2026Abstract:Physical implementations of neural computation now extend far beyond silicon hardware, encompassing substrates such as memristive devices, photonic circuits, mechanical metamaterials, microfluidic networks, chemical reaction systems, and living neural tissue. By exploiting intrinsic physical processes such as charge transport, wave interference, elastic deformation, mass transport, and biochemical regulation, these substrates can realize neural inference and adaptation directly in matter. As silicon GPU-centered AI faces growing energy and data-movement constraints, physical neural computation is becoming increasingly relevant as a complementary path beyond conventional digital accelerators. This trend is driven in particular by pervasive intelligence, i.e., the deployment of on-device and edge AI across large numbers of resource-constrained systems. In such settings, co-locating computation with sensing and memory can reduce data shuttling and improve efficiency. Meanwhile, physical neural approaches have emerged across disparate disciplines, yet progress remains fragmented, with limited shared terminology and few principled ways to compare platforms. This survey unifies the field by mapping neural primitives to substrate-specific mechanisms, analyzing architectural and training paradigms, and identifying key engineering constraints including scalability, precision, programmability, and I/O interfacing overhead. To enable cross-domain comparison, we introduce a first-order benchmarking scheme based on standardized static and dynamic tasks and physically interpretable performance dimensions. We show that no single substrate dominates across the considered dimensions; instead, physical neural systems occupy complementary operating regimes, enabling applications ranging from ultrafast signal processing and in-memory inference to embodied control and in-sample biochemical decision making.
Deep Learning for automatic head and neck lymph node level delineation
Aug 28, 2022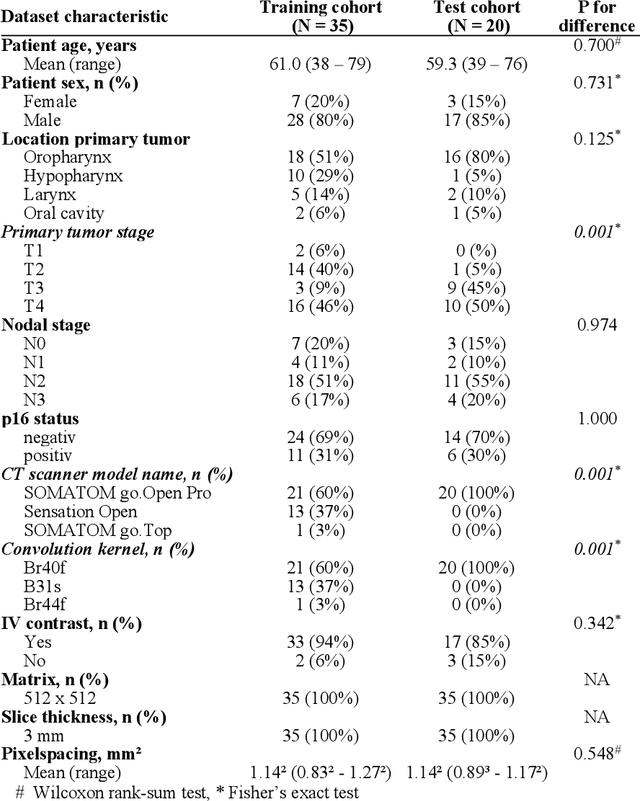
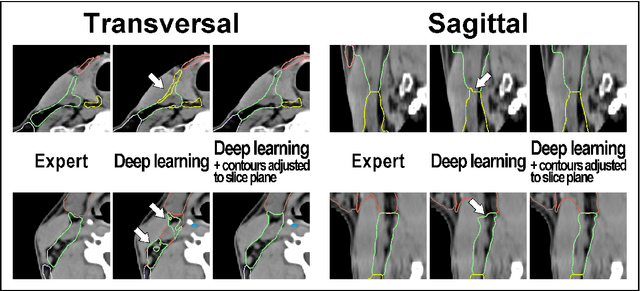
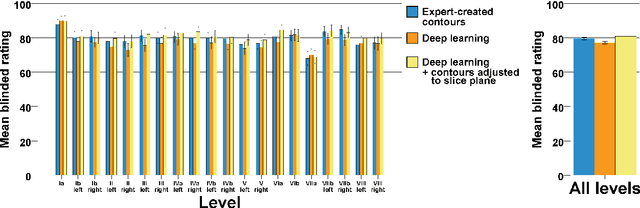
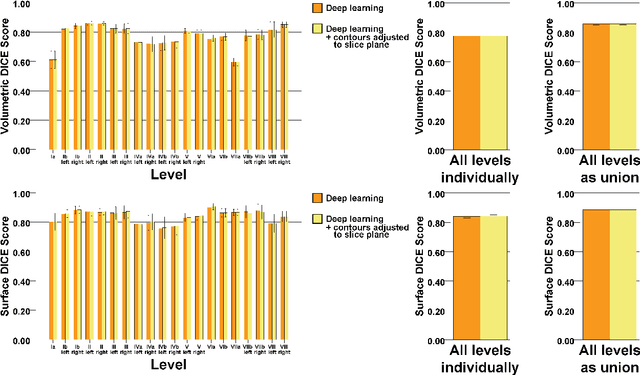
Abstract:Background: Deep learning-based head and neck lymph node level (HN_LNL) autodelineation is of high relevance to radiotherapy research and clinical treatment planning but still understudied in academic literature. Methods: An expert-delineated cohort of 35 planning CTs was used for training of an nnU-net 3D-fullres/2D-ensemble model for autosegmentation of 20 different HN_LNL. Validation was performed in an independent test set (n=20). In a completely blinded evaluation, 3 clinical experts rated the quality of deep learning autosegmentations in a head-to-head comparison with expert-created contours. For a subgroup of 10 cases, intraobserver variability was compared to deep learning autosegmentation performance. The effect of autocontour consistency with CT slice plane orientation on geometric accuracy and expert rating was investigated. Results: Mean blinded expert rating per level was significantly better for deep learning segmentations with CT slice plane adjustment than for expert-created contours (81.0 vs. 79.6, p<0.001), but deep learning segmentations without slice plane adjustment were rated significantly worse than expert-created contours (77.2 vs. 79.6, p<0.001). Geometric accuracy of deep learning segmentations was non-different from intraobserver variability (mean Dice per level, 0.78 vs. 0.77, p=0.064) with variance in accuracy between levels being improved (p<0.001). Clinical significance of contour consistency with CT slice plane orientation was not represented by geometric accuracy metrics (Dice, 0.78 vs. 0.78, p=0.572) Conclusions: We show that a nnU-net 3D-fullres/2D-ensemble model can be used for highly accurate autodelineation of HN_LNL using only a limited training dataset that is ideally suited for large-scale standardized autodelineation of HN_LNL in the research setting. Geometric accuracy metrics are only an imperfect surrogate for blinded expert rating.
Continual Learning for Peer-to-Peer Federated Learning: A Study on Automated Brain Metastasis Identification
Apr 30, 2022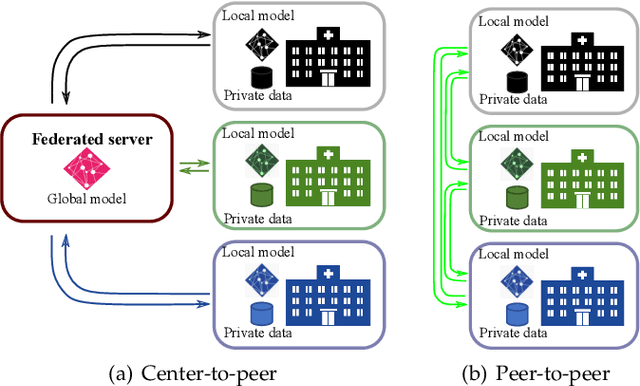
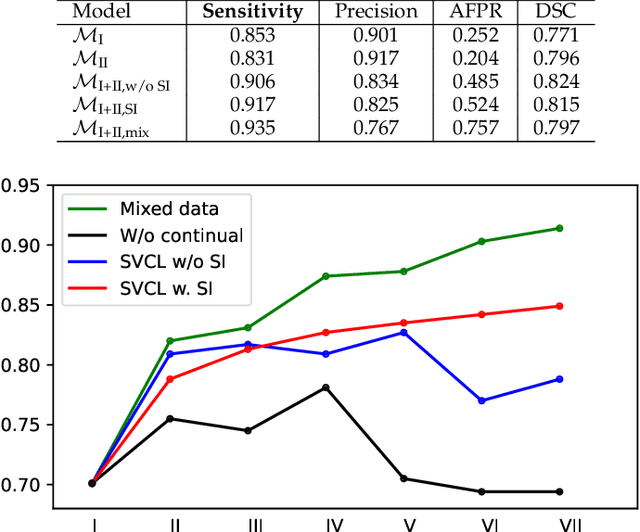
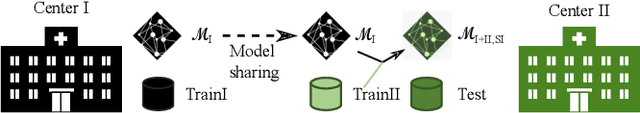
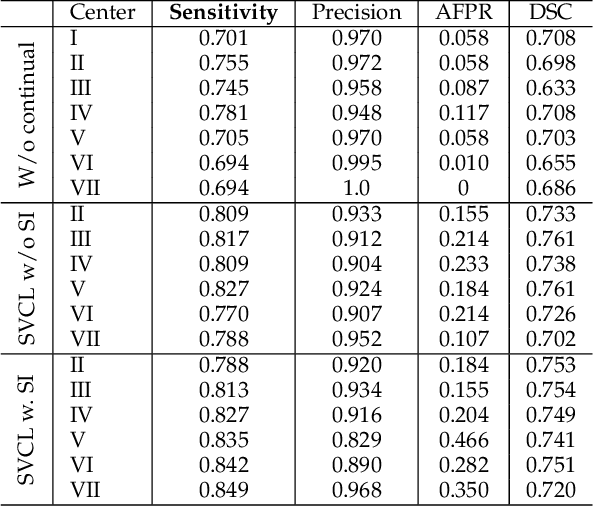
Abstract:Due to data privacy constraints, data sharing among multiple centers is restricted. Continual learning, as one approach to peer-to-peer federated learning, can promote multicenter collaboration on deep learning algorithm development by sharing intermediate models instead of training data. This work aims to investigate the feasibility of continual learning for multicenter collaboration on an exemplary application of brain metastasis identification using DeepMedic. 920 T1 MRI contrast enhanced volumes are split to simulate multicenter collaboration scenarios. A continual learning algorithm, synaptic intelligence (SI), is applied to preserve important model weights for training one center after another. In a bilateral collaboration scenario, continual learning with SI achieves a sensitivity of 0.917, and naive continual learning without SI achieves a sensitivity of 0.906, while two models trained on internal data solely without continual learning achieve sensitivity of 0.853 and 0.831 only. In a seven-center multilateral collaboration scenario, the models trained on internal datasets (100 volumes each center) without continual learning obtain a mean sensitivity value of 0.699. With single-visit continual learning (i.e., the shared model visits each center only once during training), the sensitivity is improved to 0.788 and 0.849 without SI and with SI, respectively. With iterative continual learning (i.e., the shared model revisits each center multiple times during training), the sensitivity is further improved to 0.914, which is identical to the sensitivity using mixed data for training. Our experiments demonstrate that continual learning can improve brain metastasis identification performance for centers with limited data. This study demonstrates the feasibility of applying continual learning for peer-to-peer federated learning in multicenter collaboration.
Lifting DecPOMDPs for Nanoscale Systems -- A Work in Progress
Oct 18, 2021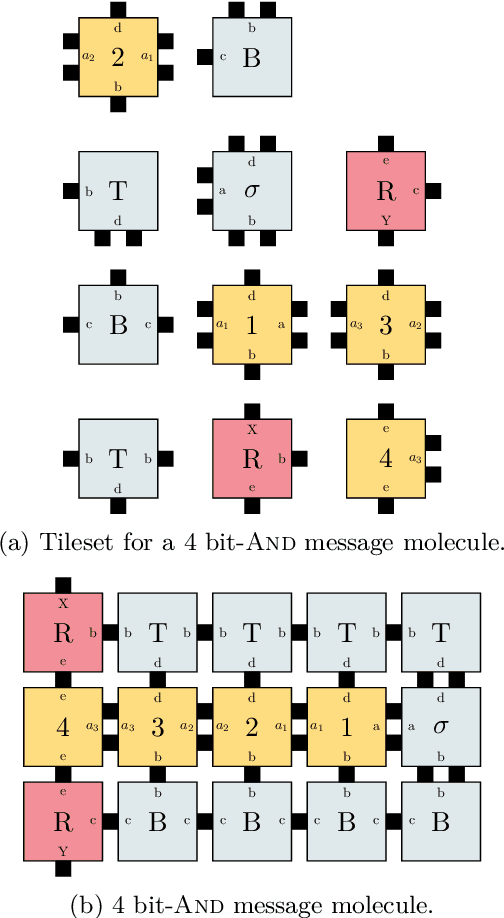
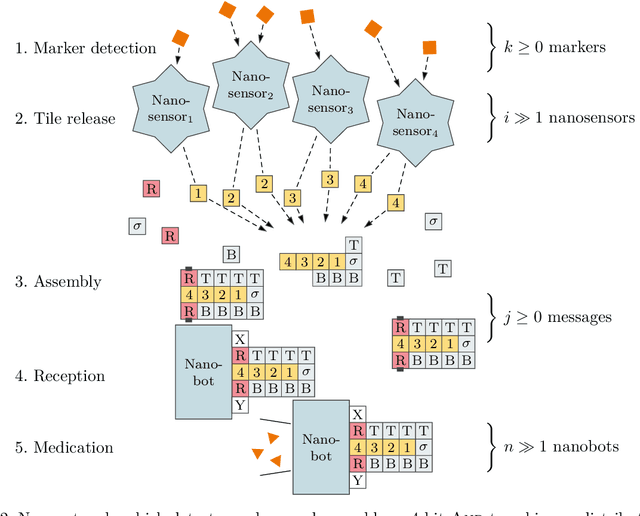
Abstract:DNA-based nanonetworks have a wide range of promising use cases, especially in the field of medicine. With a large set of agents, a partially observable stochastic environment, and noisy observations, such nanoscale systems can be modelled as a decentralised, partially observable, Markov decision process (DecPOMDP). As the agent set is a dominating factor, this paper presents (i) lifted DecPOMDPs, partitioning the agent set into sets of indistinguishable agents, reducing the worst-case space required, and (ii) a nanoscale medical system as an application. Future work turns to solving and implementing lifted DecPOMDPs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge