Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
Improved HER2 Tumor Segmentation with Subtype Balancing using Deep Generative Networks
Nov 11, 2022



Abstract:Tumor segmentation in histopathology images is often complicated by its composition of different histological subtypes and class imbalance. Oversampling subtypes with low prevalence features is not a satisfactory solution since it eventually leads to overfitting. We propose to create synthetic images with semantically-conditioned deep generative networks and to combine subtype-balanced synthetic images with the original dataset to achieve better segmentation performance. We show the suitability of Generative Adversarial Networks (GANs) and especially diffusion models to create realistic images based on subtype-conditioning for the use case of HER2-stained histopathology. Additionally, we show the capability of diffusion models to conditionally inpaint HER2 tumor areas with modified subtypes. Combining the original dataset with the same amount of diffusion-generated images increased the tumor Dice score from 0.833 to 0.854 and almost halved the variance between the HER2 subtype recalls. These results create the basis for more reliable automatic HER2 analysis with lower performance variance between individual HER2 subtypes.
Employing Graph Representations for Cell-level Characterization of Melanoma MELC Samples
Nov 10, 2022


Abstract:Histopathology imaging is crucial for the diagnosis and treatment of skin diseases. For this reason, computer-assisted approaches have gained popularity and shown promising results in tasks such as segmentation and classification of skin disorders. However, collecting essential data and sufficiently high-quality annotations is a challenge. This work describes a pipeline that uses suspected melanoma samples that have been characterized using Multi-Epitope-Ligand Cartography (MELC). This cellular-level tissue characterisation is then represented as a graph and used to train a graph neural network. This imaging technology, combined with the methodology proposed in this work, achieves a classification accuracy of 87%, outperforming existing approaches by 10%.
Generation of Anonymous Chest Radiographs Using Latent Diffusion Models for Training Thoracic Abnormality Classification Systems
Nov 04, 2022Abstract:The availability of large-scale chest X-ray datasets is a requirement for developing well-performing deep learning-based algorithms in thoracic abnormality detection and classification. However, biometric identifiers in chest radiographs hinder the public sharing of such data for research purposes due to the risk of patient re-identification. To counteract this issue, synthetic data generation offers a solution for anonymizing medical images. This work employs a latent diffusion model to synthesize an anonymous chest X-ray dataset of high-quality class-conditional images. We propose a privacy-enhancing sampling strategy to ensure the non-transference of biometric information during the image generation process. The quality of the generated images and the feasibility of serving as exclusive training data are evaluated on a thoracic abnormality classification task. Compared to a real classifier, we achieve competitive results with a performance gap of only 3.5% in the area under the receiver operating characteristic curve.
On the Benefit of Dual-domain Denoising in a Self-supervised Low-dose CT Setting
Nov 03, 2022



Abstract:Computed tomography (CT) is routinely used for three-dimensional non-invasive imaging. Numerous data-driven image denoising algorithms were proposed to restore image quality in low-dose acquisitions. However, considerably less research investigates methods already intervening in the raw detector data due to limited access to suitable projection data or correct reconstruction algorithms. In this work, we present an end-to-end trainable CT reconstruction pipeline that contains denoising operators in both the projection and the image domain and that are optimized simultaneously without requiring ground-truth high-dose CT data. Our experiments demonstrate that including an additional projection denoising operator improved the overall denoising performance by 82.4-94.1%/12.5-41.7% (PSNR/SSIM) on abdomen CT and 1.5-2.9%/0.4-0.5% (PSNR/SSIM) on XRM data relative to the low-dose baseline. We make our entire helical CT reconstruction framework publicly available that contains a raw projection rebinning step to render helical projection data suitable for differentiable fan-beam reconstruction operators and end-to-end learning.
Neural Network based Formation of Cognitive Maps of Semantic Spaces and the Emergence of Abstract Concepts
Oct 28, 2022Abstract:The hippocampal-entorhinal complex plays a major role in the organization of memory and thought. The formation of and navigation in cognitive maps of arbitrary mental spaces via place and grid cells can serve as a representation of memories and experiences and their relations to each other. The multi-scale successor representation is proposed to be the mathematical principle underlying place and grid cell computations. Here, we present a neural network, which learns a cognitive map of a semantic space based on 32 different animal species encoded as feature vectors. The neural network successfully learns the similarities between different animal species, and constructs a cognitive map of 'animal space' based on the principle of successor representations with an accuracy of around 30% which is near to the theoretical maximum regarding the fact that all animal species have more than one possible successor, i.e. nearest neighbor in feature space. Furthermore, a hierarchical structure, i.e. different scales of cognitive maps, can be modeled based on multi-scale successor representations. We find that, in fine-grained cognitive maps, the animal vectors are evenly distributed in feature space. In contrast, in coarse-grained maps, animal vectors are highly clustered according to their biological class, i.e. amphibians, mammals and insects. This could be a possible mechanism explaining the emergence of new abstract semantic concepts. Finally, even completely new or incomplete input can be represented by interpolation of the representations from the cognitive map with remarkable high accuracy of up to 95%. We conclude that the successor representation can serve as a weighted pointer to past memories and experiences, and may therefore be a crucial building block for future machine learning to include prior knowledge, and to derive context knowledge from novel input.
Heat Demand Forecasting with Multi-Resolutional Representation of Heterogeneous Temporal Ensemble
Oct 24, 2022Abstract:One of the primal challenges faced by utility companies is ensuring efficient supply with minimal greenhouse gas emissions. The advent of smart meters and smart grids provide an unprecedented advantage in realizing an optimised supply of thermal energies through proactive techniques such as load forecasting. In this paper, we propose a forecasting framework for heat demand based on neural networks where the time series are encoded as scalograms equipped with the capacity of embedding exogenous variables such as weather, and holiday/non-holiday. Subsequently, CNNs are utilized to predict the heat load multi-step ahead. Finally, the proposed framework is compared with other state-of-the-art methods, such as SARIMAX and LSTM. The quantitative results from retrospective experiments show that the proposed framework consistently outperforms the state-of-the-art baseline method with real-world data acquired from Denmark. A minimal mean error of 7.54% for MAPE and 417kW for RMSE is achieved with the proposed framework in comparison to all other methods.
ArtFacePoints: High-resolution Facial Landmark Detection in Paintings and Prints
Oct 17, 2022
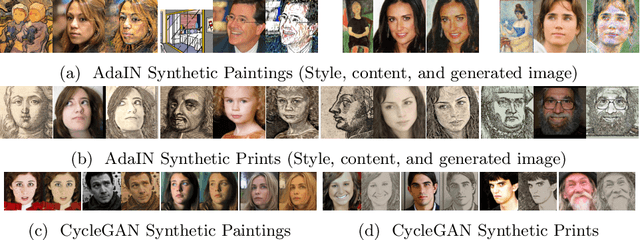
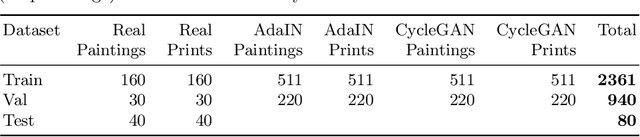

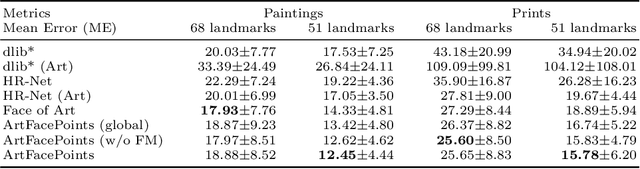
Abstract:Facial landmark detection plays an important role for the similarity analysis in artworks to compare portraits of the same or similar artists. With facial landmarks, portraits of different genres, such as paintings and prints, can be automatically aligned using control-point-based image registration. We propose a deep-learning-based method for facial landmark detection in high-resolution images of paintings and prints. It divides the task into a global network for coarse landmark prediction and multiple region networks for precise landmark refinement in regions of the eyes, nose, and mouth that are automatically determined based on the predicted global landmark coordinates. We created a synthetically augmented facial landmark art dataset including artistic style transfer and geometric landmark shifts. Our method demonstrates an accurate detection of the inner facial landmarks for our high-resolution dataset of artworks while being comparable for a public low-resolution artwork dataset in comparison to competing methods.
Self-Supervised 2D/3D Registration for X-Ray to CT Image Fusion
Oct 14, 2022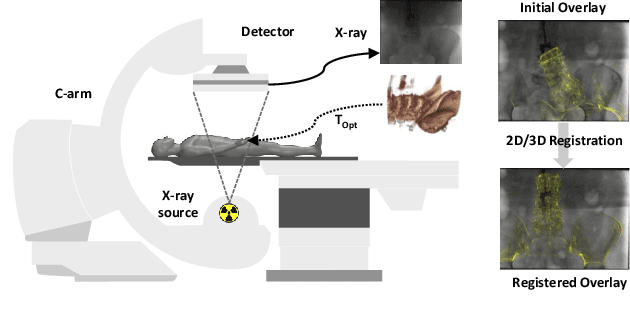
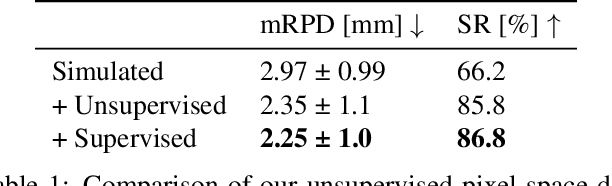
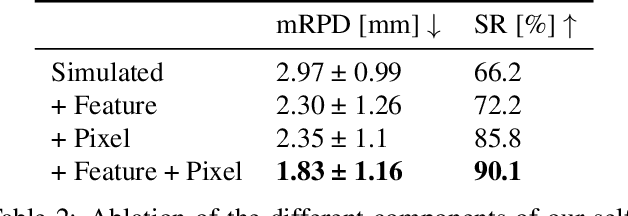

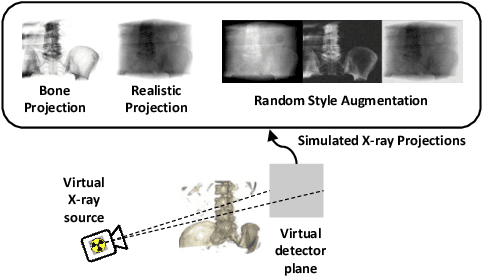

Abstract:Deep Learning-based 2D/3D registration enables fast, robust, and accurate X-ray to CT image fusion when large annotated paired datasets are available for training. However, the need for paired CT volume and X-ray images with ground truth registration limits the applicability in interventional scenarios. An alternative is to use simulated X-ray projections from CT volumes, thus removing the need for paired annotated datasets. Deep Neural Networks trained exclusively on simulated X-ray projections can perform significantly worse on real X-ray images due to the domain gap. We propose a self-supervised 2D/3D registration framework combining simulated training with unsupervised feature and pixel space domain adaptation to overcome the domain gap and eliminate the need for paired annotated datasets. Our framework achieves a registration accuracy of 1.83$\pm$1.16 mm with a high success ratio of 90.1% on real X-ray images showing a 23.9% increase in success ratio compared to reference annotation-free algorithms.
Deep Learning-based Anonymization of Chest Radiographs: A Utility-preserving Measure for Patient Privacy
Sep 23, 2022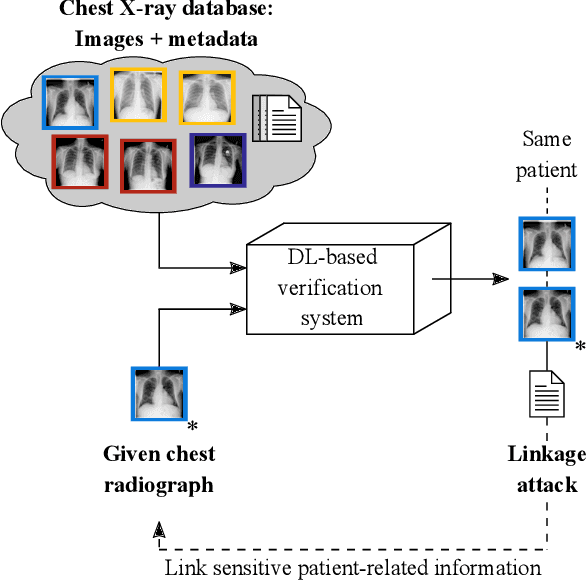
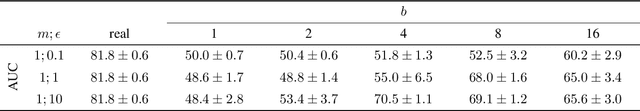
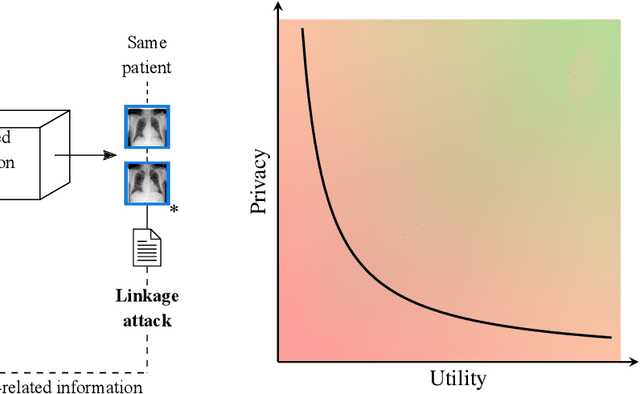
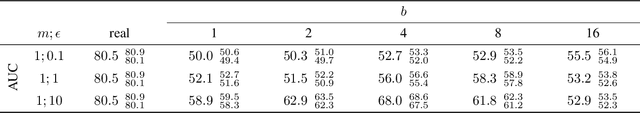
Abstract:Robust and reliable anonymization of chest radiographs constitutes an essential step before publishing large datasets of such for research purposes. The conventional anonymization process is carried out by obscuring personal information in the images with black boxes and removing or replacing meta-information. However, such simple measures retain biometric information in the chest radiographs, allowing patients to be re-identified by a linkage attack. Therefore, we see an urgent need to obfuscate the biometric information appearing in the images. To the best of our knowledge, we propose the first deep learning-based approach to targetedly anonymize chest radiographs while maintaining data utility for diagnostic and machine learning purposes. Our model architecture is a composition of three independent neural networks that, when collectively used, allow for learning a deformation field that is able to impede patient re-identification. The individual influence of each component is investigated with an ablation study. Quantitative results on the ChestX-ray14 dataset show a reduction of patient re-identification from 81.8% to 58.6% in the area under the receiver operating characteristic curve (AUC) with little impact on the abnormality classification performance. This indicates the ability to preserve underlying abnormality patterns while increasing patient privacy. Furthermore, we compare the proposed deep learning-based anonymization approach with differentially private image pixelization, and demonstrate the superiority of our method towards resolving the privacy-utility trade-off for chest radiographs.
Metal Inpainting in CBCT Projections Using Score-based Generative Model
Sep 20, 2022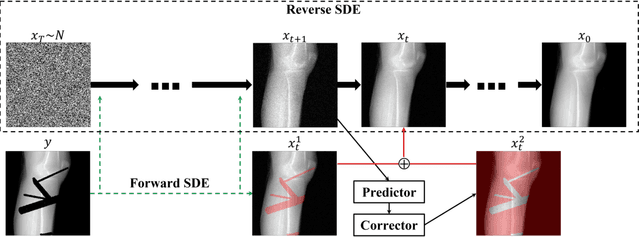
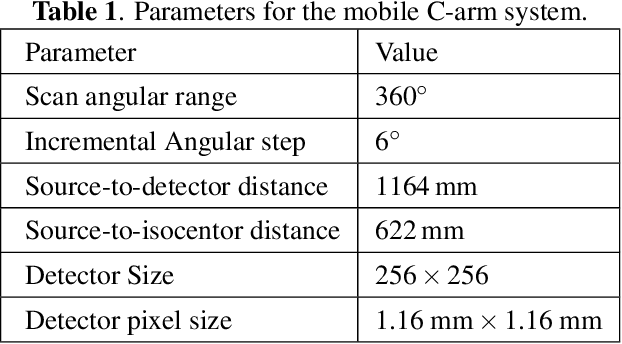
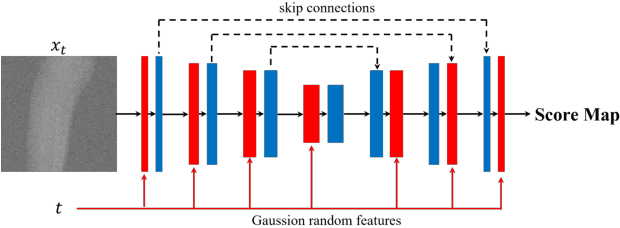
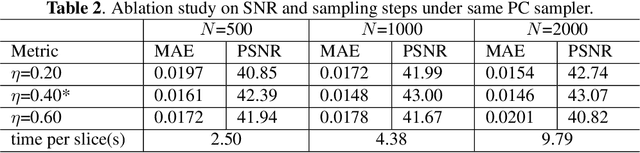
Abstract:During orthopaedic surgery, the inserting of metallic implants or screws are often performed under mobile C-arm systems. Due to the high attenuation of metals, severe metal artifacts occur in 3D reconstructions, which degrade the image quality greatly. To reduce the artifacts, many metal artifact reduction algorithms have been developed and metal inpainting in projection domain is an essential step. In this work, a score-based generative model is trained on simulated knee projections and the inpainted image is obtained by removing the noise in conditional resampling process. The result implies that the inpainted images by score-based generative model have more detailed information and achieve the lowest mean absolute error and the highest peak-signal-to-noise-ratio compared with interpolation and CNN based method. Besides, the score-based model can also recover projections with big circlar and rectangular masks, showing its generalization in inpainting task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge