Alexander Becker
Making Foundation Models Probabilistic via Singular Value Ensembles
Jan 29, 2026Abstract:Foundation models have become a dominant paradigm in machine learning, achieving remarkable performance across diverse tasks through large-scale pretraining. However, these models often yield overconfident, uncalibrated predictions. The standard approach to quantifying epistemic uncertainty, training an ensemble of independent models, incurs prohibitive computational costs that scale linearly with ensemble size, making it impractical for large foundation models. We propose Singular Value Ensemble (SVE), a parameter-efficient implicit ensemble method that builds on a simple, but powerful core assumption: namely, that the singular vectors of the weight matrices constitute meaningful subspaces of the model's knowledge. Pretrained foundation models encode rich, transferable information in their weight matrices. If the singular vectors are indeed meaningful (orthogonal) "knowledge directions". To obtain a model ensemble, we modulate only how strongly each direction contributes to the output. Rather than learning entirely new parameters, we freeze the singular vectors and only train per-member singular values that rescale the contribution of each direction in that shared knowledge basis. Ensemble diversity emerges naturally as stochastic initialization and random sampling of mini-batches during joint training cause different members to converge to different combinations of the same underlying knowledge. SVE achieves uncertainty quantification comparable to explicit deep ensembles while increasing the parameter count of the base model by less than 1%, making principled uncertainty estimation accessible in resource-constrained settings. We validate SVE on NLP and vision tasks with various different backbones and show that it improves calibration while maintaining predictive accuracy.
Marigold-DC: Zero-Shot Monocular Depth Completion with Guided Diffusion
Dec 18, 2024



Abstract:Depth completion upgrades sparse depth measurements into dense depth maps guided by a conventional image. Existing methods for this highly ill-posed task operate in tightly constrained settings and tend to struggle when applied to images outside the training domain or when the available depth measurements are sparse, irregularly distributed, or of varying density. Inspired by recent advances in monocular depth estimation, we reframe depth completion as an image-conditional depth map generation guided by sparse measurements. Our method, Marigold-DC, builds on a pretrained latent diffusion model for monocular depth estimation and injects the depth observations as test-time guidance via an optimization scheme that runs in tandem with the iterative inference of denoising diffusion. The method exhibits excellent zero-shot generalization across a diverse range of environments and handles even extremely sparse guidance effectively. Our results suggest that contemporary monocular depth priors greatly robustify depth completion: it may be better to view the task as recovering dense depth from (dense) image pixels, guided by sparse depth; rather than as inpainting (sparse) depth, guided by an image. Project website: https://MarigoldDepthCompletion.github.io/
The unrealized potential of agroforestry for an emissions-intensive agricultural commodity
Oct 28, 2024


Abstract:Reconciling agricultural production with climate-change mitigation and adaptation is one of the most formidable problems in sustainability. One proposed strategy for addressing this problem is the judicious retention of trees in agricultural systems. However, the magnitude of the current and future-potential benefit that trees contribute remains uncertain, particularly in the agricultural sector where trees can also limit production. Here we help to resolve these issues across a West African region responsible for producing $\approx$60% of the world's cocoa, a crop that contributes one of the highest per unit carbon footprints of all foods. We use machine learning to generate spatially-explicit estimates of shade-tree cover and carbon stocks across the region. We find that existing shade-tree cover is low, and not spatially aligned with climate threat. But we also find enormous unrealized potential for the sector to counterbalance a large proportion of their high carbon footprint annually, without threatening production. Our methods can be applied to other globally significant commodities that can be grown in agroforests, and align with accounting requirements of carbon markets, and emerging legislative requirements for sustainability reporting.
LoRA-Ensemble: Efficient Uncertainty Modelling for Self-attention Networks
May 23, 2024Abstract:Numerous crucial tasks in real-world decision-making rely on machine learning algorithms with calibrated uncertainty estimates. However, modern methods often yield overconfident and uncalibrated predictions. Various approaches involve training an ensemble of separate models to quantify the uncertainty related to the model itself, known as epistemic uncertainty. In an explicit implementation, the ensemble approach has high computational cost and high memory requirements. This particular challenge is evident in state-of-the-art neural networks such as transformers, where even a single network is already demanding in terms of compute and memory. Consequently, efforts are made to emulate the ensemble model without actually instantiating separate ensemble members, referred to as implicit ensembling. We introduce LoRA-Ensemble, a parameter-efficient deep ensemble method for self-attention networks, which is based on Low-Rank Adaptation (LoRA). Initially developed for efficient LLM fine-tuning, we extend LoRA to an implicit ensembling approach. By employing a single pre-trained self-attention network with weights shared across all members, we train member-specific low-rank matrices for the attention projections. Our method exhibits superior calibration compared to explicit ensembles and achieves similar or better accuracy across various prediction tasks and datasets.
Neural Fields with Thermal Activations for Arbitrary-Scale Super-Resolution
Nov 29, 2023Abstract:Recent approaches for arbitrary-scale single image super-resolution (ASSR) have used local neural fields to represent continuous signals that can be sampled at different rates. However, in such formulation, the point-wise query of field values does not naturally match the point spread function (PSF) of a given pixel. In this work we present a novel way to design neural fields such that points can be queried with a Gaussian PSF, which serves as anti-aliasing when moving across resolutions for ASSR. We achieve this using a novel activation function derived from Fourier theory and the heat equation. This comes at no additional cost: querying a point with a Gaussian PSF in our framework does not affect computational cost, unlike filtering in the image domain. Coupled with a hypernetwork, our method not only provides theoretically guaranteed anti-aliasing, but also sets a new bar for ASSR while also being more parameter-efficient than previous methods.
Certified Data Removal in Sum-Product Networks
Oct 04, 2022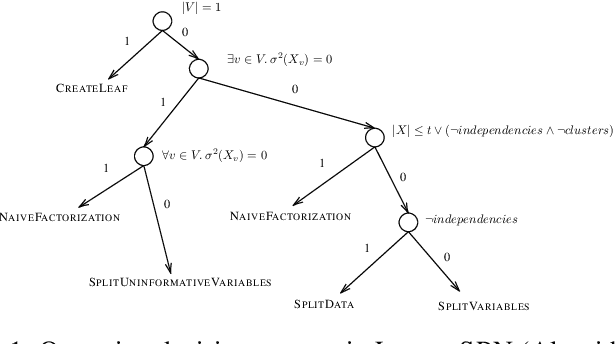
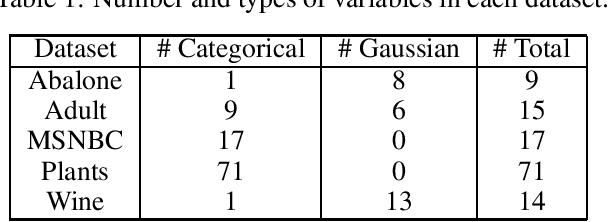
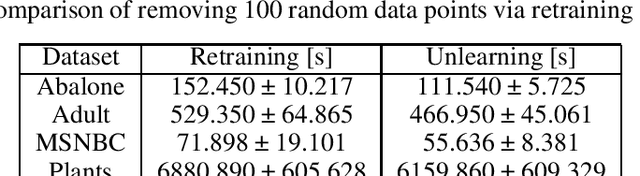
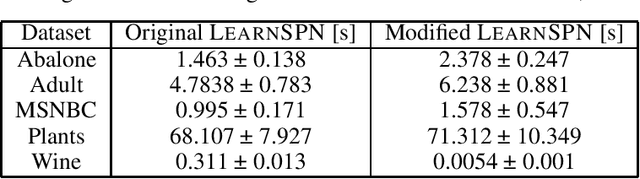
Abstract:Data protection regulations like the GDPR or the California Consumer Privacy Act give users more control over the data that is collected about them. Deleting the collected data is often insufficient to guarantee data privacy since it is often used to train machine learning models, which can expose information about the training data. Thus, a guarantee that a trained model does not expose information about its training data is additionally needed. In this paper, we present UnlearnSPN -- an algorithm that removes the influence of single data points from a trained sum-product network and thereby allows fulfilling data privacy requirements on demand.
Evaluating Machine Unlearning via Epistemic Uncertainty
Aug 23, 2022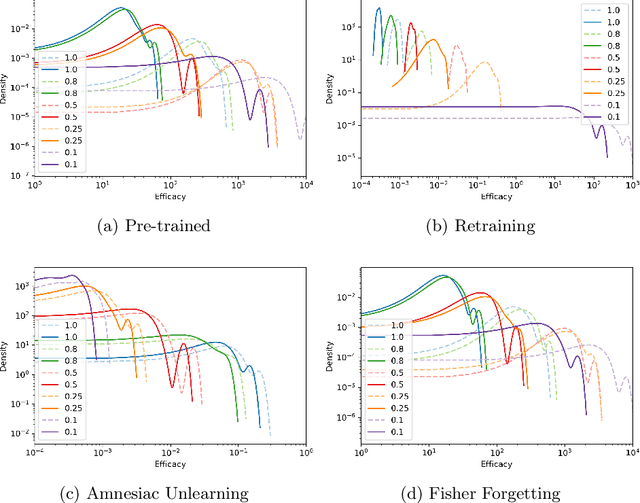
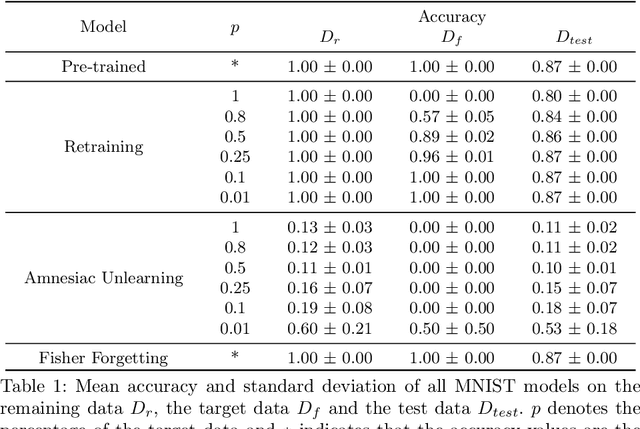
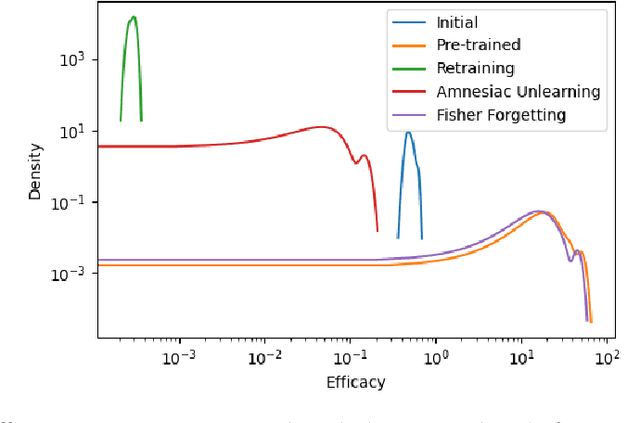
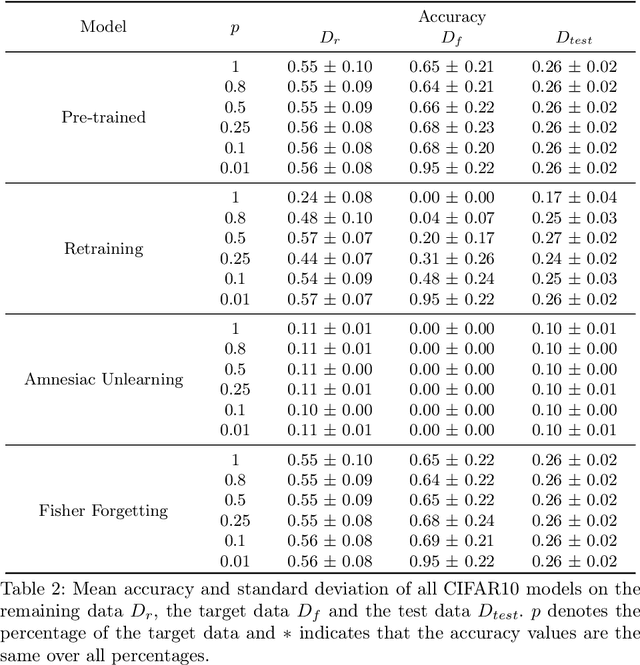
Abstract:There has been a growing interest in Machine Unlearning recently, primarily due to legal requirements such as the General Data Protection Regulation (GDPR) and the California Consumer Privacy Act. Thus, multiple approaches were presented to remove the influence of specific target data points from a trained model. However, when evaluating the success of unlearning, current approaches either use adversarial attacks or compare their results to the optimal solution, which usually incorporates retraining from scratch. We argue that both ways are insufficient in practice. In this work, we present an evaluation metric for Machine Unlearning algorithms based on epistemic uncertainty. This is the first definition of a general evaluation metric for Machine Unlearning to our best knowledge.
FiLM-Ensemble: Probabilistic Deep Learning via Feature-wise Linear Modulation
May 31, 2022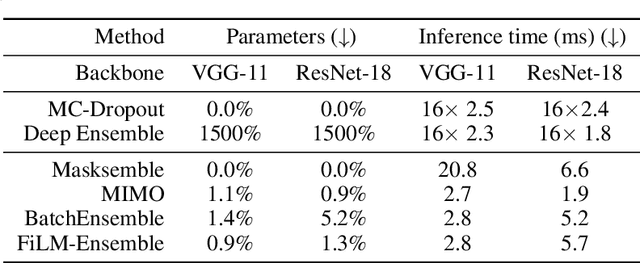
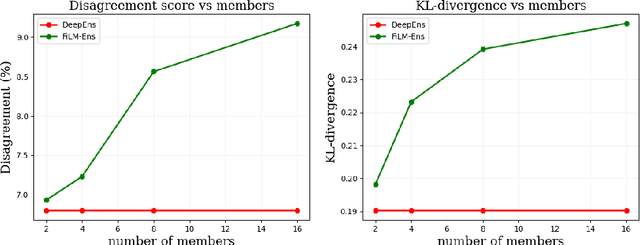
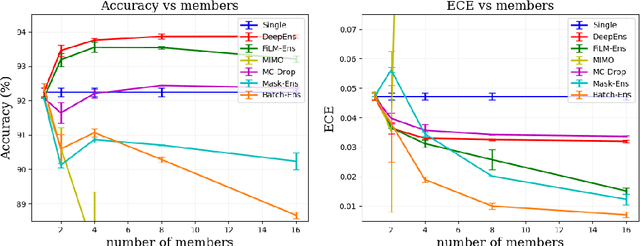
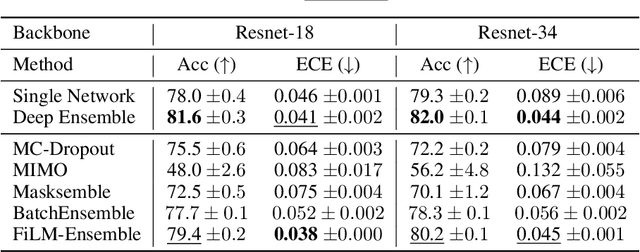
Abstract:The ability to estimate epistemic uncertainty is often crucial when deploying machine learning in the real world, but modern methods often produce overconfident, uncalibrated uncertainty predictions. A common approach to quantify epistemic uncertainty, usable across a wide class of prediction models, is to train a model ensemble. In a naive implementation, the ensemble approach has high computational cost and high memory demand. This challenges in particular modern deep learning, where even a single deep network is already demanding in terms of compute and memory, and has given rise to a number of attempts to emulate the model ensemble without actually instantiating separate ensemble members. We introduce FiLM-Ensemble, a deep, implicit ensemble method based on the concept of Feature-wise Linear Modulation (FiLM). That technique was originally developed for multi-task learning, with the aim of decoupling different tasks. We show that the idea can be extended to uncertainty quantification: by modulating the network activations of a single deep network with FiLM, one obtains a model ensemble with high diversity, and consequently well-calibrated estimates of epistemic uncertainty, with low computational overhead in comparison. Empirically, FiLM-Ensemble outperforms other implicit ensemble methods, and it and comes very close to the upper bound of an explicit ensemble of networks (sometimes even beating it), at a fraction of the memory cost.
Learning Graph Regularisation for Guided Super-Resolution
Mar 27, 2022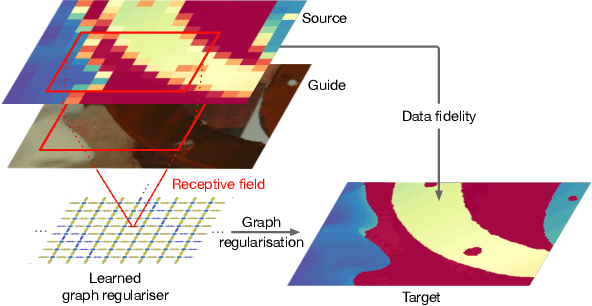
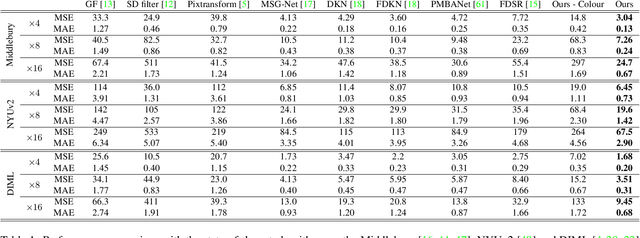
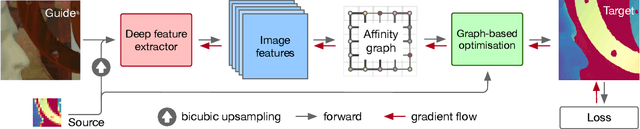
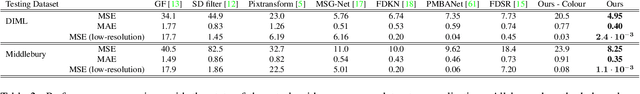
Abstract:We introduce a novel formulation for guided super-resolution. Its core is a differentiable optimisation layer that operates on a learned affinity graph. The learned graph potentials make it possible to leverage rich contextual information from the guide image, while the explicit graph optimisation within the architecture guarantees rigorous fidelity of the high-resolution target to the low-resolution source. With the decision to employ the source as a constraint rather than only as an input to the prediction, our method differs from state-of-the-art deep architectures for guided super-resolution, which produce targets that, when downsampled, will only approximately reproduce the source. This is not only theoretically appealing, but also produces crisper, more natural-looking images. A key property of our method is that, although the graph connectivity is restricted to the pixel lattice, the associated edge potentials are learned with a deep feature extractor and can encode rich context information over large receptive fields. By taking advantage of the sparse graph connectivity, it becomes possible to propagate gradients through the optimisation layer and learn the edge potentials from data. We extensively evaluate our method on several datasets, and consistently outperform recent baselines in terms of quantitative reconstruction errors, while also delivering visually sharper outputs. Moreover, we demonstrate that our method generalises particularly well to new datasets not seen during training.
Country-wide Retrieval of Forest Structure From Optical and SAR Satellite Imagery With Bayesian Deep Learning
Nov 25, 2021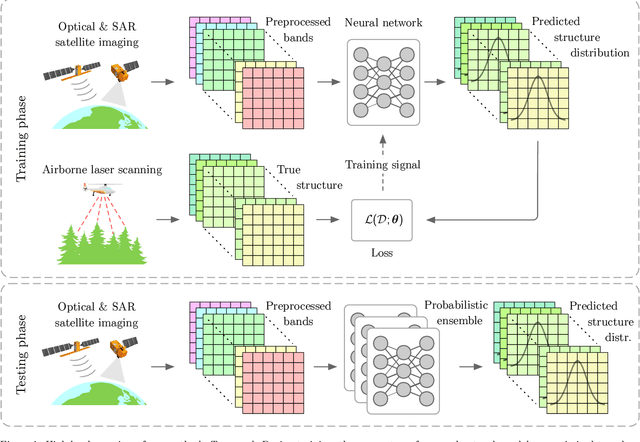
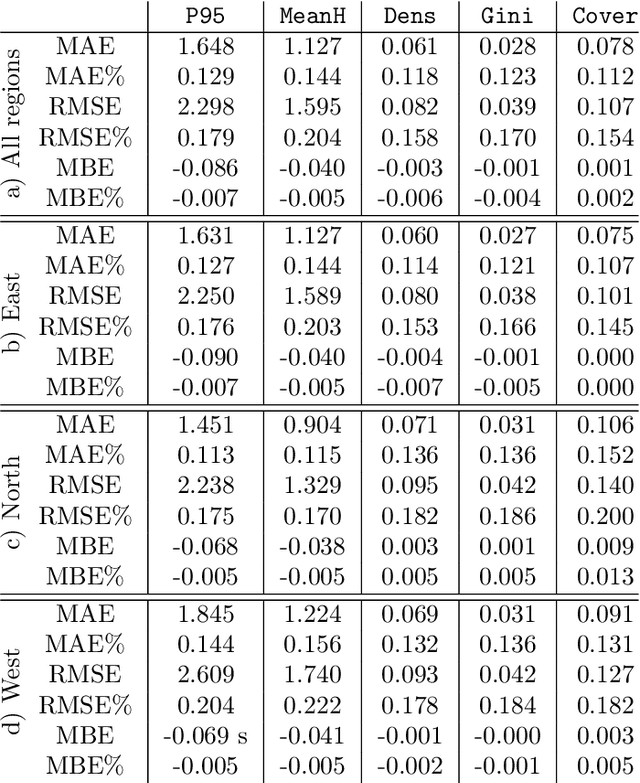
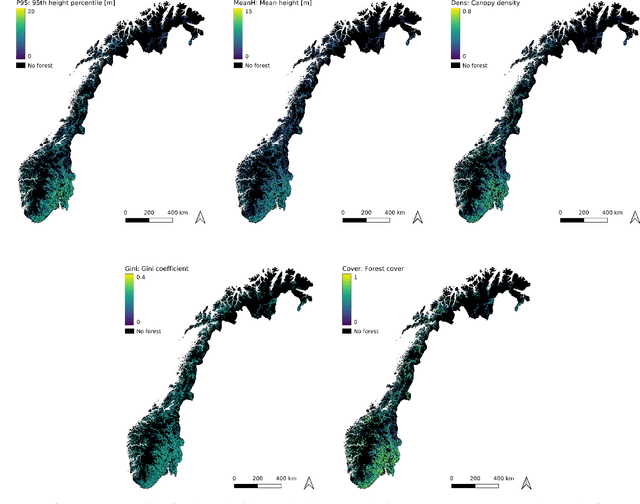
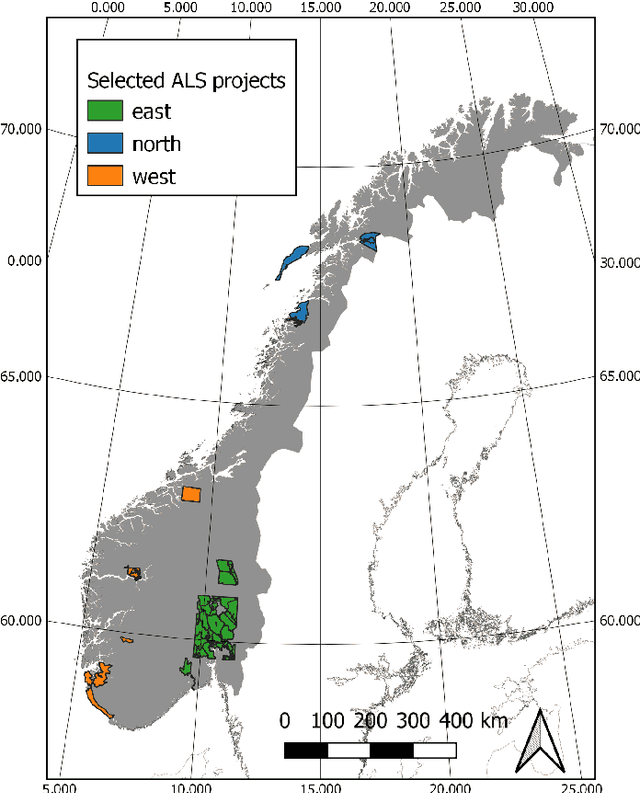
Abstract:Monitoring and managing Earth's forests in an informed manner is an important requirement for addressing challenges like biodiversity loss and climate change. While traditional in situ or aerial campaigns for forest assessments provide accurate data for analysis at regional level, scaling them to entire countries and beyond with high temporal resolution is hardly possible. In this work, we propose a Bayesian deep learning approach to densely estimate forest structure variables at country-scale with 10-meter resolution, using freely available satellite imagery as input. Our method jointly transforms Sentinel-2 optical images and Sentinel-1 synthetic aperture radar images into maps of five different forest structure variables: 95th height percentile, mean height, density, Gini coefficient, and fractional cover. We train and test our model on reference data from 41 airborne laser scanning missions across Norway and demonstrate that it is able to generalize to unseen test regions, achieving normalized mean absolute errors between 11% and 15%, depending on the variable. Our work is also the first to propose a Bayesian deep learning approach so as to predict forest structure variables with well-calibrated uncertainty estimates. These increase the trustworthiness of the model and its suitability for downstream tasks that require reliable confidence estimates, such as informed decision making. We present an extensive set of experiments to validate the accuracy of the predicted maps as well as the quality of the predicted uncertainties. To demonstrate scalability, we provide Norway-wide maps for the five forest structure variables.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge