Aidong Zhang
GCAV: A Global Concept Activation Vector Framework for Cross-Layer Consistency in Interpretability
Aug 28, 2025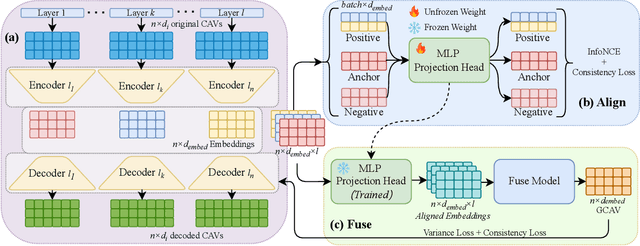

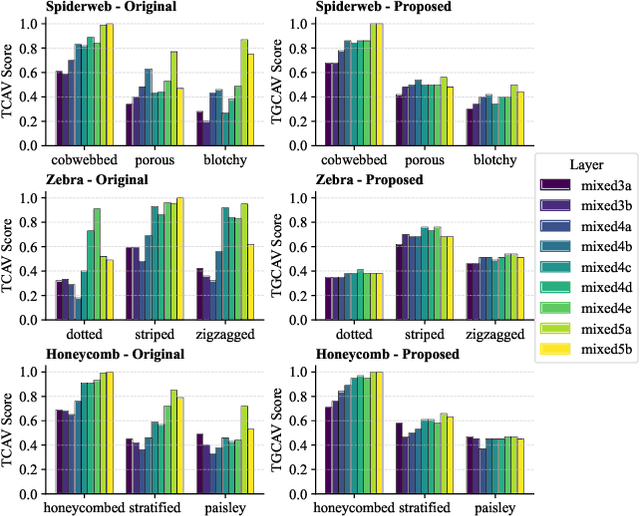

Abstract:Concept Activation Vectors (CAVs) provide a powerful approach for interpreting deep neural networks by quantifying their sensitivity to human-defined concepts. However, when computed independently at different layers, CAVs often exhibit inconsistencies, making cross-layer comparisons unreliable. To address this issue, we propose the Global Concept Activation Vector (GCAV), a novel framework that unifies CAVs into a single, semantically consistent representation. Our method leverages contrastive learning to align concept representations across layers and employs an attention-based fusion mechanism to construct a globally integrated CAV. By doing so, our method significantly reduces the variance in TCAV scores while preserving concept relevance, ensuring more stable and reliable concept attributions. To evaluate the effectiveness of GCAV, we introduce Testing with Global Concept Activation Vectors (TGCAV) as a method to apply TCAV to GCAV-based representations. We conduct extensive experiments on multiple deep neural networks, demonstrating that our method effectively mitigates concept inconsistency across layers, enhances concept localization, and improves robustness against adversarial perturbations. By integrating cross-layer information into a coherent framework, our method offers a more comprehensive and interpretable understanding of how deep learning models encode human-defined concepts. Code and models are available at https://github.com/Zhenghao-He/GCAV.
MedCite: Can Language Models Generate Verifiable Text for Medicine?
Jun 07, 2025Abstract:Existing LLM-based medical question-answering systems lack citation generation and evaluation capabilities, raising concerns about their adoption in practice. In this work, we introduce \name, the first end-to-end framework that facilitates the design and evaluation of citation generation with LLMs for medical tasks. Meanwhile, we introduce a novel multi-pass retrieval-citation method that generates high-quality citations. Our evaluation highlights the challenges and opportunities of citation generation for medical tasks, while identifying important design choices that have a significant impact on the final citation quality. Our proposed method achieves superior citation precision and recall improvements compared to strong baseline methods, and we show that evaluation results correlate well with annotation results from professional experts.
NeuronTune: Towards Self-Guided Spurious Bias Mitigation
May 29, 2025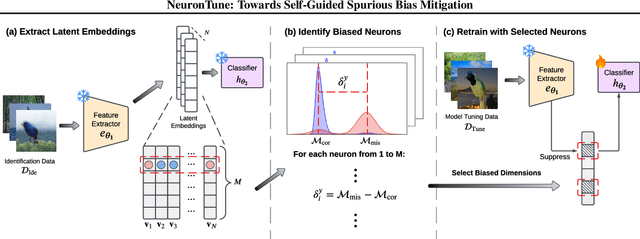
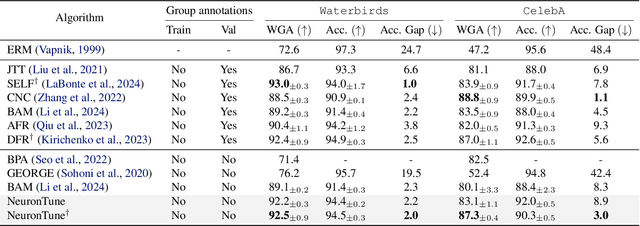
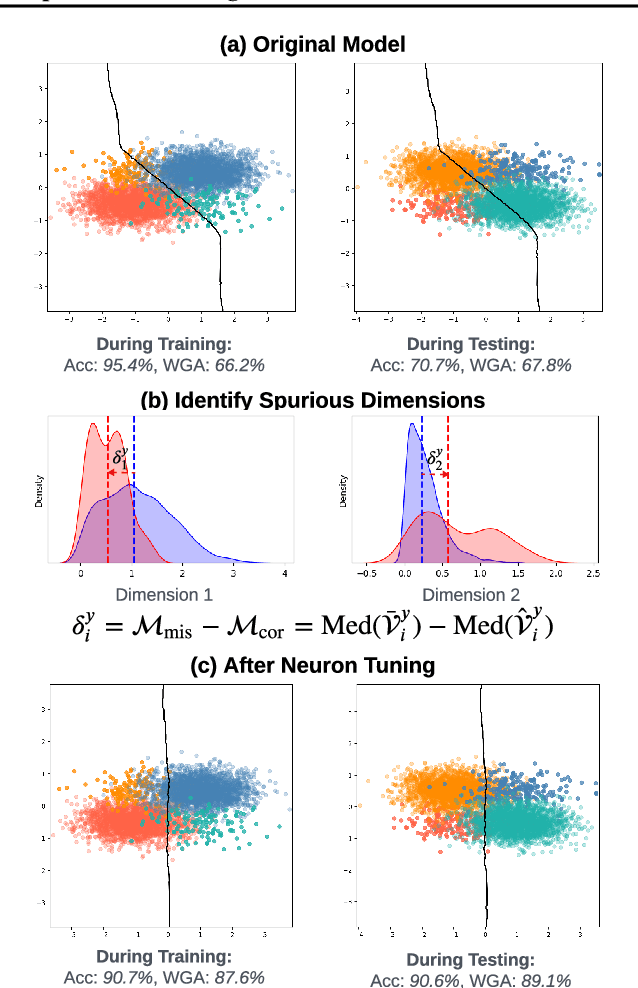
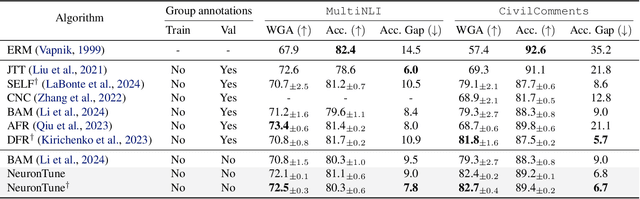
Abstract:Deep neural networks often develop spurious bias, reliance on correlations between non-essential features and classes for predictions. For example, a model may identify objects based on frequently co-occurring backgrounds rather than intrinsic features, resulting in degraded performance on data lacking these correlations. Existing mitigation approaches typically depend on external annotations of spurious correlations, which may be difficult to obtain and are not relevant to the spurious bias in a model. In this paper, we take a step towards self-guided mitigation of spurious bias by proposing NeuronTune, a post hoc method that directly intervenes in a model's internal decision process. Our method probes in a model's latent embedding space to identify and regulate neurons that lead to spurious prediction behaviors. We theoretically justify our approach and show that it brings the model closer to an unbiased one. Unlike previous methods, NeuronTune operates without requiring spurious correlation annotations, making it a practical and effective tool for improving model robustness. Experiments across different architectures and data modalities demonstrate that our method significantly mitigates spurious bias in a self-guided way.
Toward Reliable Biomedical Hypothesis Generation: Evaluating Truthfulness and Hallucination in Large Language Models
May 20, 2025Abstract:Large language models (LLMs) have shown significant potential in scientific disciplines such as biomedicine, particularly in hypothesis generation, where they can analyze vast literature, identify patterns, and suggest research directions. However, a key challenge lies in evaluating the truthfulness of generated hypotheses, as verifying their accuracy often requires substantial time and resources. Additionally, the hallucination problem in LLMs can lead to the generation of hypotheses that appear plausible but are ultimately incorrect, undermining their reliability. To facilitate the systematic study of these challenges, we introduce TruthHypo, a benchmark for assessing the capabilities of LLMs in generating truthful biomedical hypotheses, and KnowHD, a knowledge-based hallucination detector to evaluate how well hypotheses are grounded in existing knowledge. Our results show that LLMs struggle to generate truthful hypotheses. By analyzing hallucinations in reasoning steps, we demonstrate that the groundedness scores provided by KnowHD serve as an effective metric for filtering truthful hypotheses from the diverse outputs of LLMs. Human evaluations further validate the utility of KnowHD in identifying truthful hypotheses and accelerating scientific discovery. Our data and source code are available at https://github.com/Teddy-XiongGZ/TruthHypo.
ShortcutProbe: Probing Prediction Shortcuts for Learning Robust Models
May 20, 2025Abstract:Deep learning models often achieve high performance by inadvertently learning spurious correlations between targets and non-essential features. For example, an image classifier may identify an object via its background that spuriously correlates with it. This prediction behavior, known as spurious bias, severely degrades model performance on data that lacks the learned spurious correlations. Existing methods on spurious bias mitigation typically require a variety of data groups with spurious correlation annotations called group labels. However, group labels require costly human annotations and often fail to capture subtle spurious biases such as relying on specific pixels for predictions. In this paper, we propose a novel post hoc spurious bias mitigation framework without requiring group labels. Our framework, termed ShortcutProbe, identifies prediction shortcuts that reflect potential non-robustness in predictions in a given model's latent space. The model is then retrained to be invariant to the identified prediction shortcuts for improved robustness. We theoretically analyze the effectiveness of the framework and empirically demonstrate that it is an efficient and practical tool for improving a model's robustness to spurious bias on diverse datasets.
TAMA: A Human-AI Collaborative Thematic Analysis Framework Using Multi-Agent LLMs for Clinical Interviews
Mar 26, 2025Abstract:Thematic analysis (TA) is a widely used qualitative approach for uncovering latent meanings in unstructured text data. TA provides valuable insights in healthcare but is resource-intensive. Large Language Models (LLMs) have been introduced to perform TA, yet their applications in healthcare remain unexplored. Here, we propose TAMA: A Human-AI Collaborative Thematic Analysis framework using Multi-Agent LLMs for clinical interviews. We leverage the scalability and coherence of multi-agent systems through structured conversations between agents and coordinate the expertise of cardiac experts in TA. Using interview transcripts from parents of children with Anomalous Aortic Origin of a Coronary Artery (AAOCA), a rare congenital heart disease, we demonstrate that TAMA outperforms existing LLM-assisted TA approaches, achieving higher thematic hit rate, coverage, and distinctiveness. TAMA demonstrates strong potential for automated TA in clinical settings by leveraging multi-agent LLM systems with human-in-the-loop integration by enhancing quality while significantly reducing manual workload.
RAG-Gym: Optimizing Reasoning and Search Agents with Process Supervision
Feb 19, 2025



Abstract:Retrieval-augmented generation (RAG) has shown great potential for knowledge-intensive tasks, but its traditional architectures rely on static retrieval, limiting their effectiveness for complex questions that require sequential information-seeking. While agentic reasoning and search offer a more adaptive approach, most existing methods depend heavily on prompt engineering. In this work, we introduce RAG-Gym, a unified optimization framework that enhances information-seeking agents through fine-grained process supervision at each search step. We also propose ReSearch, a novel agent architecture that synergizes answer reasoning and search query generation within the RAG-Gym framework. Experiments on four challenging datasets show that RAG-Gym improves performance by up to 25.6\% across various agent architectures, with ReSearch consistently outperforming existing baselines. Further analysis highlights the effectiveness of advanced LLMs as process reward judges and the transferability of trained reward models as verifiers for different LLMs. Additionally, we examine the scaling properties of training and inference in agentic RAG. The project homepage is available at https://rag-gym.github.io/.
Client-Centric Federated Adaptive Optimization
Jan 17, 2025



Abstract:Federated Learning (FL) is a distributed learning paradigm where clients collaboratively train a model while keeping their own data private. With an increasing scale of clients and models, FL encounters two key challenges, client drift due to a high degree of statistical/system heterogeneity, and lack of adaptivity. However, most existing FL research is based on unrealistic assumptions that virtually ignore system heterogeneity. In this paper, we propose Client-Centric Federated Adaptive Optimization, which is a class of novel federated adaptive optimization approaches. We enable several features in this framework such as arbitrary client participation, asynchronous server aggregation, and heterogeneous local computing, which are ubiquitous in real-world FL systems but are missed in most existing works. We provide a rigorous convergence analysis of our proposed framework for general nonconvex objectives, which is shown to converge with the best-known rate. Extensive experiments show that our approaches consistently outperform the baseline by a large margin across benchmarks.
Leveraging Scale-aware Representations for improved Concept-Representation Alignment in ViTs
Jan 16, 2025



Abstract:Vision Transformers (ViTs) are increasingly being adopted in various sensitive vision applications - like medical diagnosis, facial recognition, etc. To improve the interpretability of such models, many approaches attempt to forward-align them with carefully annotated abstract, human-understandable semantic entities - concepts. Concepts provide global rationales to the model predictions and can be quickly understood/intervened on by domain experts. Most current research focuses on designing model-agnostic, plug-and-play generic concept-based explainability modules that do not incorporate the inner workings of foundation models (e.g., inductive biases, scale invariance, etc.) during training. To alleviate this issue for ViTs, in this paper, we propose a novel Concept Representation Alignment Module (CRAM) which learns both scale and position-aware representations from multi-scale feature pyramids and patch representations respectively. CRAM further aligns these representations with concept annotations through an attention matrix. The proposed CRAM module improves the predictive performance of ViT architectures and also provides accurate and robust concept explanations as demonstrated on five datasets - including three widely used benchmarks (CUB, Pascal APY, Concept-MNIST) and 2 real-world datasets (AWA2, KITS).
Ensuring Safety and Trust: Analyzing the Risks of Large Language Models in Medicine
Nov 20, 2024



Abstract:The remarkable capabilities of Large Language Models (LLMs) make them increasingly compelling for adoption in real-world healthcare applications. However, the risks associated with using LLMs in medical applications have not been systematically characterized. We propose using five key principles for safe and trustworthy medical AI: Truthfulness, Resilience, Fairness, Robustness, and Privacy, along with ten specific aspects. Under this comprehensive framework, we introduce a novel MedGuard benchmark with 1,000 expert-verified questions. Our evaluation of 11 commonly used LLMs shows that the current language models, regardless of their safety alignment mechanisms, generally perform poorly on most of our benchmarks, particularly when compared to the high performance of human physicians. Despite recent reports indicate that advanced LLMs like ChatGPT can match or even exceed human performance in various medical tasks, this study underscores a significant safety gap, highlighting the crucial need for human oversight and the implementation of AI safety guardrails.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge