"Time": models, code, and papers
Palm Vein Recognition via Multi-task Loss Function and Attention Layer
Nov 11, 2022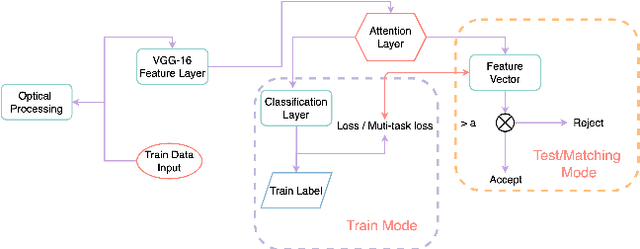
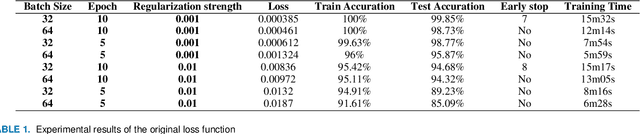
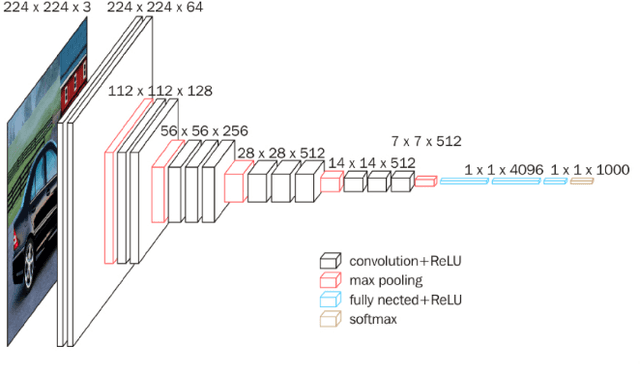
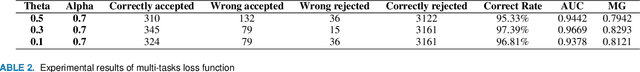
With the improvement of arithmetic power and algorithm accuracy of personal devices, biological features are increasingly widely used in personal identification, and palm vein recognition has rich extractable features and has been widely studied in recent years. However, traditional recognition methods are poorly robust and susceptible to environmental influences such as reflections and noise. In this paper, a convolutional neural network based on VGG-16 transfer learning fused attention mechanism is used as the feature extraction network on the infrared palm vein dataset. The palm vein classification task is first trained using palmprint classification methods, followed by matching using a similarity function, in which we propose the multi-task loss function to improve the accuracy of the matching task. In order to verify the robustness of the model, some experiments were carried out on datasets from different sources. Then, we used K-means clustering to determine the adaptive matching threshold and finally achieved an accuracy rate of 98.89% on prediction set. At the same time, the matching is with high efficiency which takes an average of 0.13 seconds per palm vein pair, and that means our method can be adopted in practice.
Efficient HLA imputation from sequential SNPs data by Transformer
Nov 11, 2022
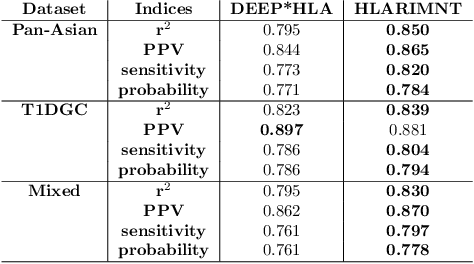
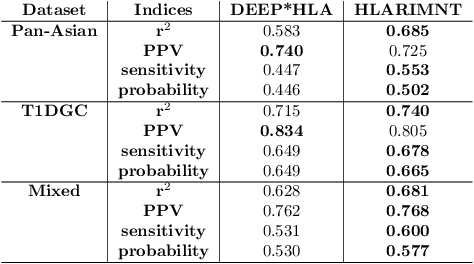
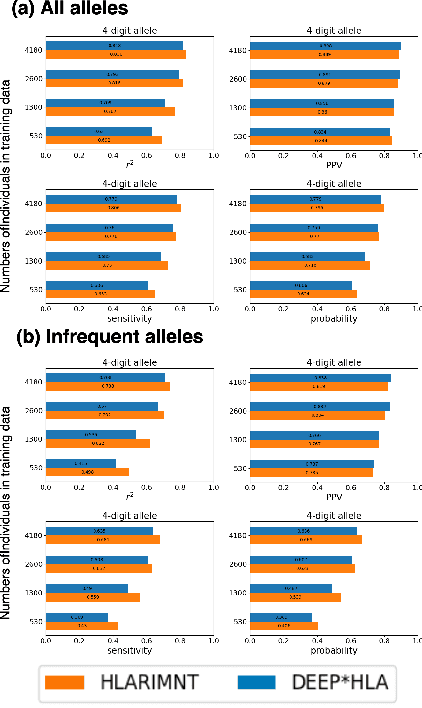
Human leukocyte antigen (HLA) genes are associated with a variety of diseases, however direct typing of HLA is time and cost consuming. Thus various imputation methods using sequential SNPs data have been proposed based on statistical or deep learning models, e.g. CNN-based model, named DEEP*HLA. However, imputation efficiency is not sufficient for in frequent alleles and a large size of reference panel is required. Here, we developed a Transformer-based model to impute HLA alleles, named "HLA Reliable IMputatioN by Transformer (HLARIMNT)" to take advantage of sequential nature of SNPs data. We validated the performance of HLARIMNT using two different reference panels; Pan-Asian reference panel (n = 530) and Type 1 Diabetes Genetics Consortium (T1DGC) reference panel (n = 5,225), as well as the mixture of those two panels (n = 1,060). HLARIMNT achieved higher accuracy than DEEP*HLA by several indices, especially for infrequent alleles. We also varied the size of data used for training, and HLARIMNT imputed more accurately among any size of training data. These results suggest that Transformer-based model may impute efficiently not only HLA types but also any other gene types from sequential SNPs data.
Beyond Homophily with Graph Echo State Networks
Oct 27, 2022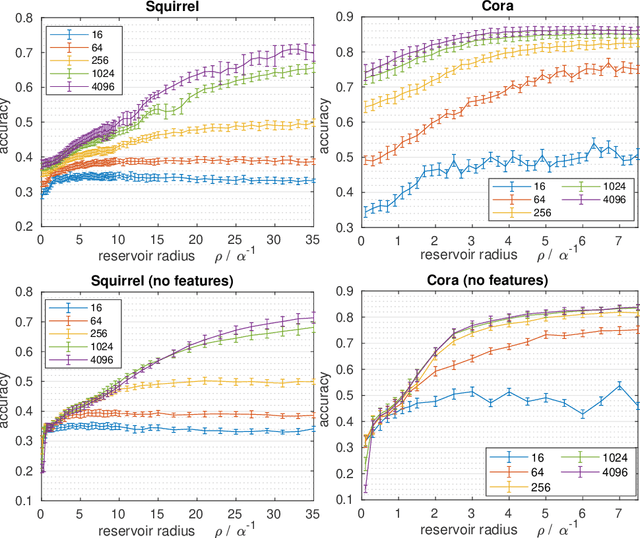
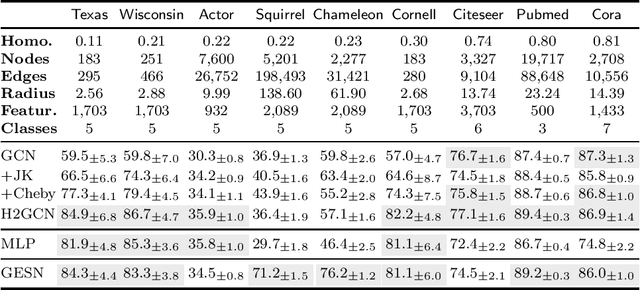
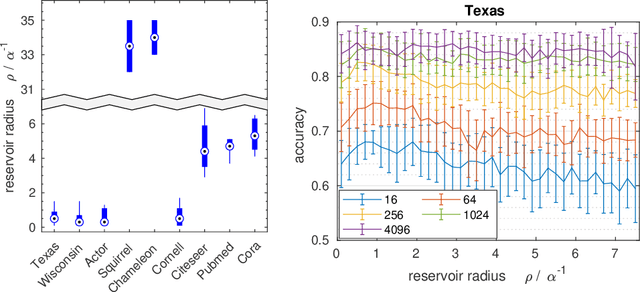
Graph Echo State Networks (GESN) have already demonstrated their efficacy and efficiency in graph classification tasks. However, semi-supervised node classification brought out the problem of over-smoothing in end-to-end trained deep models, which causes a bias towards high homophily graphs. We evaluate for the first time GESN on node classification tasks with different degrees of homophily, analyzing also the impact of the reservoir radius. Our experiments show that reservoir models are able to achieve better or comparable accuracy with respect to fully trained deep models that implement ad hoc variations in the architectural bias, with a gain in terms of efficiency.
* Accepted for oral presentation at ESANN 2022
HQNAS: Auto CNN deployment framework for joint quantization and architecture search
Oct 16, 2022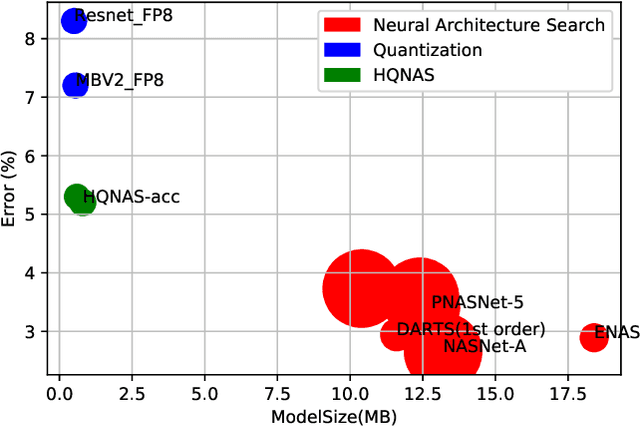
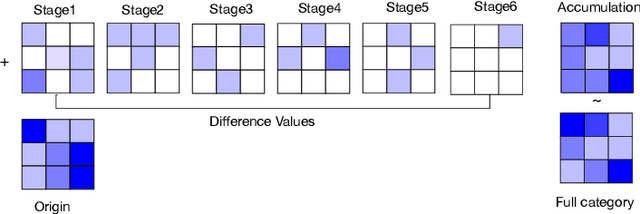
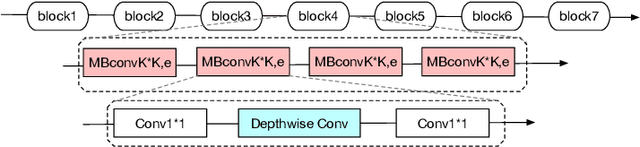
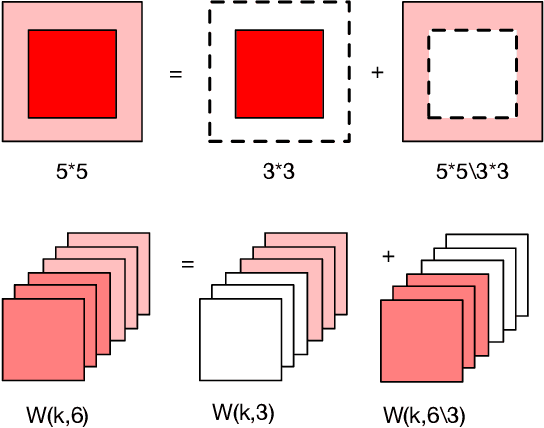
Deep learning applications are being transferred from the cloud to edge with the rapid development of embedded computing systems. In order to achieve higher energy efficiency with the limited resource budget, neural networks(NNs) must be carefully designed in two steps, the architecture design and the quantization policy choice. Neural Architecture Search(NAS) and Quantization have been proposed separately when deploying NNs onto embedded devices. However, taking the two steps individually is time-consuming and leads to a sub-optimal final deployment. To this end, we propose a novel neural network design framework called Hardware-aware Quantized Neural Architecture Search(HQNAS) framework which combines the NAS and Quantization together in a very efficient manner using weight-sharing and bit-sharing. It takes only 4 GPU hours to discover an outstanding NN policy on CIFAR10. It also takes only %10 GPU time to generate a comparable model on Imagenet compared to the traditional NAS method with 1.8x decrease of latency and a negligible accuracy loss of only 0.7%. Besides, our method can be adapted in a lifelong situation where the neural network needs to evolve occasionally due to changes of local data, environment and user preference.
Estimation of Shade Losses in Unlabeled PV Data
Sep 20, 2022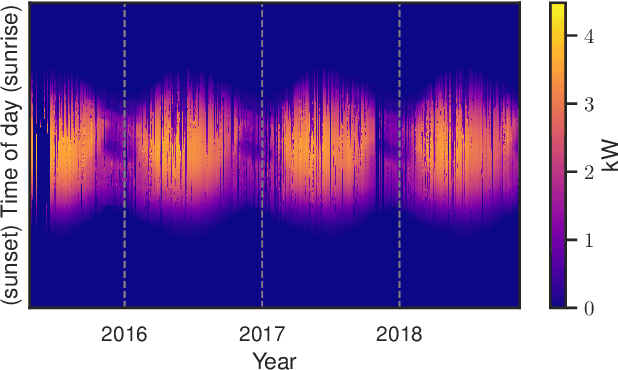
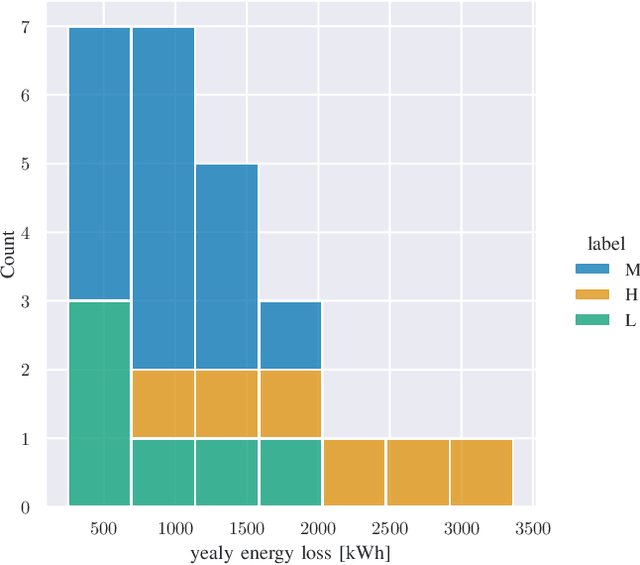
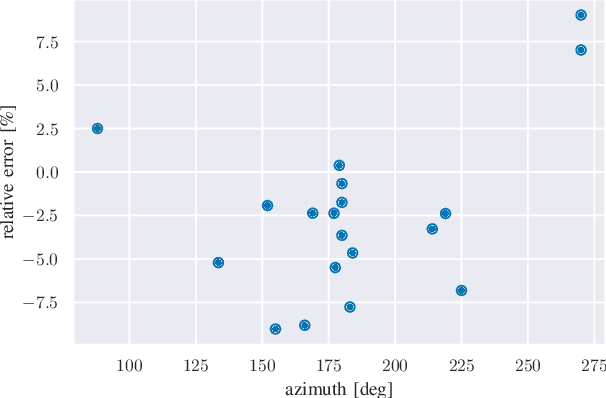
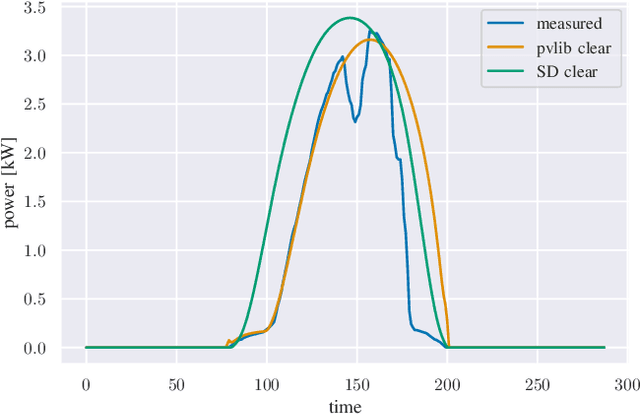
We provide a methodology for estimating the losses due to shade in power generation data sets produced by real-world photovoltaic (PV) systems. We focus this work on estimating shade loss from data that are unlabeled, i.e. power measurements with time stamps but no other information such as site configuration or meteorological data. This approach enables, for the first time, the analysis of data generated by small scale, distributed PV systems, which do not have the data quality or richness of large, utility-scale PV systems or research-grade installations. This work is an application of the newly published signal decomposition (SD) framework, which provides an extensible approach for estimating hidden components in time-series data.
Travel time optimization on multi-AGV routing by reverse annealing
Apr 25, 2022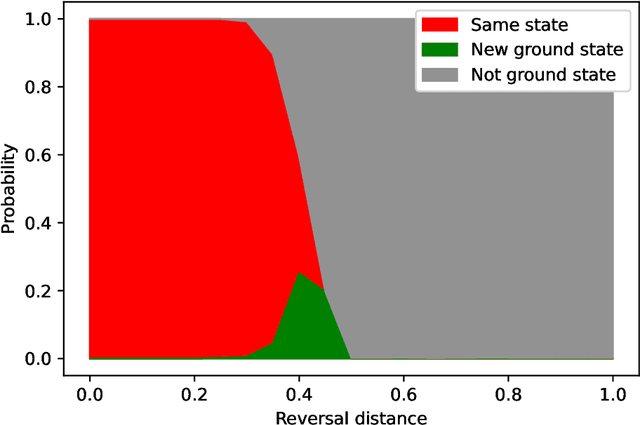

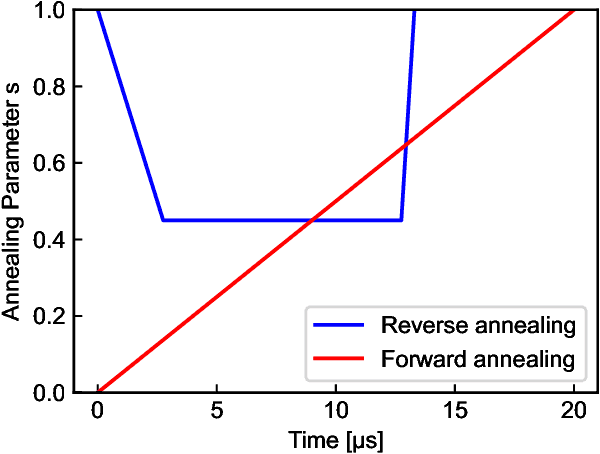
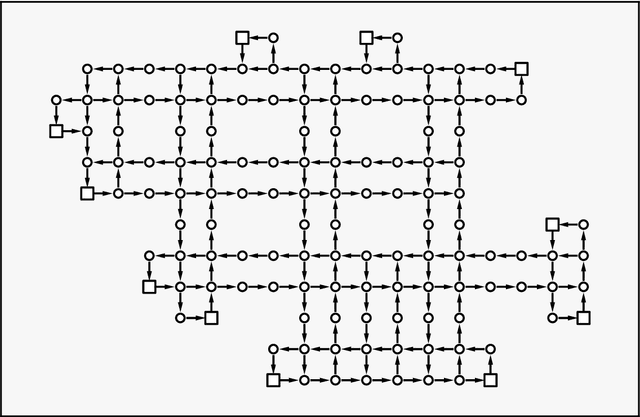
Quantum annealing has been actively researched since D-Wave Systems produced the first commercial machine in 2011. Controlling a large fleet of automated guided vehicles is one of the real-world applications utilizing quantum annealing. In this study, we propose a formulation to control the traveling routes to minimize the travel time. We validate our formulation through simulation in a virtual plant and authenticate the effectiveness for faster distribution compared to a greedy algorithm that does not consider the overall detour distance. Furthermore, we utilize reverse annealing to maximize the advantage of the D-Wave's quantum annealer. Starting from relatively good solutions obtained by a fast greedy algorithm, reverse annealing searches for better solutions around them. Our reverse annealing method improves the performance compared to standard quantum annealing alone and performs up to 10 times faster than the strong classical solver, Gurobi. This study extends a use of optimization with general problem solvers in the application of multi-AGV systems and reveals the potential of reverse annealing as an optimizer.
On the Multidimensional Augmentation of Fingerprint Data for Indoor Localization in A Large-Scale Building Complex Based on Multi-Output Gaussian Process
Nov 19, 2022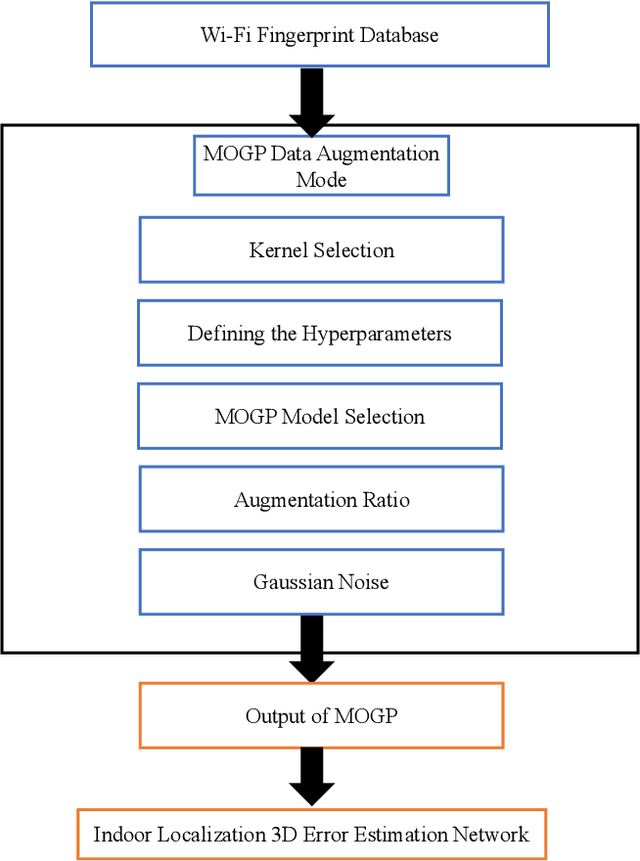
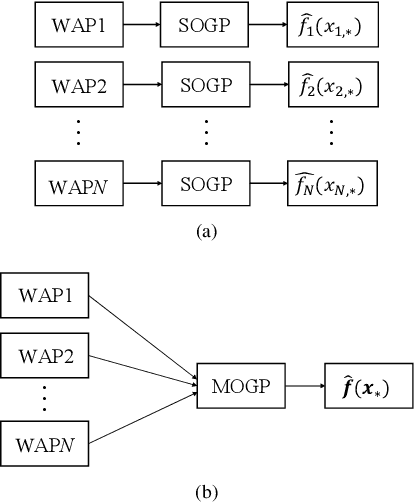
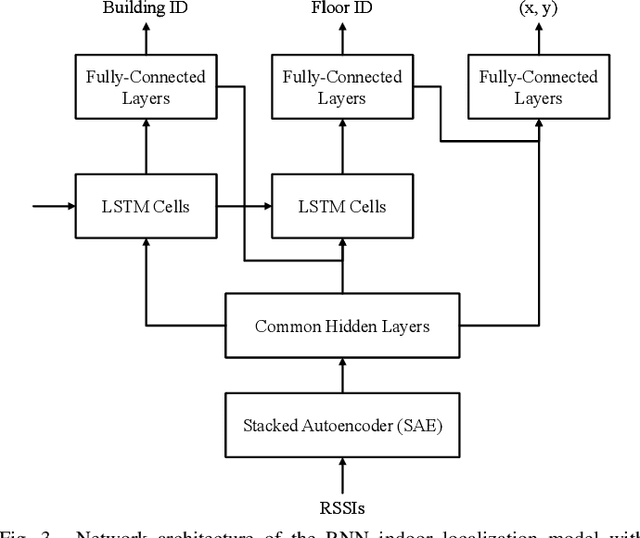
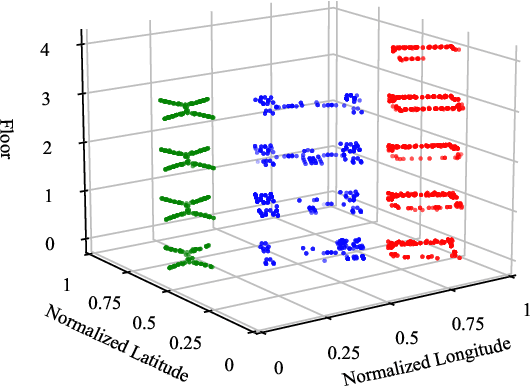
Wi-Fi fingerprinting becomes a dominant solution for large-scale indoor localization due to its major advantage of not requiring new infrastructure and dedicated devices. The number and the distribution of Reference Points (RPs) for the measurement of localization fingerprints like RSSI during the offline phase, however, greatly affects the localization accuracy; for instance, the UJIIndoorLoc is known to have the issue of uneven spatial distribution of RPs over buildings and floors. Data augmentation has been proposed as a feasible solution to not only improve the smaller number and the uneven distribution of RPs in the existing fingerprint databases but also reduce the labor and time costs of constructing new fingerprint databases. In this paper, we propose the multidimensional augmentation of fingerprint data for indoor localization in a large-scale building complex based on Multi-Output Gaussian Process (MOGP) and systematically investigate the impact of augmentation ratio as well as MOGP kernel functions and models with their hyperparameters on the performance of indoor localization using the UJIIndoorLoc database and the state-of-the-art neural network indoor localization model based on a hierarchical RNN. The investigation based on experimental results suggests that we can generate synthetic RSSI fingerprint data up to ten times the original data -- i.e., the augmentation ratio of 10 -- through the proposed multidimensional MOGP-based data augmentation without significantly affecting the indoor localization performance compared to that of the original data alone, which extends the spatial coverage of the combined RPs and thereby could improve the localization performance at the locations that are not part of the test dataset.
Solving 3D Inverse Problems using Pre-trained 2D Diffusion Models
Nov 19, 2022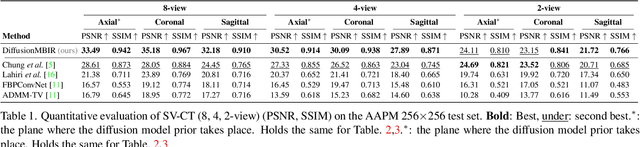
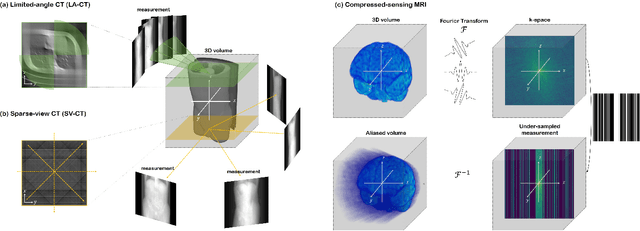
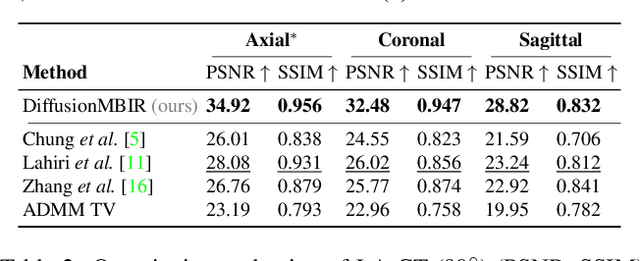
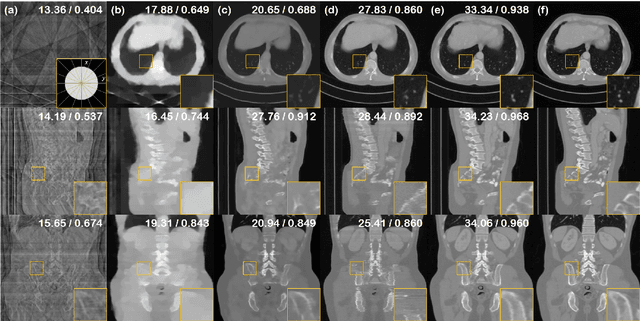
Diffusion models have emerged as the new state-of-the-art generative model with high quality samples, with intriguing properties such as mode coverage and high flexibility. They have also been shown to be effective inverse problem solvers, acting as the prior of the distribution, while the information of the forward model can be granted at the sampling stage. Nonetheless, as the generative process remains in the same high dimensional (i.e. identical to data dimension) space, the models have not been extended to 3D inverse problems due to the extremely high memory and computational cost. In this paper, we combine the ideas from the conventional model-based iterative reconstruction with the modern diffusion models, which leads to a highly effective method for solving 3D medical image reconstruction tasks such as sparse-view tomography, limited angle tomography, compressed sensing MRI from pre-trained 2D diffusion models. In essence, we propose to augment the 2D diffusion prior with a model-based prior in the remaining direction at test time, such that one can achieve coherent reconstructions across all dimensions. Our method can be run in a single commodity GPU, and establishes the new state-of-the-art, showing that the proposed method can perform reconstructions of high fidelity and accuracy even in the most extreme cases (e.g. 2-view 3D tomography). We further reveal that the generalization capacity of the proposed method is surprisingly high, and can be used to reconstruct volumes that are entirely different from the training dataset.
Automatic Quantitative Analysis of Brain Organoids via Deep Learning
Nov 01, 2022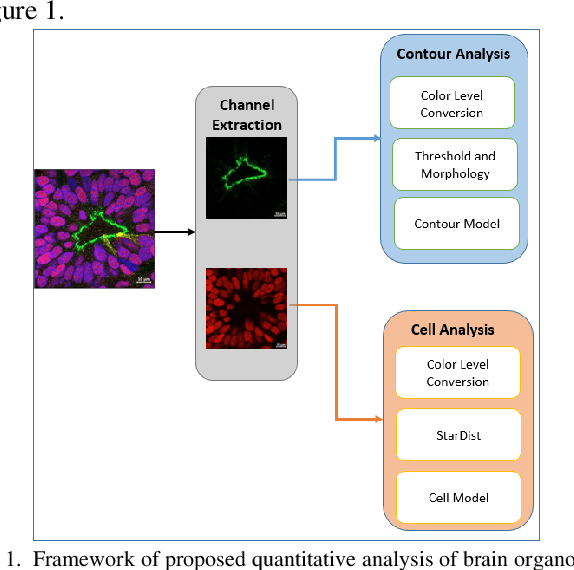
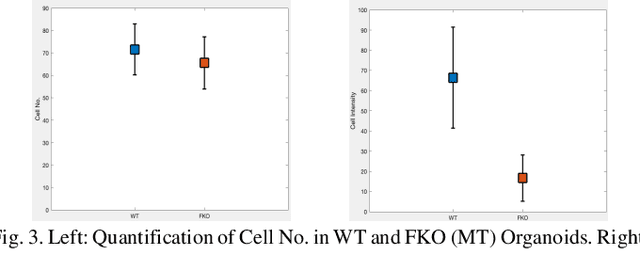
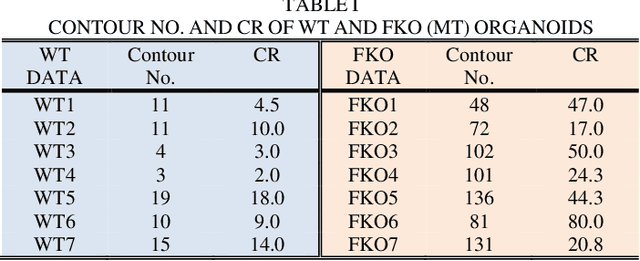
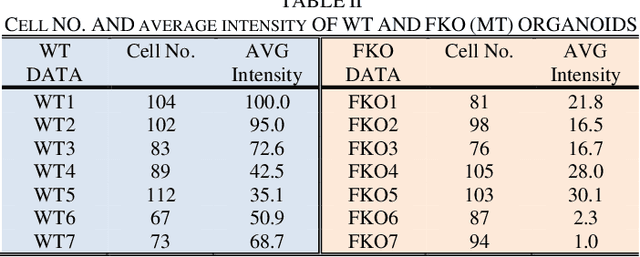
Recent advances in brain organoid technology are exciting new ways, which have the potential to change the way how doctors and researchers understand and treat cerebral diseases. Despite the remarkable use of brain organoids derived from human stem cells in new drug testing, disease modeling, and scientific research, it is still heavily time-consuming work to observe and analyze the internal structure, cells, and neural inside the organoid by humans, specifically no standard quantitative analysis method combined growing AI technology for brain organoid. In this paper, an automated computer-assisted analysis method is proposed for brain organoid slice channels tagged with different fluorescent. We applied the method on two channels of two group microscopy images and the experiment result shows an obvious difference between Wild Type and Mutant Type cerebral organoids.
Text-Only Training for Image Captioning using Noise-Injected CLIP
Nov 01, 2022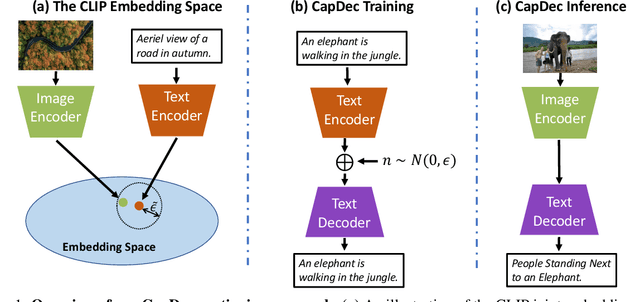
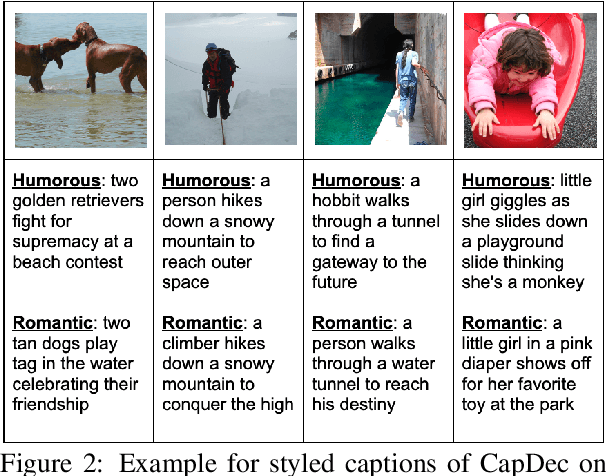
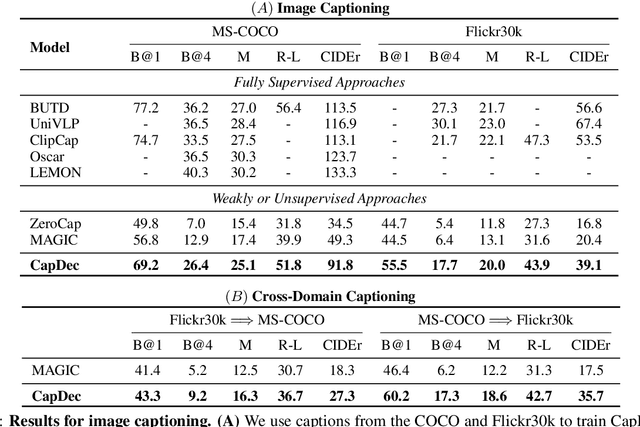
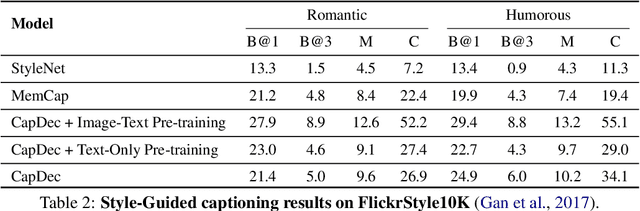

We consider the task of image-captioning using only the CLIP model and additional text data at training time, and no additional captioned images. Our approach relies on the fact that CLIP is trained to make visual and textual embeddings similar. Therefore, we only need to learn how to translate CLIP textual embeddings back into text, and we can learn how to do this by learning a decoder for the frozen CLIP text encoder using only text. We argue that this intuition is "almost correct" because of a gap between the embedding spaces, and propose to rectify this via noise injection during training. We demonstrate the effectiveness of our approach by showing SOTA zero-shot image captioning across four benchmarks, including style transfer. Code, data, and models are available on GitHub.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge