"Image": models, code, and papers
Multi-task Image Classification via Collaborative, Hierarchical Spike-and-Slab Priors
Jan 30, 2015

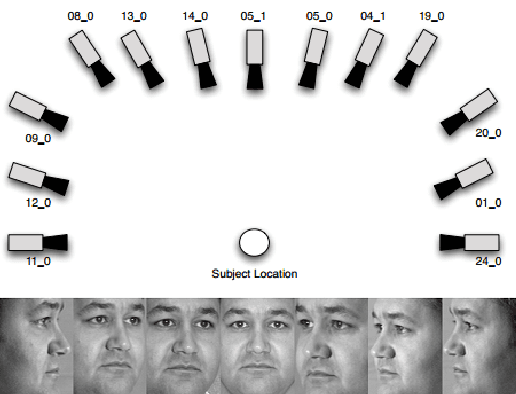
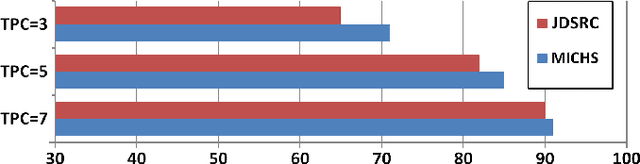
Promising results have been achieved in image classification problems by exploiting the discriminative power of sparse representations for classification (SRC). Recently, it has been shown that the use of \emph{class-specific} spike-and-slab priors in conjunction with the class-specific dictionaries from SRC is particularly effective in low training scenarios. As a logical extension, we build on this framework for multitask scenarios, wherein multiple representations of the same physical phenomena are available. We experimentally demonstrate the benefits of mining joint information from different camera views for multi-view face recognition.
Saliency Methods for Explaining Adversarial Attacks
Oct 02, 2019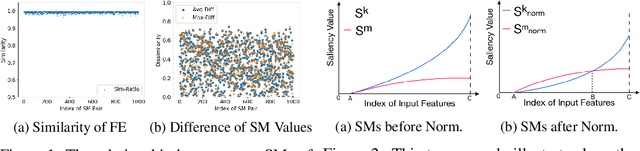
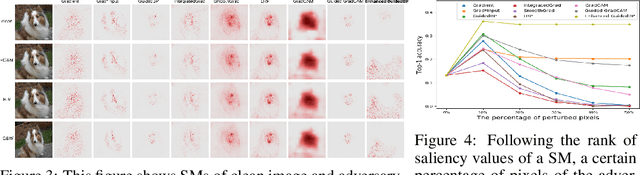

The classification decisions of neural networks can be misled by small imperceptible perturbations. This work aims to explain the misled classifications using saliency methods. The idea behind saliency methods is to explain the classification decisions of neural networks by creating so-called saliency maps. Unfortunately, a number of recent publications have shown that many of the proposed saliency methods do not provide insightful explanations. A prominent example is Guided Backpropagation (GuidedBP), which simply performs (partial) image recovery. However, our numerical analysis shows the saliency maps created by GuidedBP do indeed contain class-discriminative information. We propose a simple and efficient way to enhance the saliency maps. The proposed enhanced GuidedBP shows the state-of-the-art performance to explain adversary classifications.
Segmentation of Retinal Low-Cost Optical Coherence Tomography Images using Deep Learning
Jan 23, 2020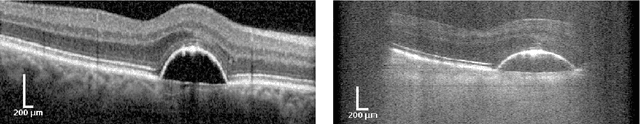
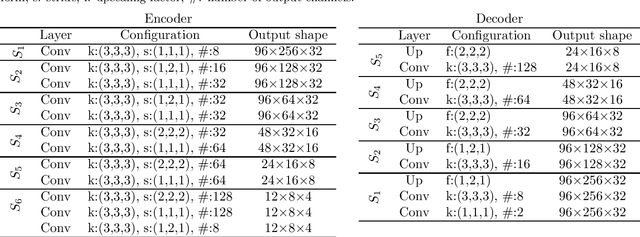
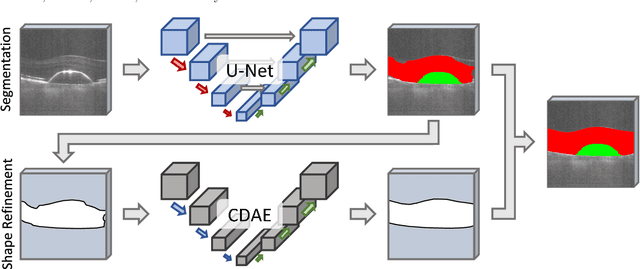
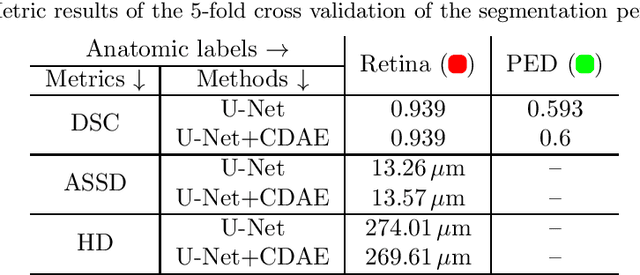
The treatment of age-related macular degeneration (AMD) requires continuous eye exams using optical coherence tomography (OCT). The need for treatment is determined by the presence or change of disease-specific OCT-based biomarkers. Therefore, the monitoring frequency has a significant influence on the success of AMD therapy. However, the monitoring frequency of current treatment schemes is not individually adapted to the patient and therefore often insufficient. While a higher monitoring frequency would have a positive effect on the success of treatment, in practice it can only be achieved with a home monitoring solution. One of the key requirements of a home monitoring OCT system is a computer-aided diagnosis to automatically detect and quantify pathological changes using specific OCT-based biomarkers. In this paper, for the first time, retinal scans of a novel self-examination low-cost full-field OCT (SELF-OCT) are segmented using a deep learning-based approach. A convolutional neural network (CNN) is utilized to segment the total retina as well as pigment epithelial detachments (PED). It is shown that the CNN-based approach can segment the retina with high accuracy, whereas the segmentation of the PED proves to be challenging. In addition, a convolutional denoising autoencoder (CDAE) refines the CNN prediction, which has previously learned retinal shape information. It is shown that the CDAE refinement can correct segmentation errors caused by artifacts in the OCT image.
Robust Compressive Sensing MRI Reconstruction using Generative Adversarial Networks
Oct 14, 2019
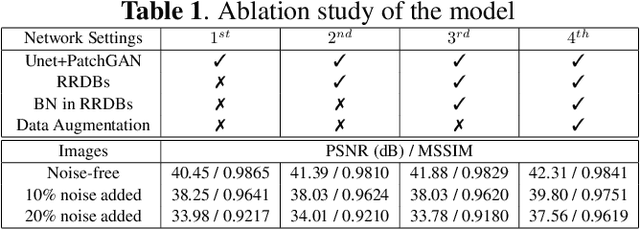
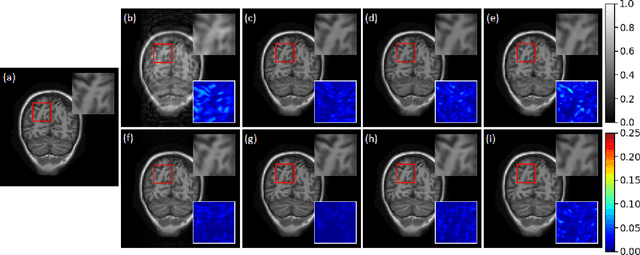
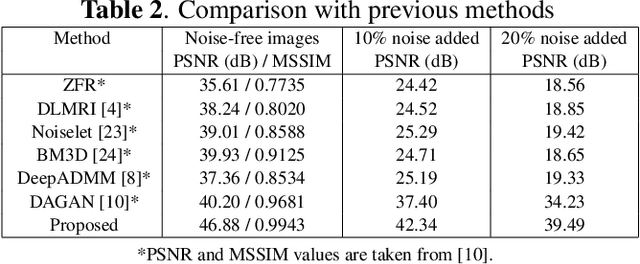
Compressive sensing magnetic resonance imaging (CS-MRI) accelerates the acquisition of MR images by breaking the Nyquist sampling limit. In this work, a novel generative adversarial network (GAN) based framework for CS-MRI reconstruction is proposed. Leveraging a combination of patchGAN discriminator and structural similarity index based loss, our model focuses on preserving high frequency content as well as fine textural details in the reconstructed image. Dense and residual connections have been incorporated in a U-net based generator architecture to allow easier transfer of information as well as variable network length. We show that our algorithm outperforms state-of-the-art methods in terms of quality of reconstruction and robustness to noise. Also, the reconstruction time, which is of the order of milliseconds, makes it highly suitable for real-time clinical use.
A Novel Image Segmentation Enhancement Technique based on Active Contour and Topological Alignments
Jun 02, 2011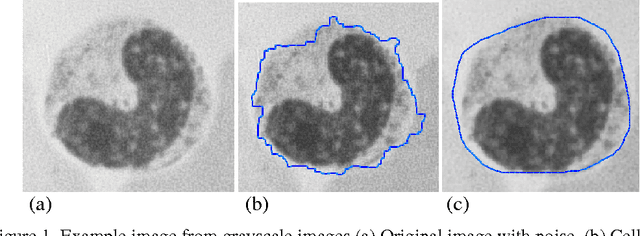
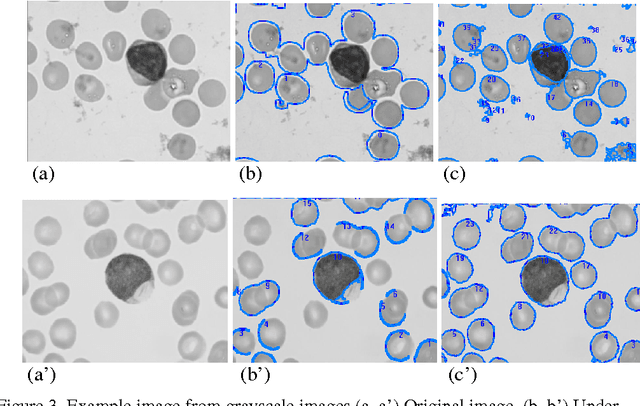
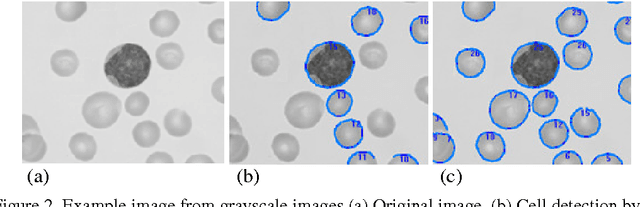

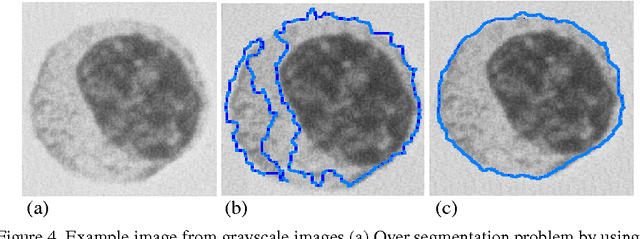
Topological alignments and snakes are used in image processing, particularly in locating object boundaries. Both of them have their own advantages and limitations. To improve the overall image boundary detection system, we focused on developing a novel algorithm for image processing. The algorithm we propose to develop will based on the active contour method in conjunction with topological alignments method to enhance the image detection approach. The algorithm presents novel technique to incorporate the advantages of both Topological Alignments and snakes. Where the initial segmentation by Topological Alignments is firstly transformed into the input of the snake model and begins its evolvement to the interested object boundary. The results show that the algorithm can deal with low contrast images and shape cells, demonstrate the segmentation accuracy under weak image boundaries, which responsible for lacking accuracy in image detecting techniques. We have achieved better segmentation and boundary detecting for the image, also the ability of the system to improve the low contrast and deal with over and under segmentation.
* 7 pages
A deep learning based tool for automatic brain extraction from functional magnetic resonance images in rodents
Dec 06, 2019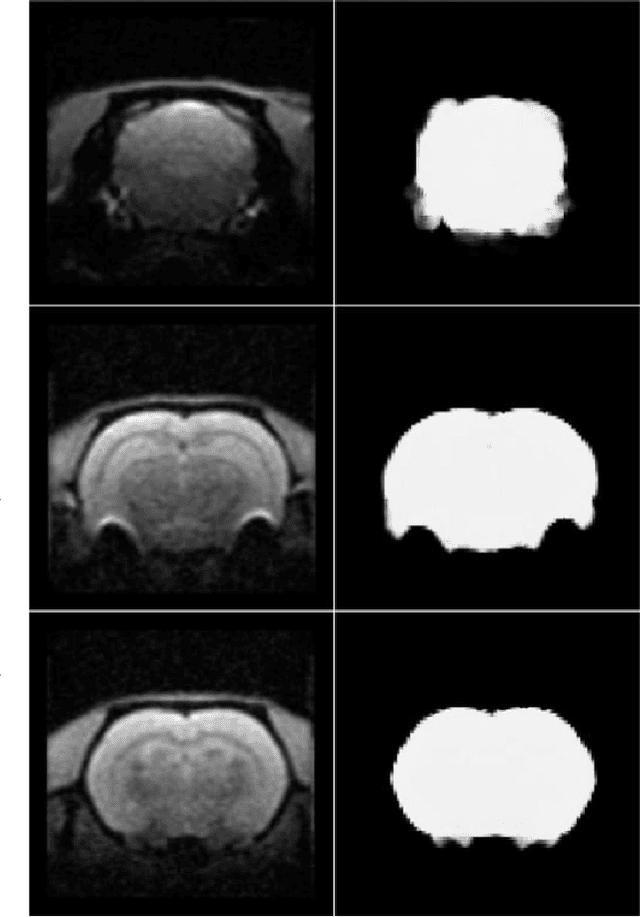
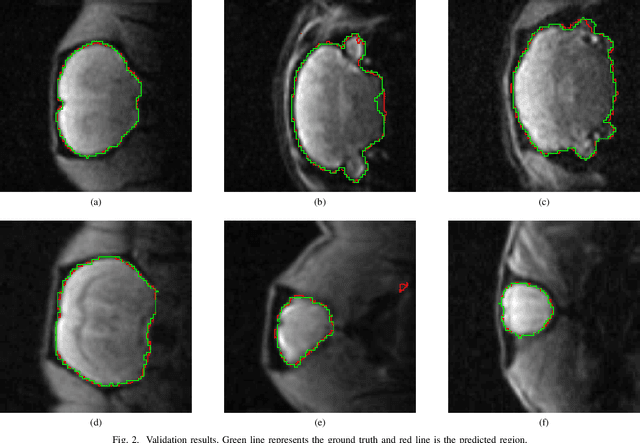
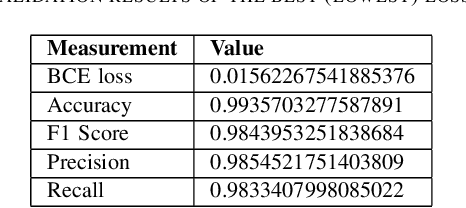
Removing skull artifacts from functional magnetic images (fMRI) is a well understood and frequently encountered problem. Because the fMRI field has grown mostly due to human studies, many new tools were developed to handle human data. Nonetheless, these tools are not equally useful to handle the data derived from animal studies, especially from rodents. This represents a major problem to the field because rodent studies generate larger datasets from larger populations, which implies that preprocessing these images manually to remove the skull becomes a bottleneck in the data analysis pipeline. In this study, we address this problem by implementing a neural network based method that uses a U-Net architecture to segment the brain area into a mask and removing the skull and other tissues from the image. We demonstrate several strategies to speed up the process of generating the training dataset using watershedding and several strategies for data augmentation that allowed to train faster the U-Net to perform the segmentation. Finally, we deployed the trained network freely available.
A Deep Factorization of Style and Structure in Fonts
Oct 02, 2019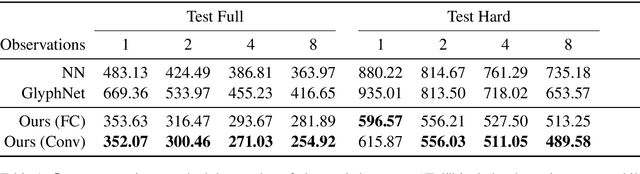
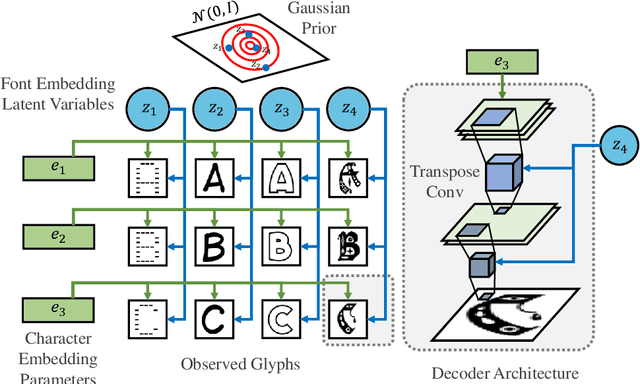
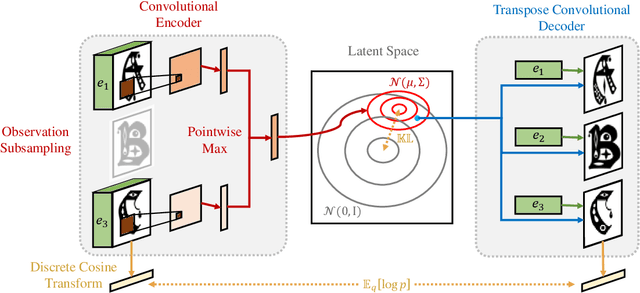
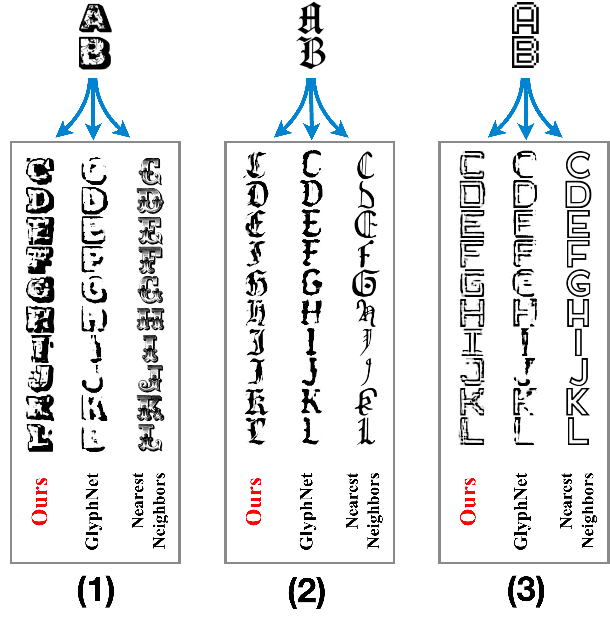
We propose a deep factorization model for typographic analysis that disentangles content from style. Specifically, a variational inference procedure factors each training glyph into the combination of a character-specific content embedding and a latent font-specific style variable. The underlying generative model combines these factors through an asymmetric transpose convolutional process to generate the image of the glyph itself. When trained on corpora of fonts, our model learns a manifold over font styles that can be used to analyze or reconstruct new, unseen fonts. On the task of reconstructing missing glyphs from an unknown font given only a small number of observations, our model outperforms both a strong nearest neighbors baseline and a state-of-the-art discriminative model from prior work.
Adaptation of a deep learning malignancy model from full-field digital mammography to digital breast tomosynthesis
Jan 23, 2020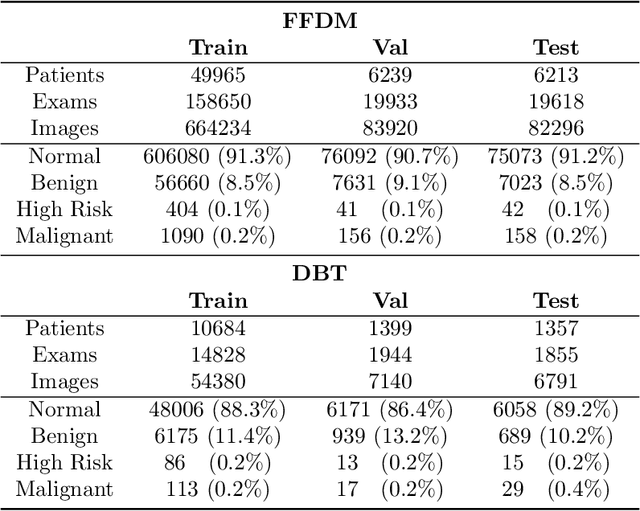
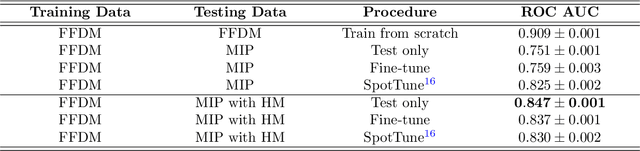
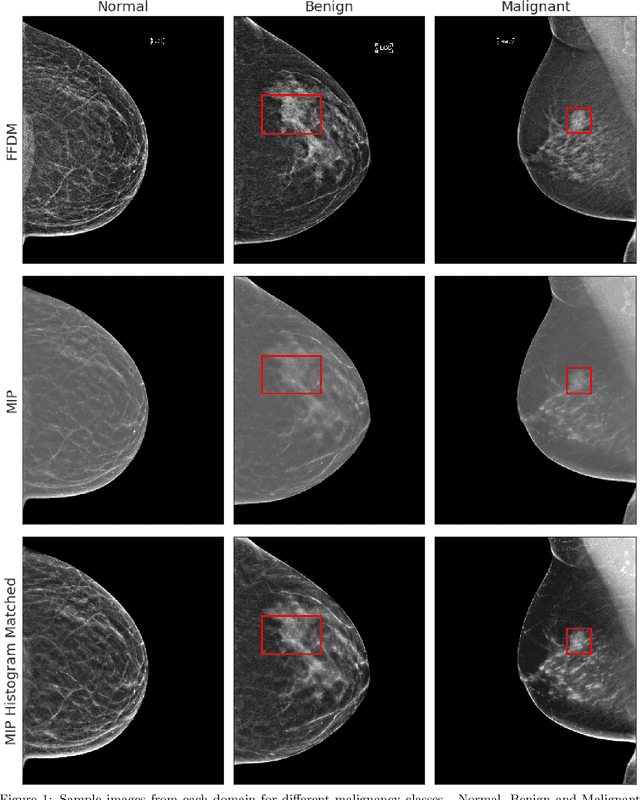
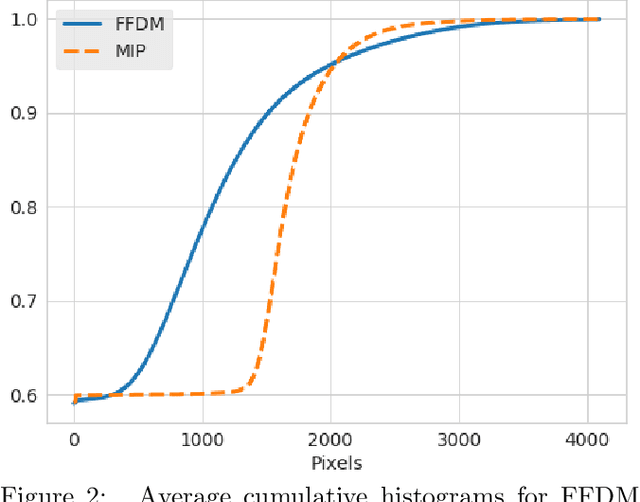

Mammography-based screening has helped reduce the breast cancer mortality rate, but has also been associated with potential harms due to low specificity, leading to unnecessary exams or procedures, and low sensitivity. Digital breast tomosynthesis (DBT) improves on conventional mammography by increasing both sensitivity and specificity and is becoming common in clinical settings. However, deep learning (DL) models have been developed mainly on conventional 2D full-field digital mammography (FFDM) or scanned film images. Due to a lack of large annotated DBT datasets, it is difficult to train a model on DBT from scratch. In this work, we present methods to generalize a model trained on FFDM images to DBT images. In particular, we use average histogram matching (HM) and DL fine-tuning methods to generalize a FFDM model to the 2D maximum intensity projection (MIP) of DBT images. In the proposed approach, the differences between the FFDM and DBT domains are reduced via HM and then the base model, which was trained on abundant FFDM images, is fine-tuned. When evaluating on image patches extracted around identified findings, we are able to achieve similar areas under the receiver operating characteristic curve (ROC AUC) of $\sim 0.9$ for FFDM and $\sim 0.85$ for MIP images, as compared to a ROC AUC of $\sim 0.75$ when tested directly on MIP images.
Scalable Nonlinear Embeddings for Semantic Category-based Image Retrieval
Sep 29, 2015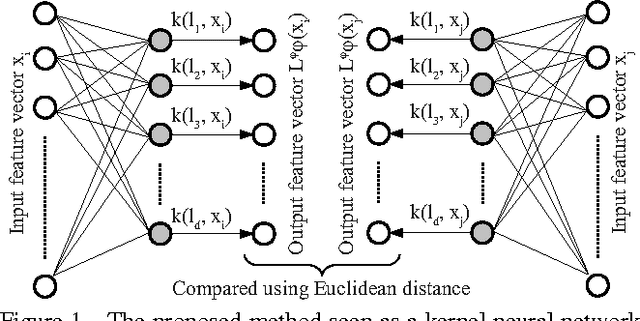
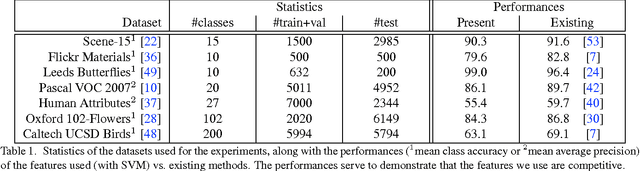
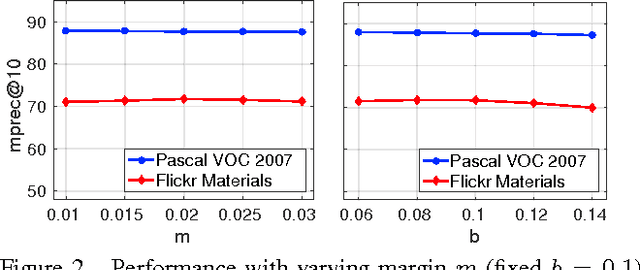
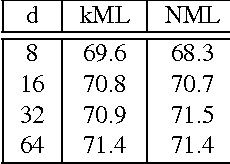
We propose a novel algorithm for the task of supervised discriminative distance learning by nonlinearly embedding vectors into a low dimensional Euclidean space. We work in the challenging setting where supervision is with constraints on similar and dissimilar pairs while training. The proposed method is derived by an approximate kernelization of a linear Mahalanobis-like distance metric learning algorithm and can also be seen as a kernel neural network. The number of model parameters and test time evaluation complexity of the proposed method are O(dD) where D is the dimensionality of the input features and d is the dimension of the projection space - this is in contrast to the usual kernelization methods as, unlike them, the complexity does not scale linearly with the number of training examples. We propose a stochastic gradient based learning algorithm which makes the method scalable (w.r.t. the number of training examples), while being nonlinear. We train the method with up to half a million training pairs of 4096 dimensional CNN features. We give empirical comparisons with relevant baselines on seven challenging datasets for the task of low dimensional semantic category based image retrieval.
Spinal Metastases Segmentation in MR Imaging using Deep Convolutional Neural Networks
Jan 08, 2020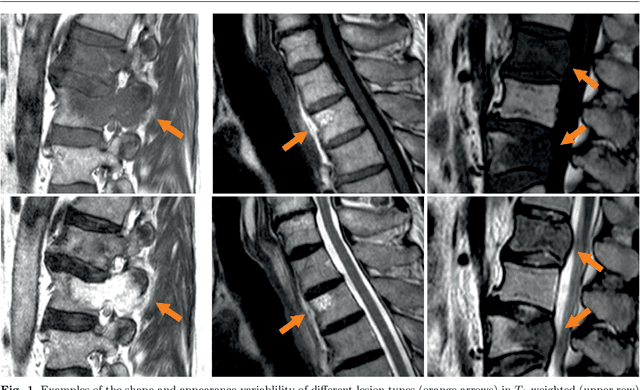
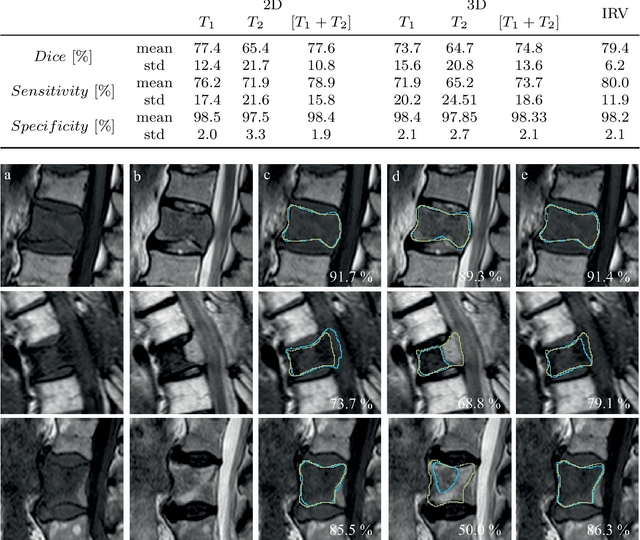
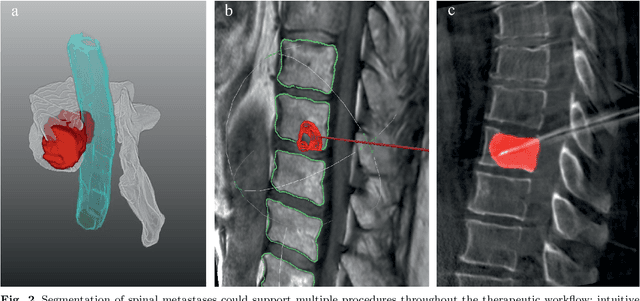
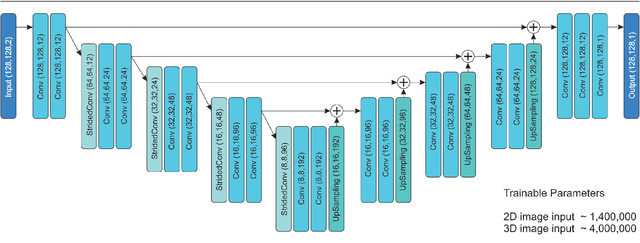

This study's objective was to segment spinal metastases in diagnostic MR images using a deep learning-based approach. Segmentation of such lesions can present a pivotal step towards enhanced therapy planning and validation, as well as intervention support during minimally invasive and image-guided surgeries like radiofrequency ablations. For this purpose, we used a U-Net like architecture trained with 40 clinical cases including both, lytic and sclerotic lesion types and various MR sequences. Our proposed method was evaluated with regards to various factors influencing the segmentation quality, e.g. the used MR sequences and the input dimension. We quantitatively assessed our experiments using Dice coefficients, sensitivity and specificity rates. Compared to expertly annotated lesion segmentations, the experiments yielded promising results with average Dice scores up to 77.6% and mean sensitivity rates up to 78.9%. To our best knowledge, our proposed study is one of the first to tackle this particular issue, which limits direct comparability with related works. In respect to similar deep learning-based lesion segmentations, e.g. in liver MR images or spinal CT images, our experiments showed similar or in some respects superior segmentation quality. Overall, our automatic approach can provide almost expert-like segmentation accuracy in this challenging and ambitious task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge