Hirid
Papers and Code
Can we generate portable representations for clinical time series data using LLMs?
Mar 25, 2026Deploying clinical ML is slow and brittle: models that work at one hospital often degrade under distribution shifts at the next. In this work, we study a simple question -- can large language models (LLMs) create portable patient embeddings i.e. representations of patients enable a downstream predictor built on one hospital to be used elsewhere with minimal-to-no retraining and fine-tuning. To do so, we map from irregular ICU time series onto concise natural language summaries using a frozen LLM, then embed each summary with a frozen text embedding model to obtain a fixed length vector capable of serving as input to a variety of downstream predictors. Across three cohorts (MIMIC-IV, HIRID, PPICU), on multiple clinically grounded forecasting and classification tasks, we find that our approach is simple, easy to use and competitive with in-distribution with grid imputation, self-supervised representation learning, and time series foundation models, while exhibiting smaller relative performance drops when transferring to new hospitals. We study the variation in performance across prompt design, with structured prompts being crucial to reducing the variance of the predictive models without altering mean accuracy. We find that using these portable representations improves few-shot learning and does not increase demographic recoverability of age or sex relative to baselines, suggesting little additional privacy risk. Our work points to the potential that LLMs hold as tools to enable the scalable deployment of production grade predictive models by reducing the engineering overhead.
Development and external validation of a multimodal artificial intelligence mortality prediction model of critically ill patients using multicenter data
Dec 15, 2025Early prediction of in-hospital mortality in critically ill patients can aid clinicians in optimizing treatment. The objective was to develop a multimodal deep learning model, using structured and unstructured clinical data, to predict in-hospital mortality risk among critically ill patients after their initial 24 hour intensive care unit (ICU) admission. We used data from MIMIC-III, MIMIC-IV, eICU, and HiRID. A multimodal model was developed on the MIMIC datasets, featuring time series components occurring within the first 24 hours of ICU admission and predicting risk of subsequent inpatient mortality. Inputs included time-invariant variables, time-variant variables, clinical notes, and chest X-ray images. External validation occurred in a temporally separated MIMIC population, HiRID, and eICU datasets. A total of 203,434 ICU admissions from more than 200 hospitals between 2001 to 2022 were included, in which mortality rate ranged from 5.2% to 7.9% across the four datasets. The model integrating structured data points had AUROC, AUPRC, and Brier scores of 0.92, 0.53, and 0.19, respectively. We externally validated the model on eight different institutions within the eICU dataset, demonstrating AUROCs ranging from 0.84-0.92. When including only patients with available clinical notes and imaging data, inclusion of notes and imaging into the model, the AUROC, AUPRC, and Brier score improved from 0.87 to 0.89, 0.43 to 0.48, and 0.37 to 0.17, respectively. Our findings highlight the importance of incorporating multiple sources of patient information for mortality prediction and the importance of external validation.
i-CardiAx: Wearable IoT-Driven System for Early Sepsis Detection Through Long-Term Vital Sign Monitoring
Jul 31, 2024



Sepsis is a significant cause of early mortality, high healthcare costs, and disability-adjusted life years. Digital interventions like continuous cardiac monitoring can help detect early warning signs and facilitate effective interventions. This paper introduces i-CardiAx, a wearable sensor utilizing low-power high-sensitivity accelerometers to measure vital signs crucial for cardiovascular health: heart rate (HR), blood pressure (BP), and respiratory rate (RR). Data collected from 10 healthy subjects using the i-CardiAx chest patch were used to develop and evaluate lightweight vital sign measurement algorithms. The algorithms demonstrated high performance: RR (-0.11 $\pm$ 0.77 breaths\min), HR (0.82 $\pm$ 2.85 beats\min), and systolic BP (-0.08 $\pm$ 6.245 mmHg). These algorithms are embedded in an ARM Cortex-M33 processor with Bluetooth Low Energy (BLE) support, achieving inference times of 4.2 ms for HR and RR, and 8.5 ms for BP. Additionally, a multi-channel quantized Temporal Convolutional Neural (TCN) Network, trained on the open-source HiRID dataset, was developed to detect sepsis onset using digitally acquired vital signs from i-CardiAx. The quantized TCN, deployed on i-CardiAx, predicted sepsis with a median time of 8.2 hours and an energy per inference of 1.29 mJ. The i-CardiAx wearable boasts a sleep power of 0.152 mW and an average power consumption of 0.77 mW, enabling a 100 mAh battery to last approximately two weeks (432 hours) with continuous monitoring of HR, BP, and RR at 30 measurements per hour and running inference every 30 minutes. In conclusion, i-CardiAx offers an energy-efficient, high-sensitivity method for long-term cardiovascular monitoring, providing predictive alerts for sepsis and other life-threatening events.
On the Importance of Step-wise Embeddings for Heterogeneous Clinical Time-Series
Nov 15, 2023



Recent advances in deep learning architectures for sequence modeling have not fully transferred to tasks handling time-series from electronic health records. In particular, in problems related to the Intensive Care Unit (ICU), the state-of-the-art remains to tackle sequence classification in a tabular manner with tree-based methods. Recent findings in deep learning for tabular data are now surpassing these classical methods by better handling the severe heterogeneity of data input features. Given the similar level of feature heterogeneity exhibited by ICU time-series and motivated by these findings, we explore these novel methods' impact on clinical sequence modeling tasks. By jointly using such advances in deep learning for tabular data, our primary objective is to underscore the importance of step-wise embeddings in time-series modeling, which remain unexplored in machine learning methods for clinical data. On a variety of clinically relevant tasks from two large-scale ICU datasets, MIMIC-III and HiRID, our work provides an exhaustive analysis of state-of-the-art methods for tabular time-series as time-step embedding models, showing overall performance improvement. In particular, we evidence the importance of feature grouping in clinical time-series, with significant performance gains when considering features within predefined semantic groups in the step-wise embedding module.
Yet Another ICU Benchmark: A Flexible Multi-Center Framework for Clinical ML
Jun 08, 2023Medical applications of machine learning (ML) have experienced a surge in popularity in recent years. The intensive care unit (ICU) is a natural habitat for ML given the abundance of available data from electronic health records. Models have been proposed to address numerous ICU prediction tasks like the early detection of complications. While authors frequently report state-of-the-art performance, it is challenging to verify claims of superiority. Datasets and code are not always published, and cohort definitions, preprocessing pipelines, and training setups are difficult to reproduce. This work introduces Yet Another ICU Benchmark (YAIB), a modular framework that allows researchers to define reproducible and comparable clinical ML experiments; we offer an end-to-end solution from cohort definition to model evaluation. The framework natively supports most open-access ICU datasets (MIMIC III/IV, eICU, HiRID, AUMCdb) and is easily adaptable to future ICU datasets. Combined with a transparent preprocessing pipeline and extensible training code for multiple ML and deep learning models, YAIB enables unified model development. Our benchmark comes with five predefined established prediction tasks (mortality, acute kidney injury, sepsis, kidney function, and length of stay) developed in collaboration with clinicians. Adding further tasks is straightforward by design. Using YAIB, we demonstrate that the choice of dataset, cohort definition, and preprocessing have a major impact on the prediction performance - often more so than model class - indicating an urgent need for YAIB as a holistic benchmarking tool. We provide our work to the clinical ML community to accelerate method development and enable real-world clinical implementations. Software Repository: https://github.com/rvandewater/YAIB.
Quantifying Health Inequalities Induced by Data and AI Models
May 03, 2022
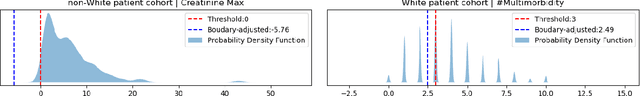
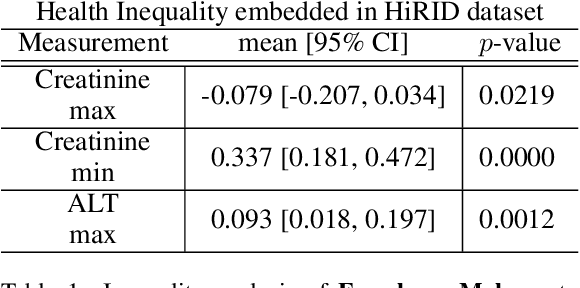

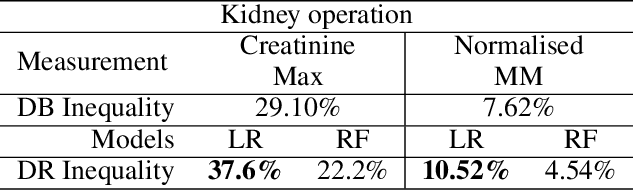
AI technologies are being increasingly tested and applied in critical environments including healthcare. Without an effective way to detect and mitigate AI induced inequalities, AI might do more harm than good, potentially leading to the widening of underlying inequalities. This paper proposes a generic allocation-deterioration framework for detecting and quantifying AI induced inequality. Specifically, AI induced inequalities are quantified as the area between two allocation-deterioration curves. To assess the framework's performance, experiments were conducted on ten synthetic datasets (N>33,000) generated from HiRID - a real-world Intensive Care Unit (ICU) dataset, showing its ability to accurately detect and quantify inequality proportionally to controlled inequalities. Extensive analyses were carried out to quantify health inequalities (a) embedded in two real-world ICU datasets; (b) induced by AI models trained for two resource allocation scenarios. Results showed that compared to men, women had up to 33% poorer deterioration in markers of prognosis when admitted to HiRID ICUs. All four AI models assessed were shown to induce significant inequalities (2.45% to 43.2%) for non-White compared to White patients. The models exacerbated data embedded inequalities significantly in 3 out of 8 assessments, one of which was >9 times worse. The codebase is at https://github.com/knowlab/DAindex-Framework.
HiRID-ICU-Benchmark -- A Comprehensive Machine Learning Benchmark on High-resolution ICU Data
Nov 18, 2021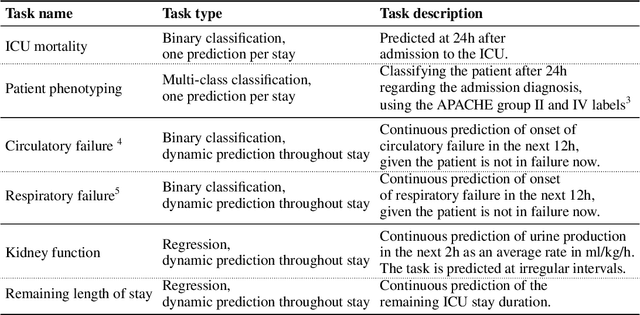
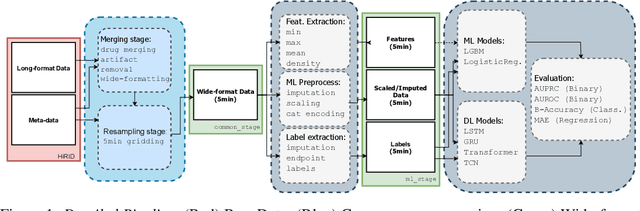
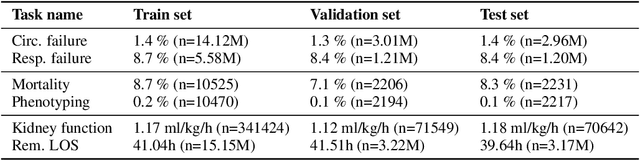
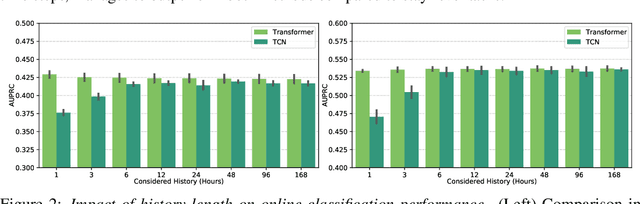
The recent success of machine learning methods applied to time series collected from Intensive Care Units (ICU) exposes the lack of standardized machine learning benchmarks for developing and comparing such methods. While raw datasets, such as MIMIC-IV or eICU, can be freely accessed on Physionet, the choice of tasks and pre-processing is often chosen ad-hoc for each publication, limiting comparability across publications. In this work, we aim to improve this situation by providing a benchmark covering a large spectrum of ICU-related tasks. Using the HiRID dataset, we define multiple clinically relevant tasks developed in collaboration with clinicians. In addition, we provide a reproducible end-to-end pipeline to construct both data and labels. Finally, we provide an in-depth analysis of current state-of-the-art sequence modeling methods, highlighting some limitations of deep learning approaches for this type of data. With this benchmark, we hope to give the research community the possibility of a fair comparison of their work.
Early prediction of respiratory failure in the intensive care unit
May 12, 2021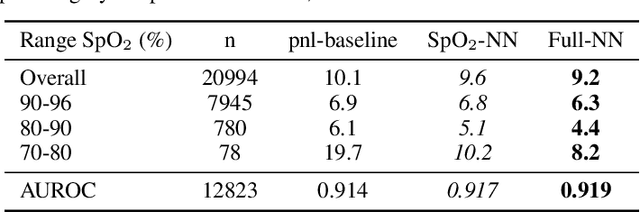
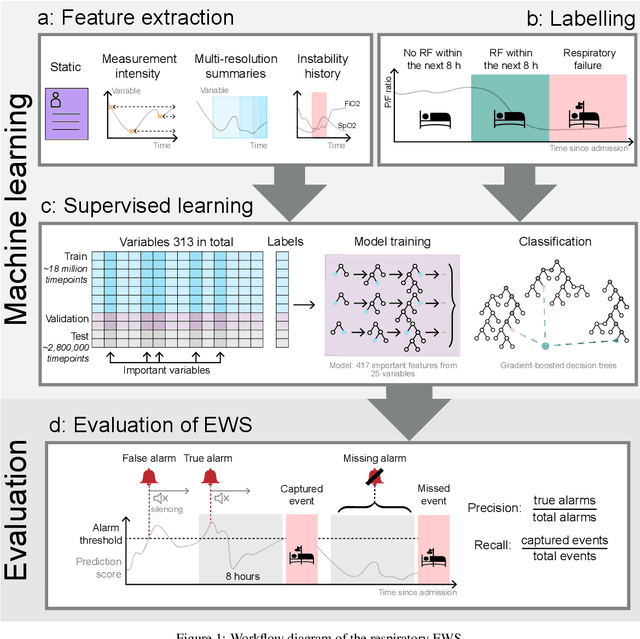
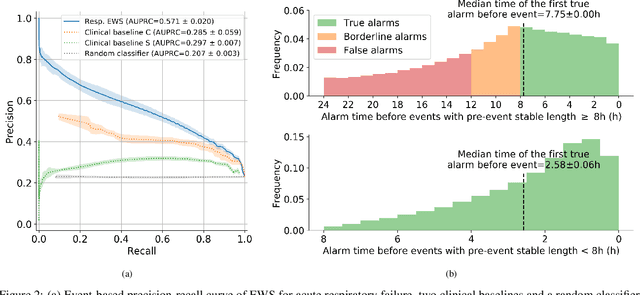
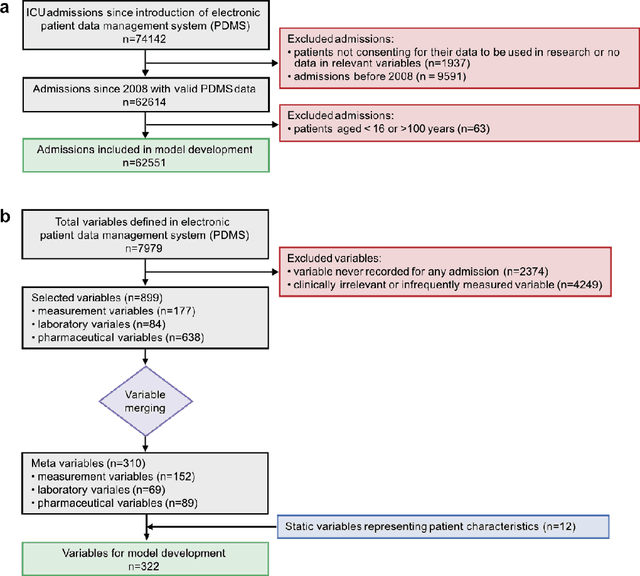
The development of respiratory failure is common among patients in intensive care units (ICU). Large data quantities from ICU patient monitoring systems make timely and comprehensive analysis by clinicians difficult but are ideal for automatic processing by machine learning algorithms. Early prediction of respiratory system failure could alert clinicians to patients at risk of respiratory failure and allow for early patient reassessment and treatment adjustment. We propose an early warning system that predicts moderate/severe respiratory failure up to 8 hours in advance. Our system was trained on HiRID-II, a data-set containing more than 60,000 admissions to a tertiary care ICU. An alarm is typically triggered several hours before the beginning of respiratory failure. Our system outperforms a clinical baseline mimicking traditional clinical decision-making based on pulse-oximetric oxygen saturation and the fraction of inspired oxygen. To provide model introspection and diagnostics, we developed an easy-to-use web browser-based system to explore model input data and predictions visually.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge