Zhiyuan Shen
GloPath: An Entity-Centric Foundation Model for Glomerular Lesion Assessment and Clinicopathological Insights
Mar 03, 2026Abstract:Glomerular pathology is central to the diagnosis and prognosis of renal diseases, yet the heterogeneity of glomerular morphology and fine-grained lesion patterns remain challenging for current AI approaches. We present GloPath, an entity-centric foundation model trained on over one million glomeruli extracted from 14,049 renal biopsy specimens using multi-scale and multi-view self-supervised learning. GloPath addresses two major challenges in nephropathology: glomerular lesion assessment and clinicopathological insights discovery. For lesion assessment, GloPath was benchmarked across three independent cohorts on 52 tasks, including lesion recognition, grading, few-shot classification, and cross-modality diagnosis-outperforming state-of-the-art methods in 42 tasks (80.8%). In the large-scale real-world study, it achieved an ROC-AUC of 91.51% for lesion recognition, demonstrating strong robustness in routine clinical settings. For clinicopathological insights, GloPath systematically revealed statistically significant associations between glomerular morphological parameters and clinical indicators across 224 morphology-clinical variable pairs, demonstrating its capacity to connect tissue-level pathology with patient-level outcomes. Together, these results position GloPath as a scalable and interpretable platform for glomerular lesion assessment and clinicopathological discovery, representing a step toward clinically translatable AI in renal pathology.
Unlocking adaptive digital pathology through dynamic feature learning
Dec 29, 2024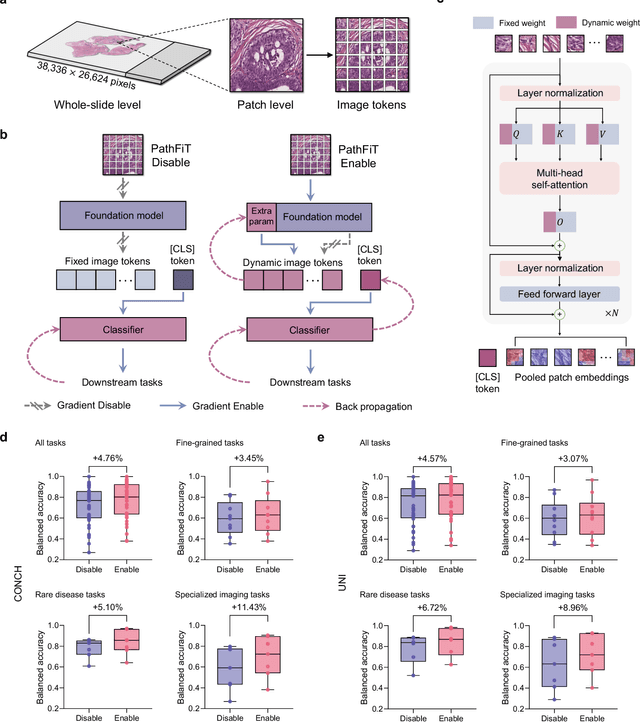
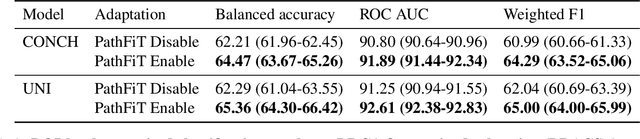
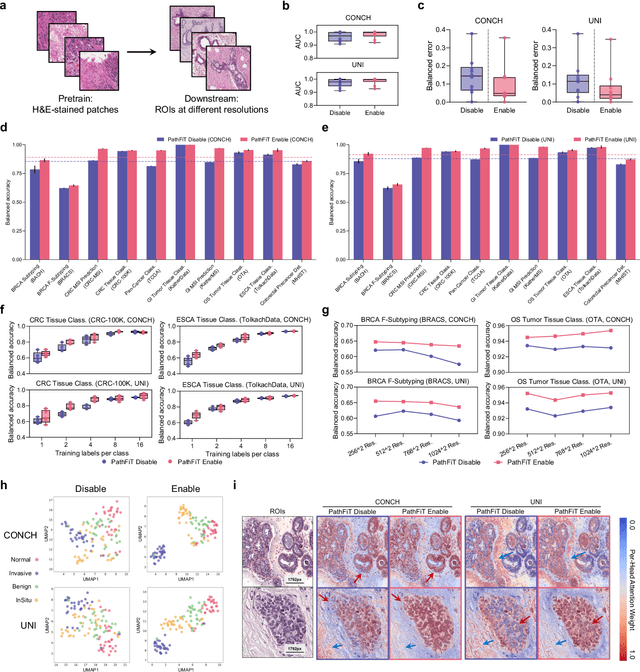

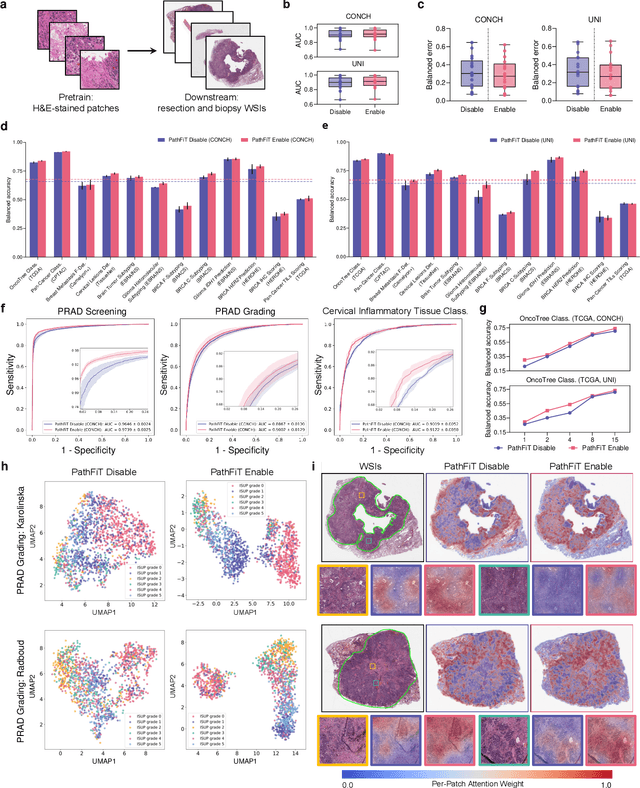
Abstract:Foundation models have revolutionized the paradigm of digital pathology, as they leverage general-purpose features to emulate real-world pathological practices, enabling the quantitative analysis of critical histological patterns and the dissection of cancer-specific signals. However, these static general features constrain the flexibility and pathological relevance in the ever-evolving needs of clinical applications, hindering the broad use of the current models. Here we introduce PathFiT, a dynamic feature learning method that can be effortlessly plugged into various pathology foundation models to unlock their adaptability. Meanwhile, PathFiT performs seamless implementation across diverse pathology applications regardless of downstream specificity. To validate PathFiT, we construct a digital pathology benchmark with over 20 terabytes of Internet and real-world data comprising 28 H\&E-stained tasks and 7 specialized imaging tasks including Masson's Trichrome staining and immunofluorescence images. By applying PathFiT to the representative pathology foundation models, we demonstrate state-of-the-art performance on 34 out of 35 tasks, with significant improvements on 23 tasks and outperforming by 10.20% on specialized imaging tasks. The superior performance and versatility of PathFiT open up new avenues in computational pathology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge