Yvan Gomez
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025


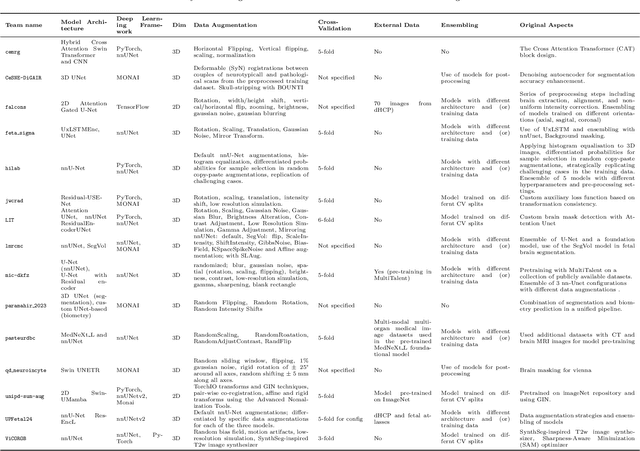
Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
FetMRQC: an open-source machine learning framework for multi-centric fetal brain MRI quality control
Nov 08, 2023



Abstract:Fetal brain MRI is becoming an increasingly relevant complement to neurosonography for perinatal diagnosis, allowing fundamental insights into fetal brain development throughout gestation. However, uncontrolled fetal motion and heterogeneity in acquisition protocols lead to data of variable quality, potentially biasing the outcome of subsequent studies. We present FetMRQC, an open-source machine-learning framework for automated image quality assessment and quality control that is robust to domain shifts induced by the heterogeneity of clinical data. FetMRQC extracts an ensemble of quality metrics from unprocessed anatomical MRI and combines them to predict experts' ratings using random forests. We validate our framework on a pioneeringly large and diverse dataset of more than 1600 manually rated fetal brain T2-weighted images from four clinical centers and 13 different scanners. Our study shows that FetMRQC's predictions generalize well to unseen data while being interpretable. FetMRQC is a step towards more robust fetal brain neuroimaging, which has the potential to shed new insights on the developing human brain.
FetMRQC: Automated Quality Control for fetal brain MRI
Apr 12, 2023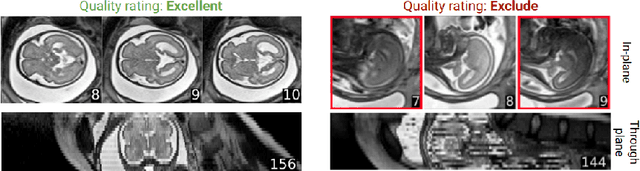
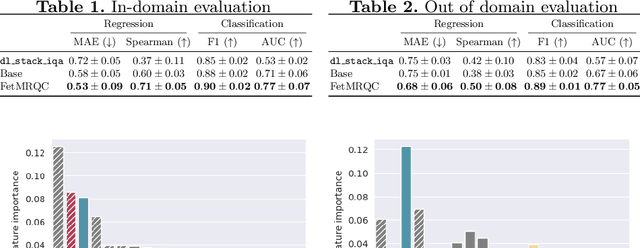
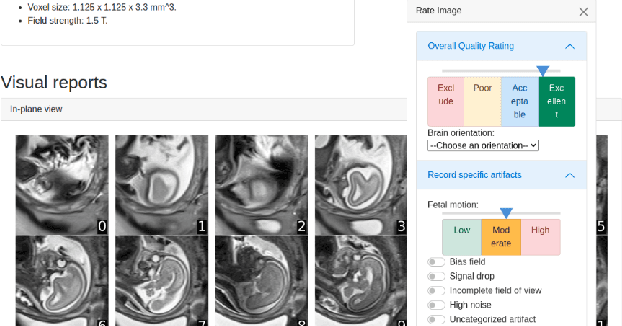
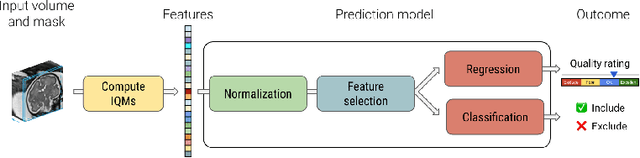
Abstract:Quality control (QC) has long been considered essential to guarantee the reliability of neuroimaging studies. It is particularly important for fetal brain MRI, where large and unpredictable fetal motion can lead to substantial artifacts in the acquired images. Existing methods for fetal brain quality assessment operate at the \textit{slice} level, and fail to get a comprehensive picture of the quality of an image, that can only be achieved by looking at the \textit{entire} brain volume. In this work, we propose FetMRQC, a machine learning framework for automated image quality assessment tailored to fetal brain MRI, which extracts an ensemble of quality metrics that are then used to predict experts' ratings. Based on the manual ratings of more than 1000 low-resolution stacks acquired across two different institutions, we show that, compared with existing quality metrics, FetMRQC is able to generalize out-of-domain, while being interpretable and data efficient. We also release a novel manual quality rating tool designed to facilitate and optimize quality rating of fetal brain images. Our tool, along with all the code to generate, train and evaluate the model will be released upon acceptance of the paper.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge