Yu-Ping Wang
A new approach to extracting coronary arteries and detecting stenosis in invasive coronary angiograms
Jan 25, 2021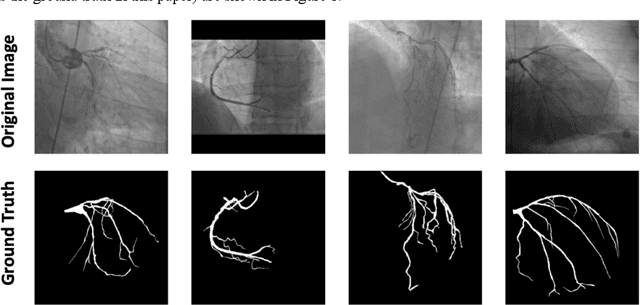

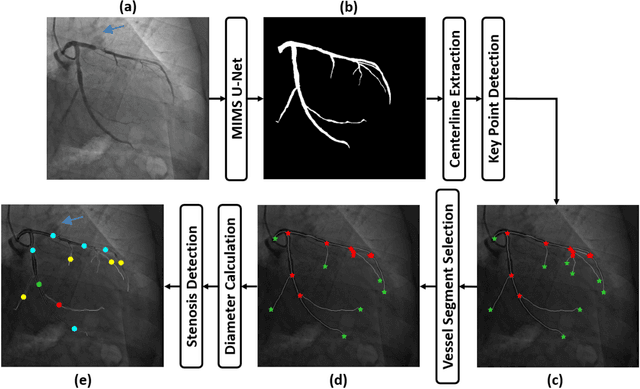
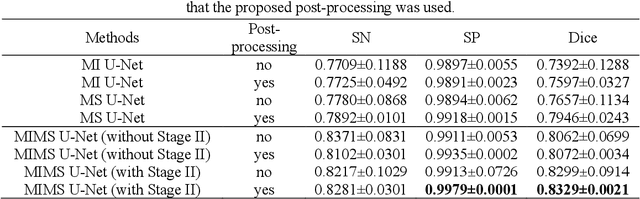
Abstract:In stable coronary artery disease (CAD), reduction in mortality and/or myocardial infarction with revascularization over medical therapy has not been reliably achieved. Coronary arteries are usually extracted to perform stenosis detection. We aim to develop an automatic algorithm by deep learning to extract coronary arteries from ICAs.In this study, a multi-input and multi-scale (MIMS) U-Net with a two-stage recurrent training strategy was proposed for the automatic vessel segmentation. Incorporating features such as the Inception residual module with depth-wise separable convolutional layers, the proposed model generated a refined prediction map with the following two training stages: (i) Stage I coarsely segmented the major coronary arteries from pre-processed single-channel ICAs and generated the probability map of vessels; (ii) during the Stage II, a three-channel image consisting of the original preprocessed image, a generated probability map, and an edge-enhanced image generated from the preprocessed image was fed to the proposed MIMS U-Net to produce the final segmentation probability map. During the training stage, the probability maps were iteratively and recurrently updated by feeding into the neural network. After segmentation, an arterial stenosis detection algorithm was developed to extract vascular centerlines and calculate arterial diameters to evaluate stenotic level. Experimental results demonstrated that the proposed method achieved an average Dice score of 0.8329, an average sensitivity of 0.8281, and an average specificity of 0.9979 in our dataset with 294 ICAs obtained from 73 patient. Moreover, our stenosis detection algorithm achieved a true positive rate of 0.6668 and a positive predictive value of 0.7043.
Ensemble manifold based regularized multi-modal graph convolutional network for cognitive ability prediction
Jan 20, 2021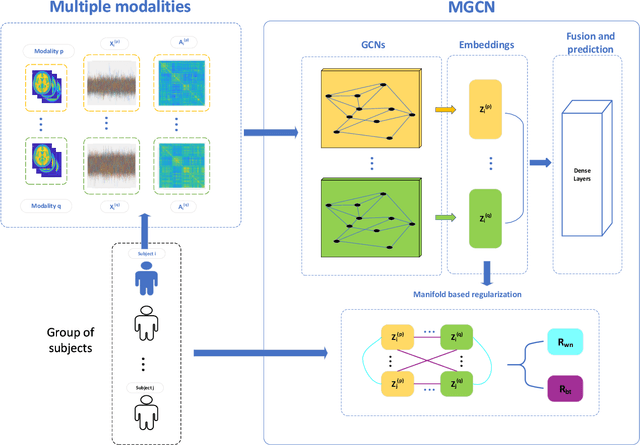
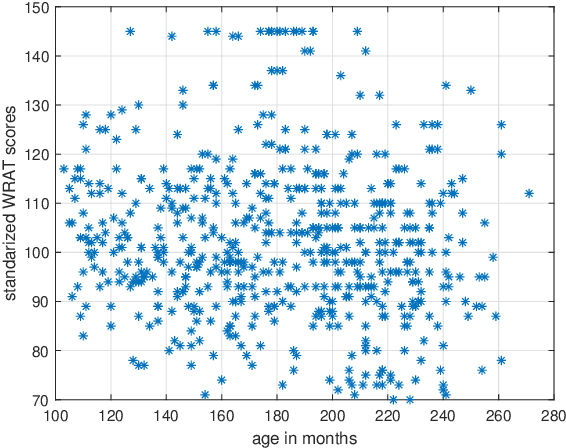
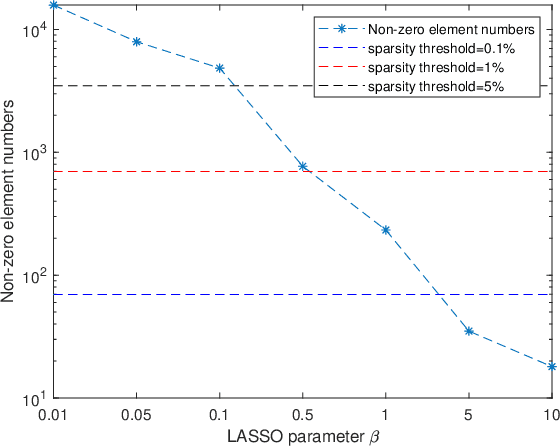
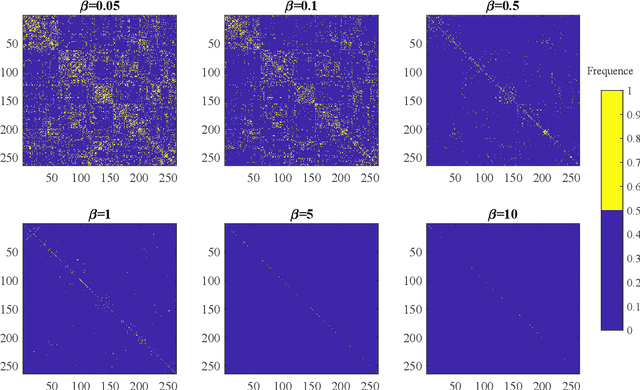
Abstract:Objective: Multi-modal functional magnetic resonance imaging (fMRI) can be used to make predictions about individual behavioral and cognitive traits based on brain connectivity networks. Methods: To take advantage of complementary information from multi-modal fMRI, we propose an interpretable multi-modal graph convolutional network (MGCN) model, incorporating the fMRI time series and the functional connectivity (FC) between each pair of brain regions. Specifically, our model learns a graph embedding from individual brain networks derived from multi-modal data. A manifold-based regularization term is then enforced to consider the relationships of subjects both within and between modalities. Furthermore, we propose the gradient-weighted regression activation mapping (Grad-RAM) and the edge mask learning to interpret the model, which is used to identify significant cognition-related biomarkers. Results: We validate our MGCN model on the Philadelphia Neurodevelopmental Cohort to predict individual wide range achievement test (WRAT) score. Our model obtains superior predictive performance over GCN with a single modality and other competing approaches. The identified biomarkers are cross-validated from different approaches. Conclusion and Significance: This paper develops a new interpretable graph deep learning framework for cognitive ability prediction, with the potential to overcome the limitations of several current data-fusion models. The results demonstrate the power of MGCN in analyzing multi-modal fMRI and discovering significant biomarkers for human brain studies.
ORBBuf: A Robust Buffering Method for Collaborative Visual SLAM
Oct 28, 2020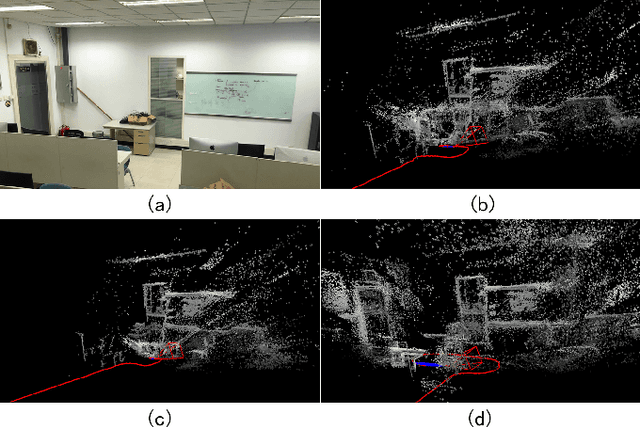
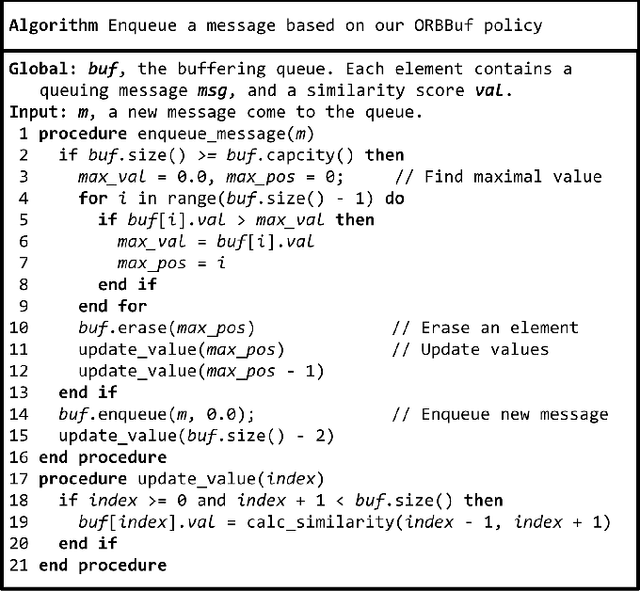

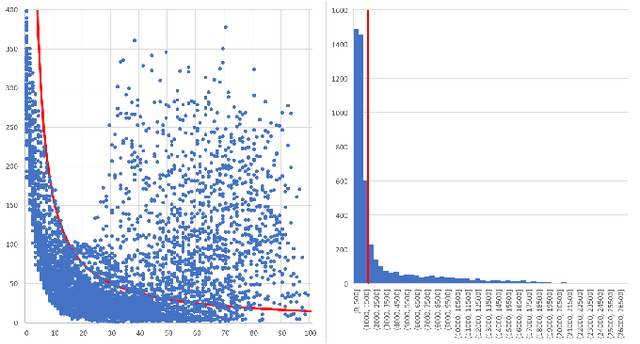
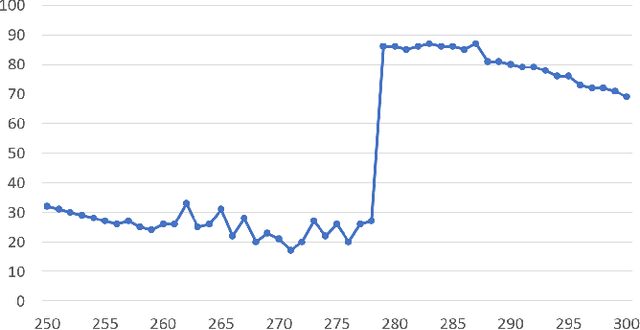
Abstract:Collaborative simultaneous localization and mapping (SLAM) approaches provide a solution for autonomous robots based on embedded devices. On the other hand, visual SLAM systems rely on correlations between visual frames. As a result, the loss of visual frames from an unreliable wireless network can easily damage the results of collaborative visual SLAM systems. From our experiment, a loss of less than 1 second of data can lead to the failure of visual SLAM algorithms. We present a novel buffering method, ORBBuf, to reduce the impact of data loss on collaborative visual SLAM systems. We model the buffering problem into an optimization problem. We use an efficient greedy-like algorithm, and our buffering method drops the frame that results in the least loss to the quality of the SLAM results. We implement our ORBBuf method on ROS, a widely used middleware framework. Through an extensive evaluation on real-world scenarios and tens of gigabytes of datasets, we demonstrate that our ORBBuf method can be applied to different algorithms, different sensor data (both monocular images and stereo images), different scenes (both indoor and outdoor), and different network environments (both WiFi networks and 4G networks). Experimental results show that the network interruptions indeed affect the SLAM results, and our ORBBuf method can reduce the RMSE up to 50 times.
Distance Correlation Based Brain Functional Connectivity Estimation and Non-Convex Multi-Task Learning for Developmental fMRI Studies
Sep 30, 2020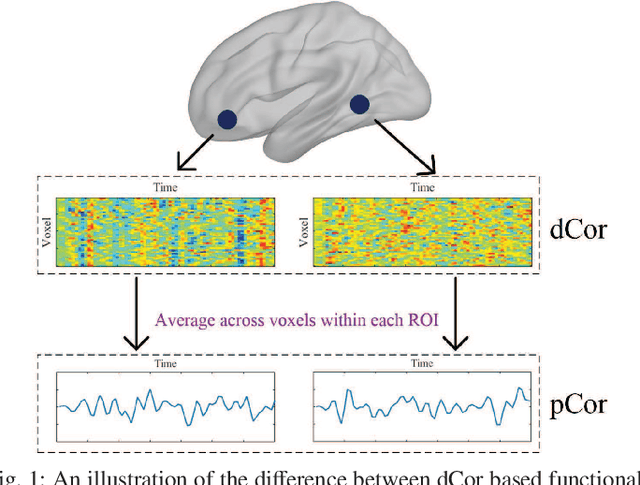
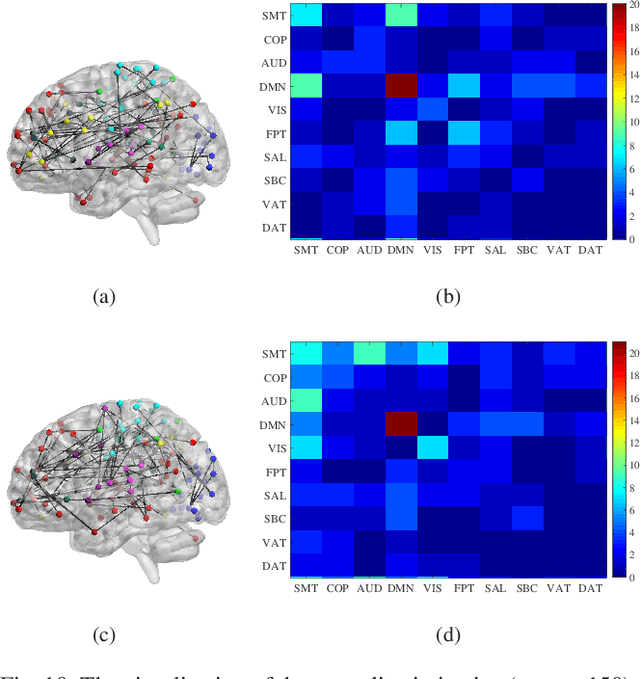
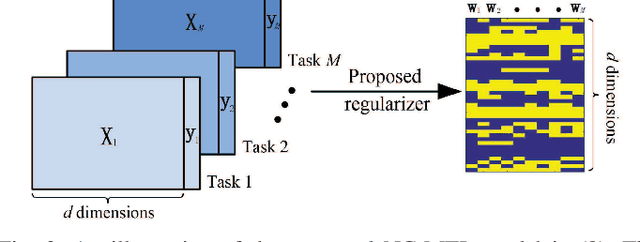
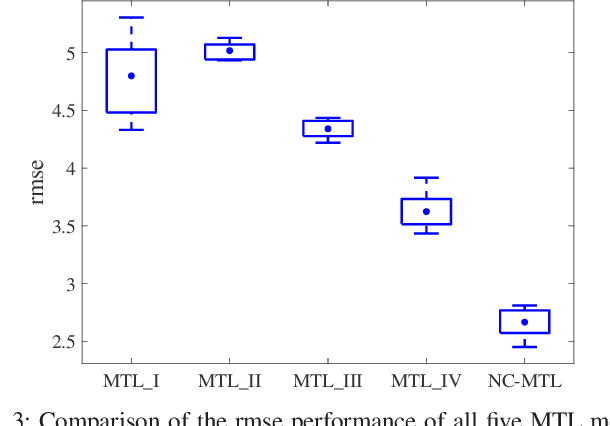
Abstract:Resting-state functional magnetic resonance imaging (rs-fMRI)-derived functional connectivity patterns have been extensively utilized to delineate global functional organization of the human brain in health, development, and neuropsychiatric disorders. In this paper, we investigate how functional connectivity in males and females differs in an age prediction framework. We first estimate functional connectivity between regions-of-interest (ROIs) using distance correlation instead of Pearson's correlation. Distance correlation, as a multivariate statistical method, explores spatial relations of voxel-wise time courses within individual ROIs and measures both linear and nonlinear dependence, capturing more complex information of between-ROI interactions. Then, a novel non-convex multi-task learning (NC-MTL) model is proposed to study age-related gender differences in functional connectivity, where age prediction for each gender group is viewed as one task. Specifically, in the proposed NC-MTL model, we introduce a composite regularizer with a combination of non-convex $\ell_{2,1-2}$ and $\ell_{1-2}$ regularization terms for selecting both common and task-specific features. Finally, we validate the proposed NC-MTL model along with distance correlation based functional connectivity on rs-fMRI of the Philadelphia Neurodevelopmental Cohort for predicting ages of both genders. The experimental results demonstrate that the proposed NC-MTL model outperforms other competing MTL models in age prediction, as well as characterizing developmental gender differences in functional connectivity patterns.
Causal inference of brain connectivity from fMRI with $ψ$-Learning Incorporated Linear non-Gaussian Acyclic Model ($ψ$-LiNGAM)
Jun 16, 2020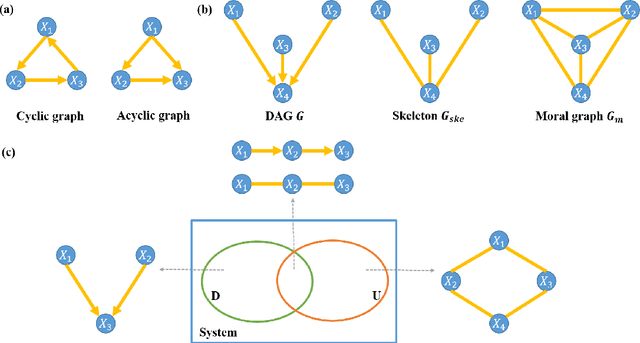
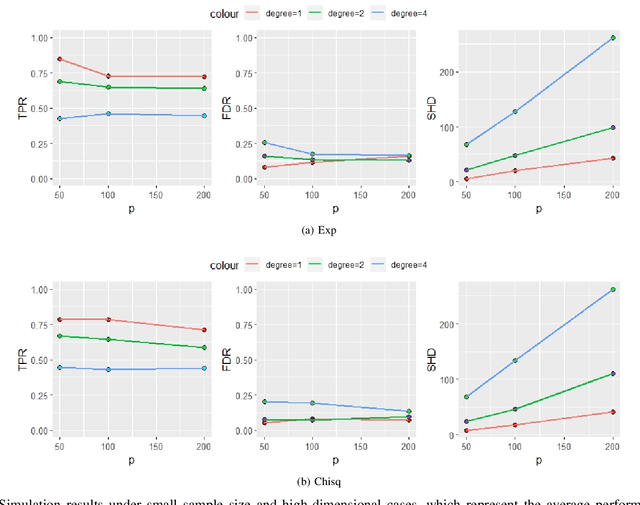
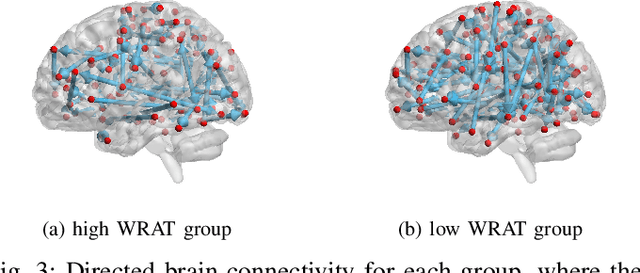
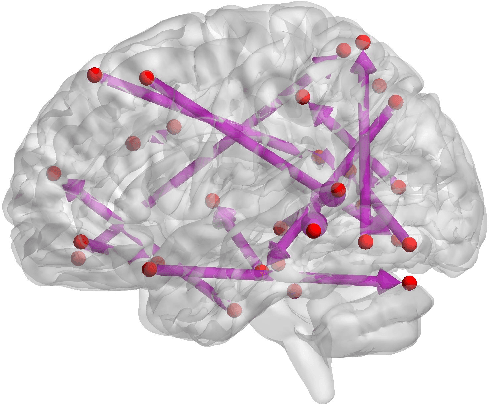
Abstract:Functional connectivity (FC) has become a primary means of understanding brain functions by identifying brain network interactions and, ultimately, how those interactions produce cognitions. A popular definition of FC is by statistical associations between measured brain regions. However, this could be problematic since the associations can only provide spatial connections but not causal interactions among regions of interests. Hence, it is necessary to study their causal relationship. Directed acyclic graph (DAG) models have been applied in recent FC studies but often encountered problems such as limited sample sizes and large number of variables (namely high-dimensional problems), which lead to both computational difficulty and convergence issues. As a result, the use of DAG models is problematic, where the identification of DAG models in general is nondeterministic polynomial time hard (NP-hard). To this end, we propose a $\psi$-learning incorporated linear non-Gaussian acyclic model ($\psi$-LiNGAM). We use the association model ($\psi$-learning) to facilitate causal inferences and the model works well especially for high-dimensional cases. Our simulation results demonstrate that the proposed method is more robust and accurate than several existing ones in detecting graph structure and direction. We then applied it to the resting state fMRI (rsfMRI) data obtained from the publicly available Philadelphia Neurodevelopmental Cohort (PNC) to study the cognitive variance, which includes 855 individuals aged 8-22 years. Therein, we have identified three types of hub structure: the in-hub, out-hub and sum-hub, which correspond to the centers of receiving, sending and relaying information, respectively. We also detected 16 most important pairs of causal flows. Several of the results have been verified to be biologically significant.
A Bayesian incorporated linear non-Gaussian acyclic model for multiple directed graph estimation to study brain emotion circuit development in adolescence
Jun 16, 2020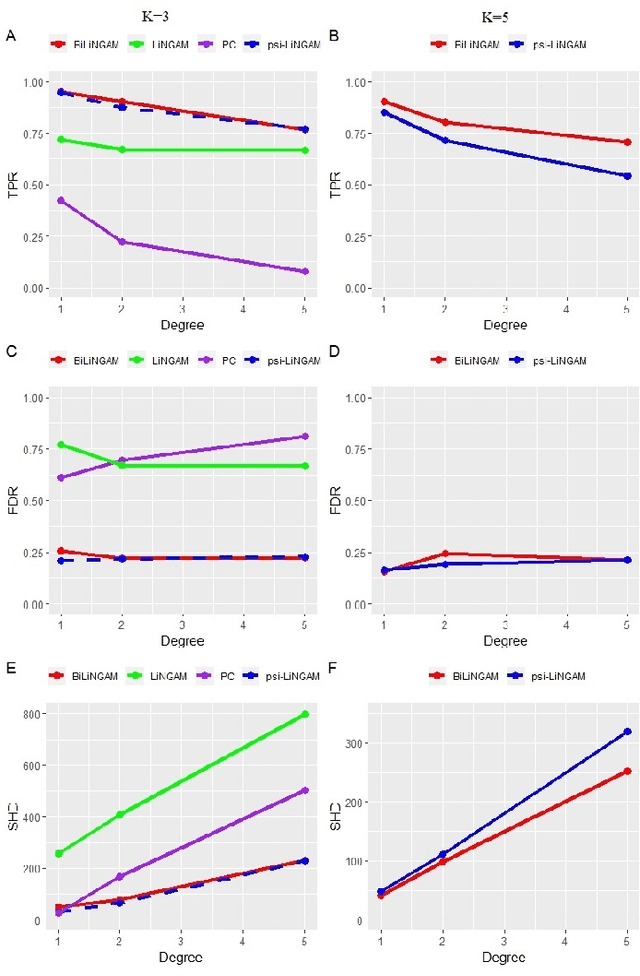
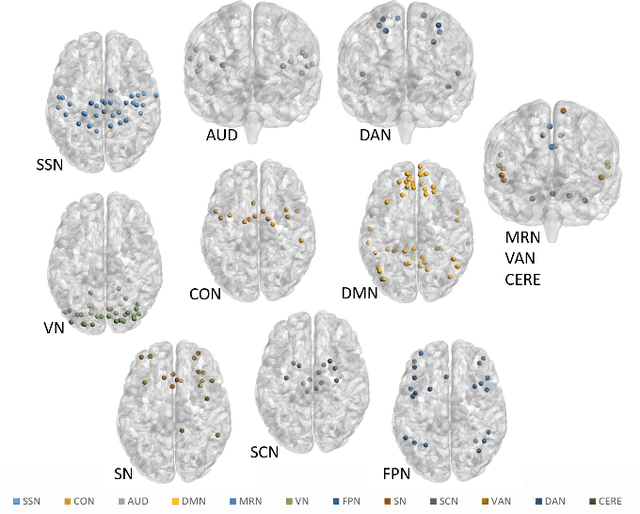
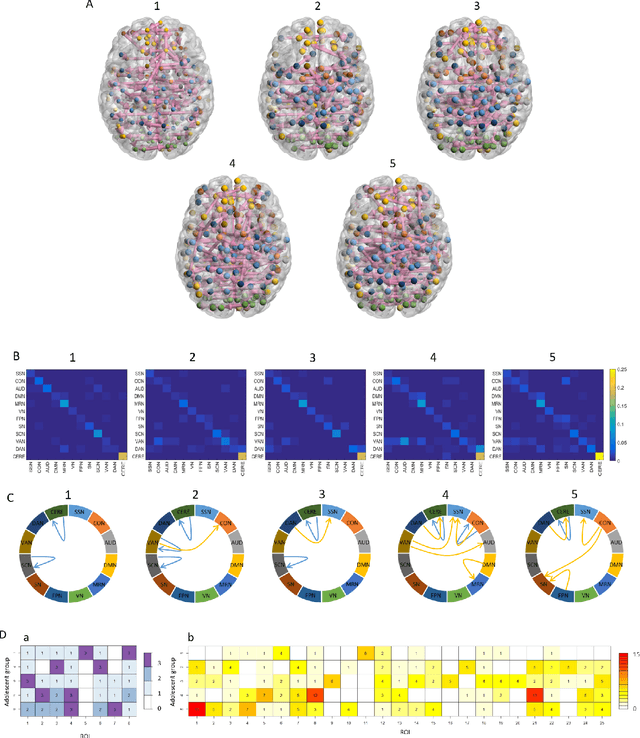
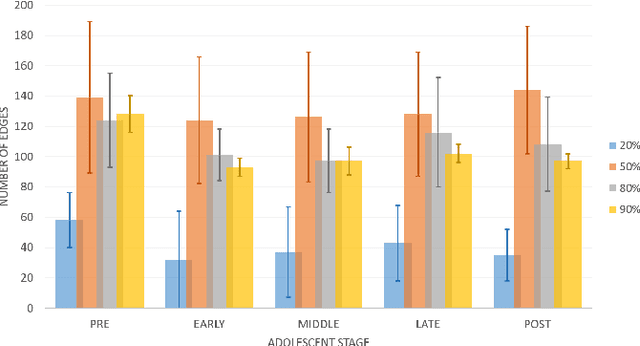
Abstract:Emotion perception is essential to affective and cognitive development which involves distributed brain circuits. The ability of emotion identification begins in infancy and continues to develop throughout childhood and adolescence. Understanding the development of brain's emotion circuitry may help us explain the emotional changes observed during adolescence. Our previous study delineated the trajectory of brain functional connectivity (FC) from late childhood to early adulthood during emotion identification tasks. In this work, we endeavour to deepen our understanding from association to causation. We proposed a Bayesian incorporated linear non-Gaussian acyclic model (BiLiNGAM), which incorporated our previous association model into the prior estimation pipeline. In particular, it can jointly estimate multiple directed acyclic graphs (DAGs) for multiple age groups at different developmental stages. Simulation results indicated more stable and accurate performance over various settings, especially when the sample size was small (high-dimensional cases). We then applied to the analysis of real data from the Philadelphia Neurodevelopmental Cohort (PNC). This included 855 individuals aged 8-22 years who were divided into five different adolescent stages. Our network analysis revealed the development of emotion-related intra- and inter- modular connectivity and pinpointed several emotion-related hubs. We further categorized the hubs into two types: in-hubs and out-hubs, as the center of receiving and distributing information. Several unique developmental hub structures and group-specific patterns were also discovered. Our findings help provide a causal understanding of emotion development in the human brain.
Interpretable multimodal fusion networks reveal mechanisms of brain cognition
Jun 16, 2020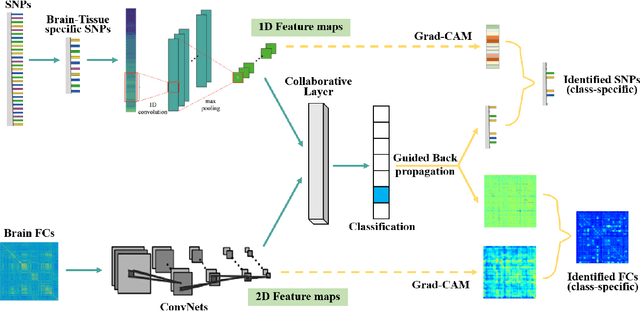
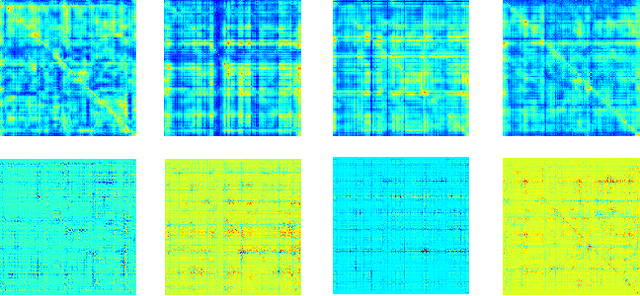
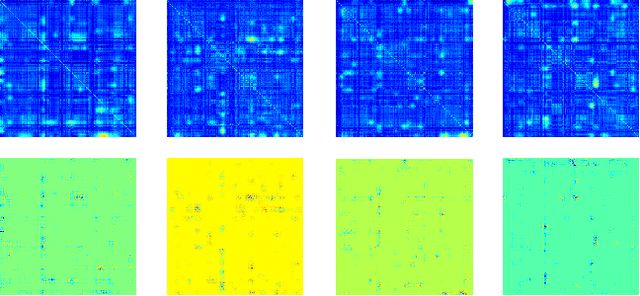
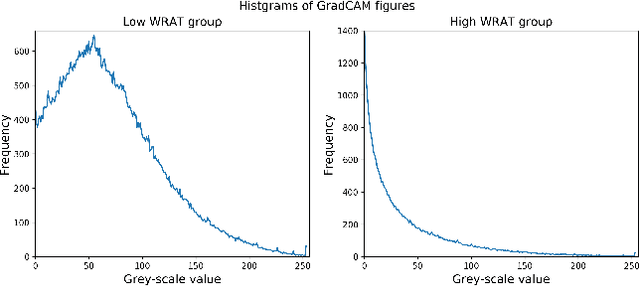
Abstract:Multimodal fusion benefits disease diagnosis by providing a more comprehensive perspective. Developing algorithms is challenging due to data heterogeneity and the complex within- and between-modality associations. Deep-network-based data-fusion models have been developed to capture the complex associations and the performance in diagnosis has been improved accordingly. Moving beyond diagnosis prediction, evaluation of disease mechanisms is critically important for biomedical research. Deep-network-based data-fusion models, however, are difficult to interpret, bringing about difficulties for studying biological mechanisms. In this work, we develop an interpretable multimodal fusion model, namely gCAM-CCL, which can perform automated diagnosis and result interpretation simultaneously. The gCAM-CCL model can generate interpretable activation maps, which quantify pixel-level contributions of the input features. This is achieved by combining intermediate feature maps using gradient-based weights. Moreover, the estimated activation maps are class-specific, and the captured cross-data associations are interest/label related, which further facilitates class-specific analysis and biological mechanism analysis. We validate the gCAM-CCL model on a brain imaging-genetic study, and show gCAM-CCL's performed well for both classification and mechanism analysis. Mechanism analysis suggests that during task-fMRI scans, several object recognition related regions of interests (ROIs) are first activated and then several downstream encoding ROIs get involved. Results also suggest that the higher cognition performing group may have stronger neurotransmission signaling while the lower cognition performing group may have problem in brain/neuron development, resulting from genetic variations.
A generalized kernel machine approach to identify higher-order composite effects in multi-view datasets
Apr 29, 2020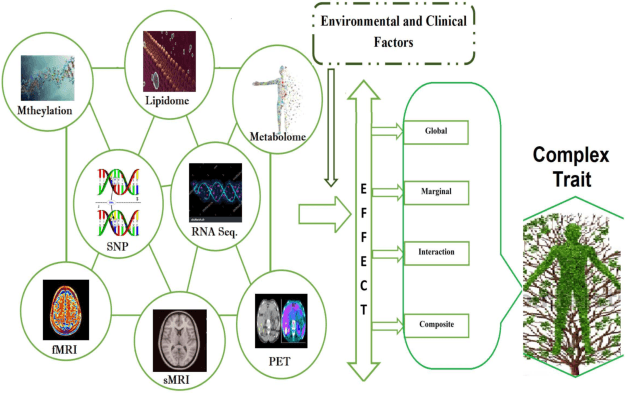
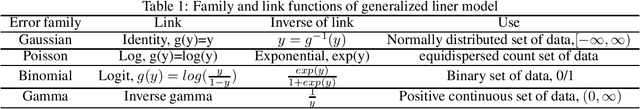

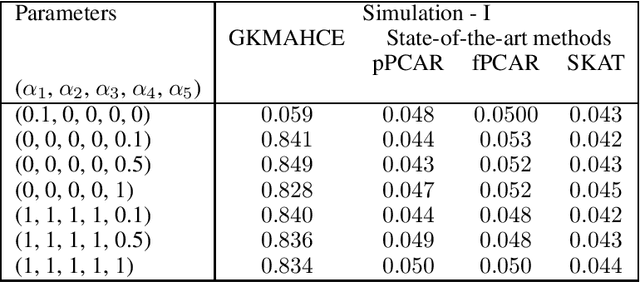
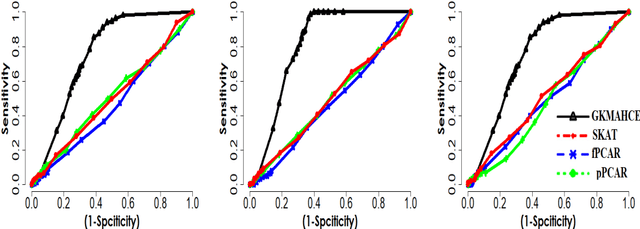
Abstract:In recent years, a comprehensive study of multi-view datasets (e.g., multi-omics and imaging scans) has been a focus and forefront in biomedical research. State-of-the-art biomedical technologies are enabling us to collect multi-view biomedical datasets for the study of complex diseases. While all the views of data tend to explore complementary information of a disease, multi-view data analysis with complex interactions is challenging for a deeper and holistic understanding of biological systems. In this paper, we propose a novel generalized kernel machine approach to identify higher-order composite effects in multi-view biomedical datasets. This generalized semi-parametric (a mixed-effect linear model) approach includes the marginal and joint Hadamard product of features from different views of data. The proposed kernel machine approach considers multi-view data as predictor variables to allow more thorough and comprehensive modeling of a complex trait. The proposed method can be applied to the study of any disease model, where multi-view datasets are available. We applied our approach to both synthesized datasets and real multi-view datasets from adolescence brain development and osteoporosis study, including an imaging scan dataset and five omics datasets. Our experiments demonstrate that the proposed method can effectively identify higher-order composite effects and suggest that corresponding features (genes, region of interests, and chemical taxonomies) function in a concerted effort. We show that the proposed method is more generalizable than existing ones.
OptSample: A Resilient Buffer Management Policy for Robotic Systems based on Optimal Message Sampling
Sep 26, 2019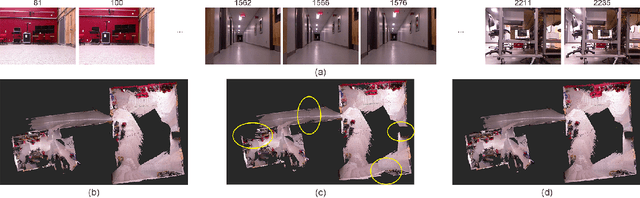
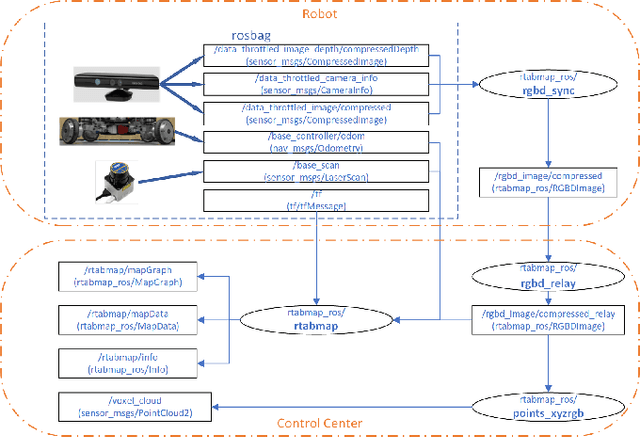
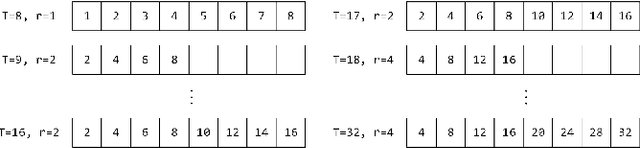
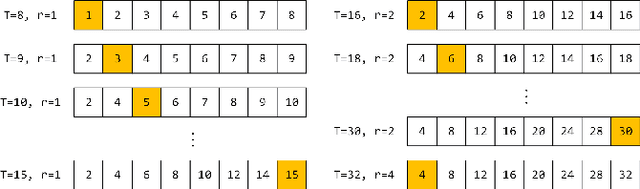
Abstract:Modern robotic systems have become an alternative to humans to perform risky or exhausting tasks. In such application scenarios, communications between robots and the control center have become one of the major problems. Buffering is a commonly used solution to relieve temporary network disruption. But the assumption that newer messages are more valuable than older ones is not true for many application scenarios such as explorations, rescue operations, and surveillance. In this paper, we proposed a novel resilient buffer management policy named OptSample. It can uniformly sampling messages and dynamically adjust the sample rate based on run-time network situation. We define an evaluation function to estimate the profit of a message sequence. Based on the function, our analysis and simulation shows that the OptSample policy can effectively prevent losing long segment of continuous messages and improve the overall profit of the received messages. We implement the proposed policy in ROS. The implementation is transparent to user and no user code need to be changed. Experimental results on several application scenarios show that the OptSample policy can help robotic systems be more resilient against network disruption.
Multimodal Sparse Classifier for Adolescent Brain Age Prediction
Apr 01, 2019
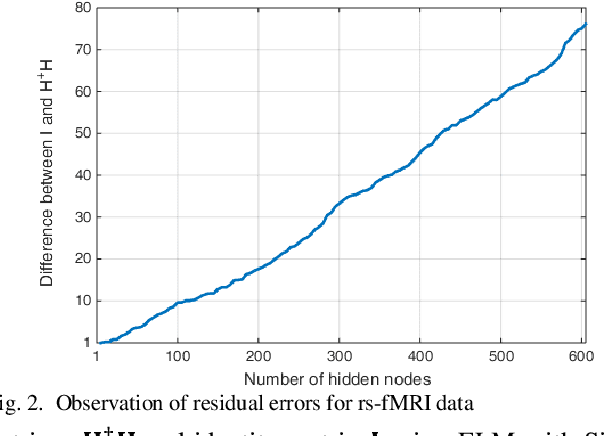
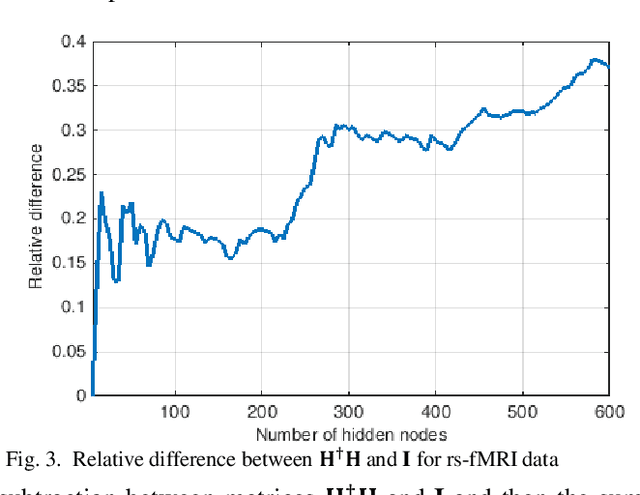
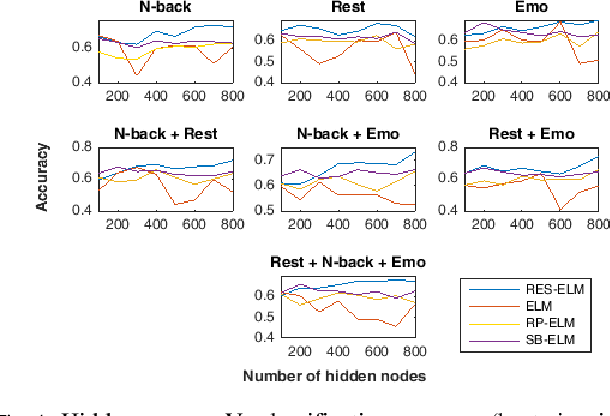
Abstract:The study of healthy brain development helps to better understand the brain transformation and brain connectivity patterns which happen during childhood to adulthood. This study presents a sparse machine learning solution across whole-brain functional connectivity (FC) measures of three sets of data, derived from resting state functional magnetic resonance imaging (rs-fMRI) and task fMRI data, including a working memory n-back task (nb-fMRI) and an emotion identification task (em-fMRI). These multi-modal image data are collected on a sample of adolescents from the Philadelphia Neurodevelopmental Cohort (PNC) for the prediction of brain ages. Due to extremely large variable-to-instance ratio of PNC data, a high dimensional matrix with several irrelevant and highly correlated features is generated and hence a pattern learning approach is necessary to extract significant features. We propose a sparse learner based on the residual errors along the estimation of an inverse problem for the extreme learning machine (ELM) neural network. The purpose of the approach is to overcome the overlearning problem through pruning of several redundant features and their corresponding output weights. The proposed multimodal sparse ELM classifier based on residual errors (RES-ELM) is highly competitive in terms of the classification accuracy compared to its counterparts such as conventional ELM, and sparse Bayesian learning ELM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge