Trieu Nguyen
MMORF: A Multi-agent Framework for Designing Multi-objective Retrosynthesis Planning Systems
Apr 06, 2026Abstract:Multi-objective retrosynthesis planning is a critical chemistry task requiring dynamic balancing of quality, safety, and cost objectives. Language model-based multi-agent systems (MAS) offer a promising approach for this task: leveraging interactions of specialized agents to incorporate multiple objectives into retrosynthesis planning. We present MMORF, a framework for constructing MAS for multi-objective retrosynthesis planning. MMORF features modular agentic components, which can be flexibly combined and configured into different systems, enabling principled evaluation and comparison of different system designs. Using MMORF, we construct two representative MAS: MASIL and RFAS. On a newly curated benchmark consisting of 218 multi-objective retrosynthesis planning tasks, MASIL achieves strong safety and cost metrics on soft-constraint tasks, frequently Pareto-dominating baseline routes, while RFAS achieves a 48.6% success rate on hard-constraint tasks, outperforming state-of-the-art baselines. Together, these results show the effectiveness of MMORF as a foundational framework for exploring MAS for multi-objective retrosynthesis planning. Code and data are available at https://anonymous.4open.science/r/MMORF/.
PepThink-R1: LLM for Interpretable Cyclic Peptide Optimization with CoT SFT and Reinforcement Learning
Aug 20, 2025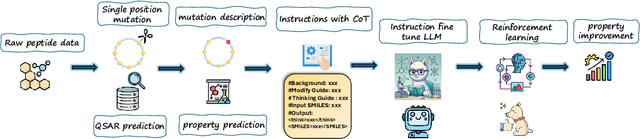
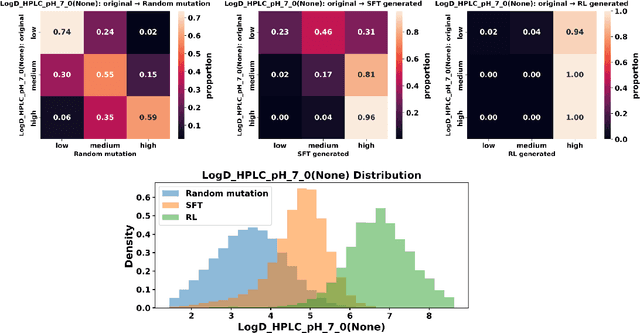
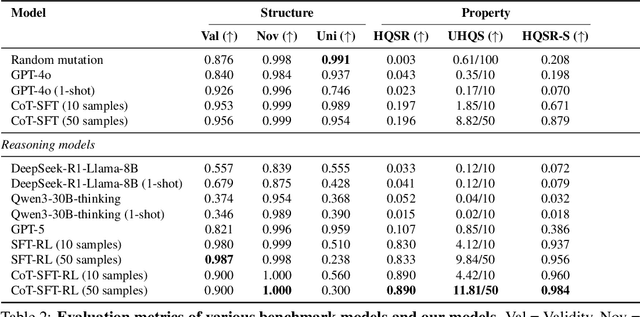
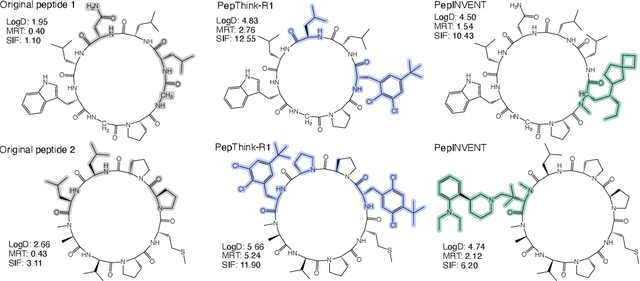
Abstract:Designing therapeutic peptides with tailored properties is hindered by the vastness of sequence space, limited experimental data, and poor interpretability of current generative models. To address these challenges, we introduce PepThink-R1, a generative framework that integrates large language models (LLMs) with chain-of-thought (CoT) supervised fine-tuning and reinforcement learning (RL). Unlike prior approaches, PepThink-R1 explicitly reasons about monomer-level modifications during sequence generation, enabling interpretable design choices while optimizing for multiple pharmacological properties. Guided by a tailored reward function balancing chemical validity and property improvements, the model autonomously explores diverse sequence variants. We demonstrate that PepThink-R1 generates cyclic peptides with significantly enhanced lipophilicity, stability, and exposure, outperforming existing general LLMs (e.g., GPT-5) and domain-specific baseline in both optimization success and interpretability. To our knowledge, this is the first LLM-based peptide design framework that combines explicit reasoning with RL-driven property control, marking a step toward reliable and transparent peptide optimization for therapeutic discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge