Thomas Lukasiewicz
Robust Graph Representation Learning via Predictive Coding
Dec 09, 2022Abstract:Predictive coding is a message-passing framework initially developed to model information processing in the brain, and now also topic of research in machine learning due to some interesting properties. One of such properties is the natural ability of generative models to learn robust representations thanks to their peculiar credit assignment rule, that allows neural activities to converge to a solution before updating the synaptic weights. Graph neural networks are also message-passing models, which have recently shown outstanding results in diverse types of tasks in machine learning, providing interdisciplinary state-of-the-art performance on structured data. However, they are vulnerable to imperceptible adversarial attacks, and unfit for out-of-distribution generalization. In this work, we address this by building models that have the same structure of popular graph neural network architectures, but rely on the message-passing rule of predictive coding. Through an extensive set of experiments, we show that the proposed models are (i) comparable to standard ones in terms of performance in both inductive and transductive tasks, (ii) better calibrated, and (iii) robust against multiple kinds of adversarial attacks.
Incremental Predictive Coding: A Parallel and Fully Automatic Learning Algorithm
Nov 16, 2022



Abstract:Neuroscience-inspired models, such as predictive coding, have the potential to play an important role in the future of machine intelligence. However, they are not yet used in industrial applications due to some limitations, such as the lack of efficiency. In this work, we address this by proposing incremental predictive coding (iPC), a variation of the original framework derived from the incremental expectation maximization algorithm, where every operation can be performed in parallel without external control. We show both theoretically and empirically that iPC is much faster than the original algorithm originally developed by Rao and Ballard, while maintaining performance comparable to backpropagation in image classification tasks. This work impacts several areas, has general applications in computational neuroscience and machine learning, and specific applications in scenarios where automatization and parallelization are important, such as distributed computing and implementations of deep learning models on analog and neuromorphic chips.
Learning to Model Multimodal Semantic Alignment for Story Visualization
Nov 14, 2022



Abstract:Story visualization aims to generate a sequence of images to narrate each sentence in a multi-sentence story, where the images should be realistic and keep global consistency across dynamic scenes and characters. Current works face the problem of semantic misalignment because of their fixed architecture and diversity of input modalities. To address this problem, we explore the semantic alignment between text and image representations by learning to match their semantic levels in the GAN-based generative model. More specifically, we introduce dynamic interactions according to learning to dynamically explore various semantic depths and fuse the different-modal information at a matched semantic level, which thus relieves the text-image semantic misalignment problem. Extensive experiments on different datasets demonstrate the improvements of our approach, neither using segmentation masks nor auxiliary captioning networks, on image quality and story consistency, compared with state-of-the-art methods.
Predictive Coding beyond Gaussian Distributions
Nov 07, 2022



Abstract:A large amount of recent research has the far-reaching goal of finding training methods for deep neural networks that can serve as alternatives to backpropagation (BP). A prominent example is predictive coding (PC), which is a neuroscience-inspired method that performs inference on hierarchical Gaussian generative models. These methods, however, fail to keep up with modern neural networks, as they are unable to replicate the dynamics of complex layers and activation functions. In this work, we solve this problem by generalizing PC to arbitrary probability distributions, enabling the training of architectures, such as transformers, that are hard to approximate with only Gaussian assumptions. We perform three experimental analyses. First, we study the gap between our method and the standard formulation of PC on multiple toy examples. Second, we test the reconstruction quality on variational autoencoders, where our method reaches the same reconstruction quality as BP. Third, we show that our method allows us to train transformer networks and achieve a performance comparable with BP on conditional language models. More broadly, this method allows neuroscience-inspired learning to be applied to multiple domains, since the internal distributions can be flexibly adapted to the data, tasks, and architectures used.
Bird-Eye Transformers for Text Generation Models
Oct 08, 2022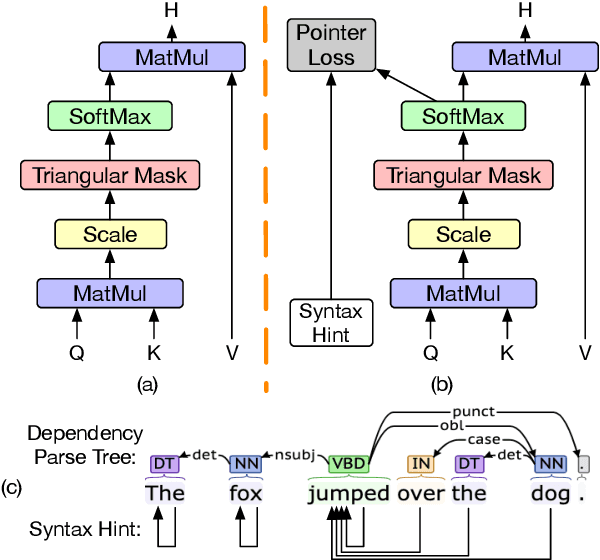
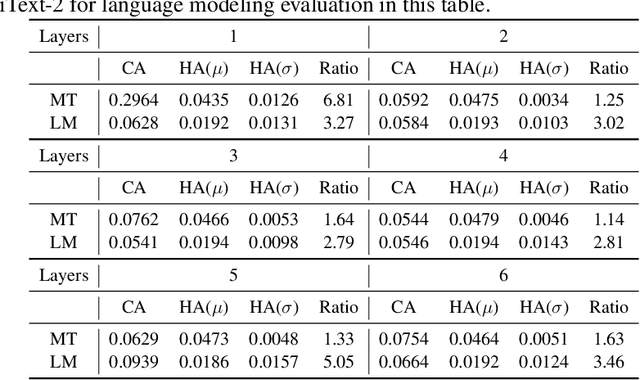
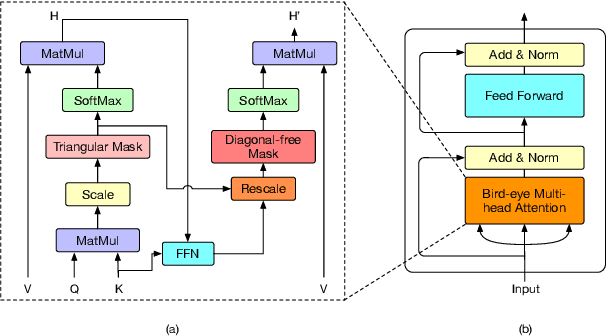
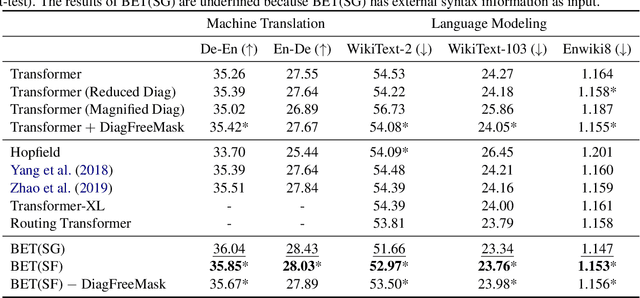
Abstract:Transformers have become an indispensable module for text generation models since their great success in machine translation. Previous works attribute the~success of transformers to the query-key-value dot-product attention, which provides a robust inductive bias by the fully connected token graphs. However, we found that self-attention has a severe limitation. When predicting the (i+1)-th token, self-attention only takes the i-th token as an information collector, and it tends to give a high attention weight to those tokens similar to itself. Therefore, most of the historical information that occurred before the i-th token is not taken into consideration. Based on this observation, in this paper, we propose a new architecture, called bird-eye transformer(BET), which goes one step further to improve the performance of transformers by reweighting self-attention to encourage it to focus more on important historical information. We have conducted experiments on multiple text generation tasks, including machine translation (2 datasets) and language models (3 datasets). These experimental~results show that our proposed model achieves a better performance than the baseline transformer architectures on~all~datasets. The code is released at: \url{https://sites.google.com/view/bet-transformer/home}.
ROAD-R: The Autonomous Driving Dataset with Logical Requirements
Oct 05, 2022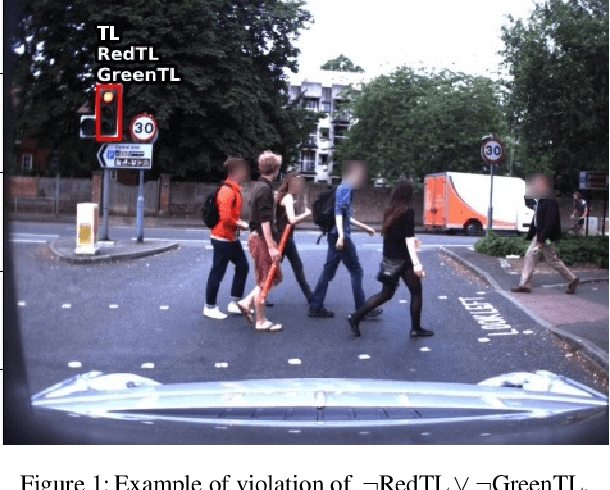
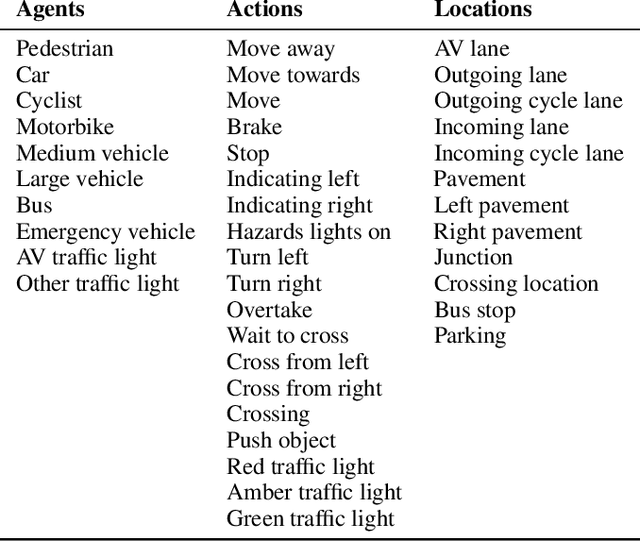


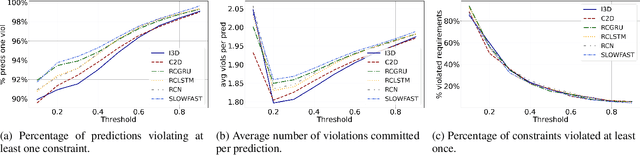
Abstract:Neural networks have proven to be very powerful at computer vision tasks. However, they often exhibit unexpected behaviours, violating known requirements expressing background knowledge. This calls for models (i) able to learn from the requirements, and (ii) guaranteed to be compliant with the requirements themselves. Unfortunately, the development of such models is hampered by the lack of datasets equipped with formally specified requirements. In this paper, we introduce the ROad event Awareness Dataset with logical Requirements (ROAD-R), the first publicly available dataset for autonomous driving with requirements expressed as logical constraints. Given ROAD-R, we show that current state-of-the-art models often violate its logical constraints, and that it is possible to exploit them to create models that (i) have a better performance, and (ii) are guaranteed to be compliant with the requirements themselves.
Efficient Deep Clustering of Human Activities and How to Improve Evaluation
Sep 17, 2022
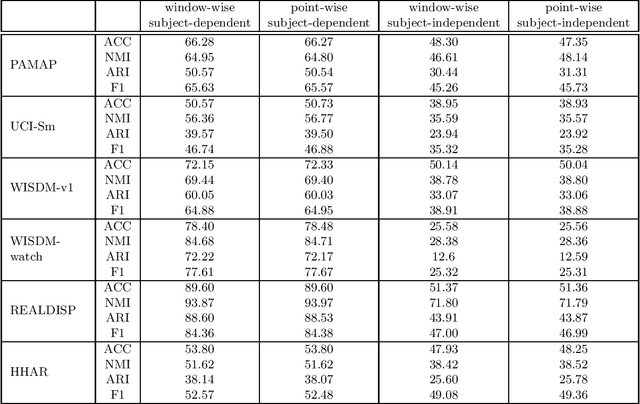
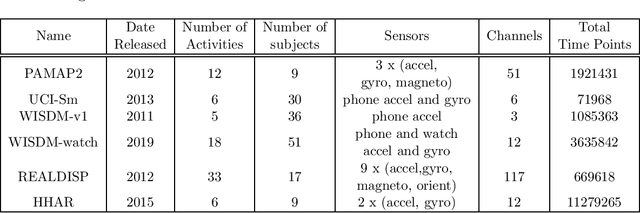
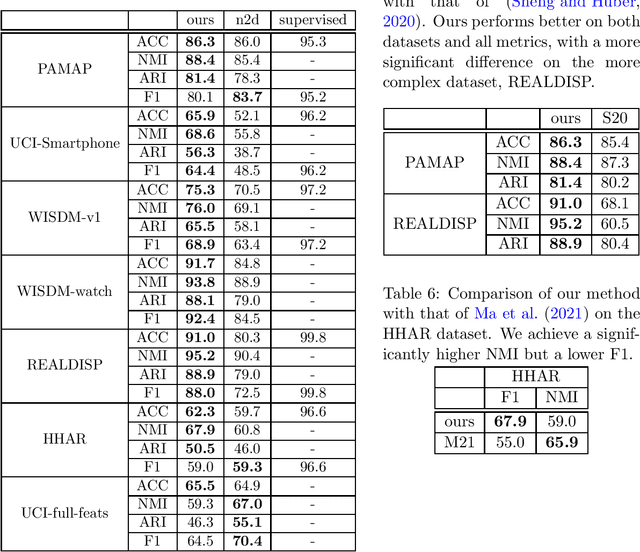
Abstract:There has been much recent research on human activity re\-cog\-ni\-tion (HAR), due to the proliferation of wearable sensors in watches and phones, and the advances of deep learning methods, which avoid the need to manually extract features from raw sensor signals. A significant disadvantage of deep learning applied to HAR is the need for manually labelled training data, which is especially difficult to obtain for HAR datasets. Progress is starting to be made in the unsupervised setting, in the form of deep HAR clustering models, which can assign labels to data without having been given any labels to train on, but there are problems with evaluating deep HAR clustering models, which makes assessing the field and devising new methods difficult. In this paper, we highlight several distinct problems with how deep HAR clustering models are evaluated, describing these problems in detail and conducting careful experiments to explicate the effect that they can have on results. We then discuss solutions to these problems, and suggest standard evaluation settings for future deep HAR clustering models. Additionally, we present a new deep clustering model for HAR. When tested under our proposed settings, our model performs better than (or on par with) existing models, while also being more efficient and better able to scale to more complex datasets by avoiding the need for an autoencoder.
Lightweight Long-Range Generative Adversarial Networks
Sep 08, 2022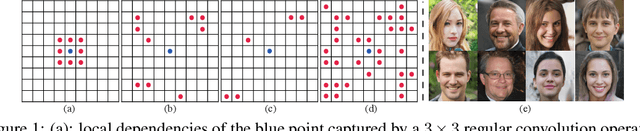
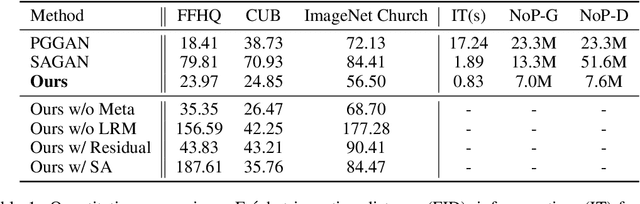
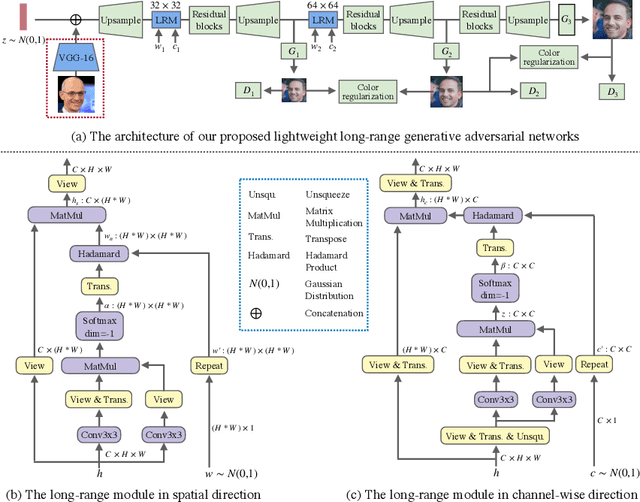

Abstract:In this paper, we introduce novel lightweight generative adversarial networks, which can effectively capture long-range dependencies in the image generation process, and produce high-quality results with a much simpler architecture. To achieve this, we first introduce a long-range module, allowing the network to dynamically adjust the number of focused sampling pixels and to also augment sampling locations. Thus, it can break the limitation of the fixed geometric structure of the convolution operator, and capture long-range dependencies in both spatial and channel-wise directions. Also, the proposed long-range module can highlight negative relations between pixels, working as a regularization to stabilize training. Furthermore, we propose a new generation strategy through which we introduce metadata into the image generation process to provide basic information about target images, which can stabilize and speed up the training process. Our novel long-range module only introduces few additional parameters and is easily inserted into existing models to capture long-range dependencies. Extensive experiments demonstrate the competitive performance of our method with a lightweight architecture.
Memory-Driven Text-to-Image Generation
Aug 15, 2022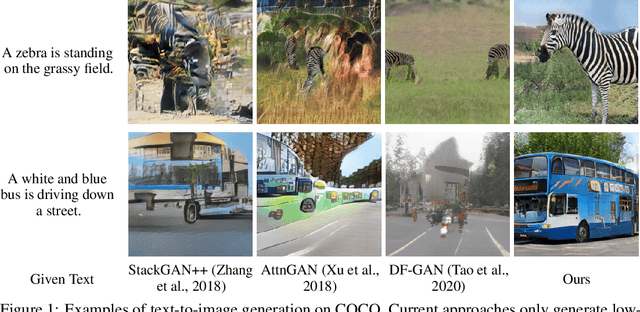
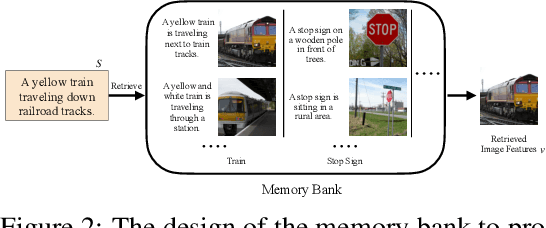
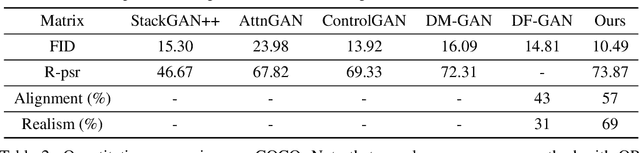

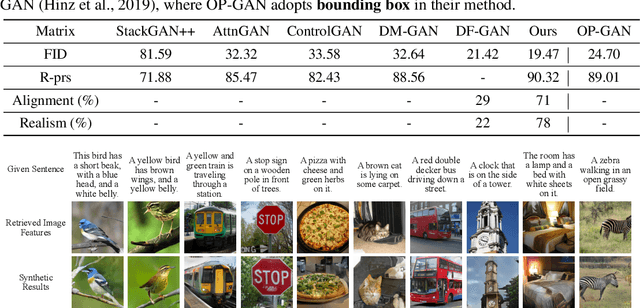
Abstract:We introduce a memory-driven semi-parametric approach to text-to-image generation, which is based on both parametric and non-parametric techniques. The non-parametric component is a memory bank of image features constructed from a training set of images. The parametric component is a generative adversarial network. Given a new text description at inference time, the memory bank is used to selectively retrieve image features that are provided as basic information of target images, which enables the generator to produce realistic synthetic results. We also incorporate the content information into the discriminator, together with semantic features, allowing the discriminator to make a more reliable prediction. Experimental results demonstrate that the proposed memory-driven semi-parametric approach produces more realistic images than purely parametric approaches, in terms of both visual fidelity and text-image semantic consistency.
A Theoretical Framework for Inference and Learning in Predictive Coding Networks
Aug 03, 2022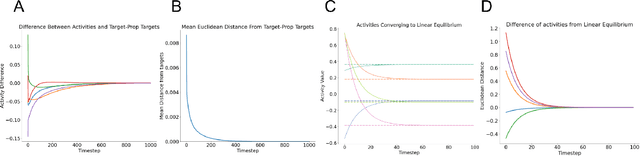
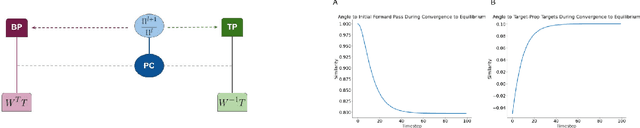
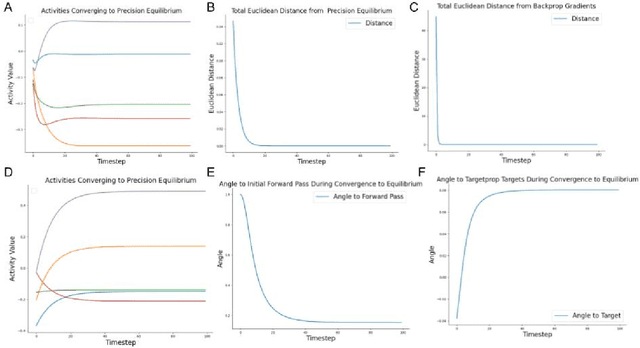
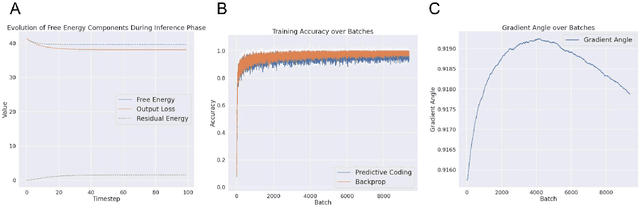
Abstract:Predictive coding (PC) is an influential theory in computational neuroscience, which argues that the cortex forms unsupervised world models by implementing a hierarchical process of prediction error minimization. PC networks (PCNs) are trained in two phases. First, neural activities are updated to optimize the network's response to external stimuli. Second, synaptic weights are updated to consolidate this change in activity -- an algorithm called \emph{prospective configuration}. While previous work has shown how in various limits, PCNs can be found to approximate backpropagation (BP), recent work has demonstrated that PCNs operating in this standard regime, which does not approximate BP, nevertheless obtain competitive training and generalization performance to BP-trained networks while outperforming them on tasks such as online, few-shot, and continual learning, where brains are known to excel. Despite this promising empirical performance, little is understood theoretically about the properties and dynamics of PCNs in this regime. In this paper, we provide a comprehensive theoretical analysis of the properties of PCNs trained with prospective configuration. We first derive analytical results concerning the inference equilibrium for PCNs and a previously unknown close connection relationship to target propagation (TP). Secondly, we provide a theoretical analysis of learning in PCNs as a variant of generalized expectation-maximization and use that to prove the convergence of PCNs to critical points of the BP loss function, thus showing that deep PCNs can, in theory, achieve the same generalization performance as BP, while maintaining their unique advantages.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge