Thomas Kröncke
Bridging MRI and PET physiology: Untangling complementarity through orthogonal representations
Apr 08, 2026Abstract:Multimodal imaging analysis often relies on joint latent representations, yet these approaches rarely define what information is shared versus modality-specific. Clarifying this distinction is clinically relevant, as it delineates the irreducible contribution of each modality and informs rational acquisition strategies. We propose a subspace decomposition framework that reframes multimodal fusion as a problem of orthogonal subspace separation rather than translation. We decompose Prostate-Specific Membrane Antigen (PSMA) PET uptake into an MRI-explainable physiological envelope and an orthogonal residual reflecting signal components not expressible within the MRI feature manifold. Using multiparametric MRI, we train an intensity-based, non-spatial implicit neural representation (INR) to map MRI feature vectors to PET uptake. We introduce a projection-based regularization using singular value decomposition to penalize residual components lying within the span of the MRI feature manifold. This enforces mathematical orthogonality between tissue-level physiological properties (structure, diffusion, perfusion) and intracellular PSMA expression. Tested on 13 prostate cancer patients, the model demonstrates that residual components spanned by MRI features are absorbed into the learned envelope, while the orthogonal residual is largest in tumour regions. This indicates that PSMA PET contains signal components not recoverable from MRI-derived physiological descriptors. The resulting decomposition provides a structured characterization of modality complementarity grounded in representation geometry rather than image translation.
An Uncertainty-Aware, Shareable and Transparent Neural Network Architecture for Brain-Age Modeling
Jul 16, 2021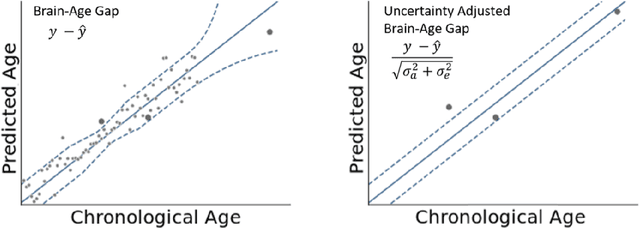
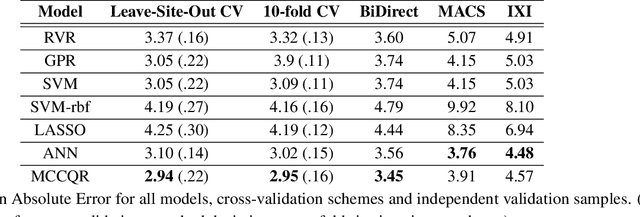
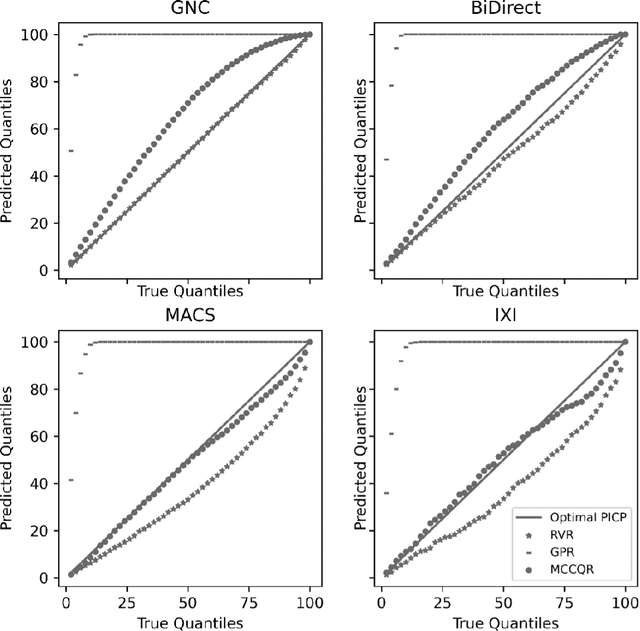
Abstract:The deviation between chronological age and age predicted from neuroimaging data has been identified as a sensitive risk-marker of cross-disorder brain changes, growing into a cornerstone of biological age-research. However, Machine Learning models underlying the field do not consider uncertainty, thereby confounding results with training data density and variability. Also, existing models are commonly based on homogeneous training sets, often not independently validated, and cannot be shared due to data protection issues. Here, we introduce an uncertainty-aware, shareable, and transparent Monte-Carlo Dropout Composite-Quantile-Regression (MCCQR) Neural Network trained on N=10,691 datasets from the German National Cohort. The MCCQR model provides robust, distribution-free uncertainty quantification in high-dimensional neuroimaging data, achieving lower error rates compared to existing models across ten recruitment centers and in three independent validation samples (N=4,004). In two examples, we demonstrate that it prevents spurious associations and increases power to detect accelerated brain-aging. We make the pre-trained model publicly available.
Predicting brain-age from raw T 1 -weighted Magnetic Resonance Imaging data using 3D Convolutional Neural Networks
Mar 22, 2021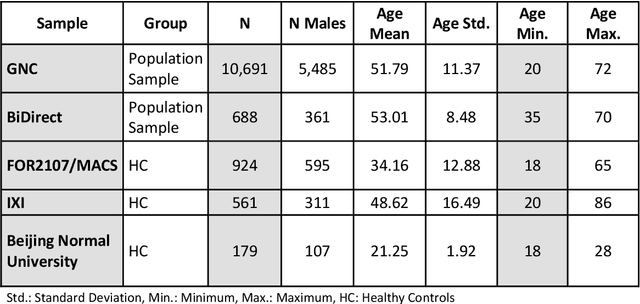
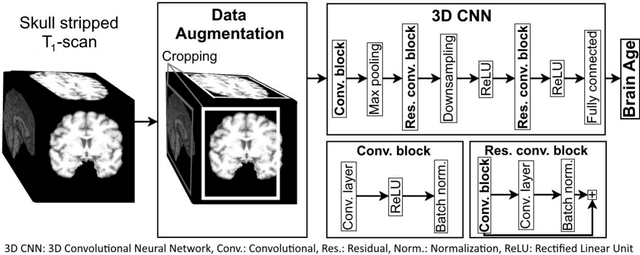
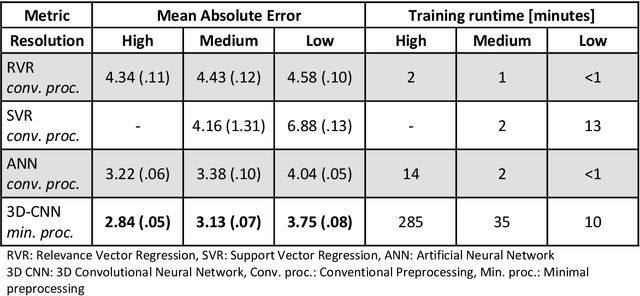
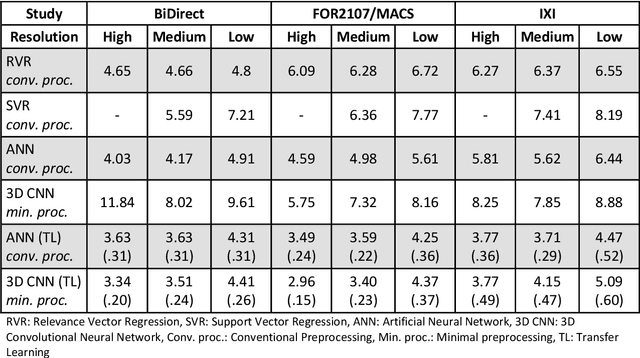
Abstract:Age prediction based on Magnetic Resonance Imaging (MRI) data of the brain is a biomarker to quantify the progress of brain diseases and aging. Current approaches rely on preparing the data with multiple preprocessing steps, such as registering voxels to a standardized brain atlas, which yields a significant computational overhead, hampers widespread usage and results in the predicted brain-age to be sensitive to preprocessing parameters. Here we describe a 3D Convolutional Neural Network (CNN) based on the ResNet architecture being trained on raw, non-registered T$_ 1$-weighted MRI data of N=10,691 samples from the German National Cohort and additionally applied and validated in N=2,173 samples from three independent studies using transfer learning. For comparison, state-of-the-art models using preprocessed neuroimaging data are trained and validated on the same samples. The 3D CNN using raw neuroimaging data predicts age with a mean average deviation of 2.84 years, outperforming the state-of-the-art brain-age models using preprocessed data. Since our approach is invariant to preprocessing software and parameter choices, it enables faster, more robust and more accurate brain-age modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge