Nils Opel
An Uncertainty-Aware, Shareable and Transparent Neural Network Architecture for Brain-Age Modeling
Jul 16, 2021
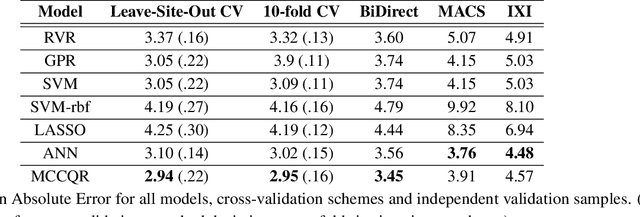
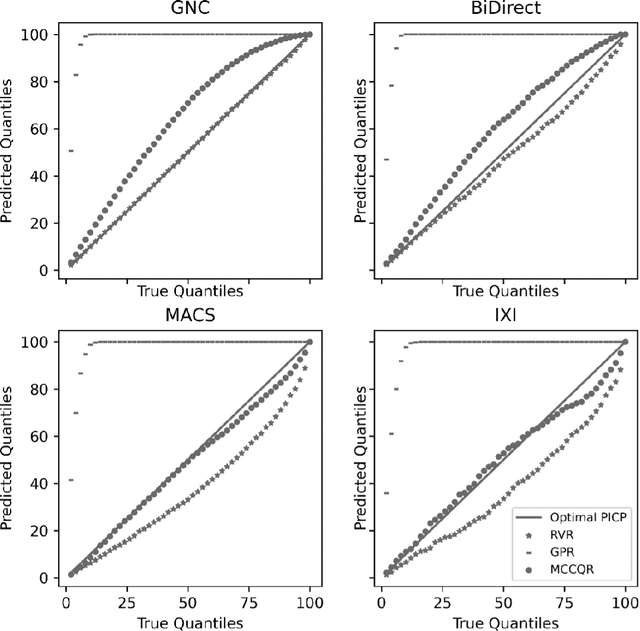
Abstract:The deviation between chronological age and age predicted from neuroimaging data has been identified as a sensitive risk-marker of cross-disorder brain changes, growing into a cornerstone of biological age-research. However, Machine Learning models underlying the field do not consider uncertainty, thereby confounding results with training data density and variability. Also, existing models are commonly based on homogeneous training sets, often not independently validated, and cannot be shared due to data protection issues. Here, we introduce an uncertainty-aware, shareable, and transparent Monte-Carlo Dropout Composite-Quantile-Regression (MCCQR) Neural Network trained on N=10,691 datasets from the German National Cohort. The MCCQR model provides robust, distribution-free uncertainty quantification in high-dimensional neuroimaging data, achieving lower error rates compared to existing models across ten recruitment centers and in three independent validation samples (N=4,004). In two examples, we demonstrate that it prevents spurious associations and increases power to detect accelerated brain-aging. We make the pre-trained model publicly available.
Predicting brain-age from raw T 1 -weighted Magnetic Resonance Imaging data using 3D Convolutional Neural Networks
Mar 22, 2021
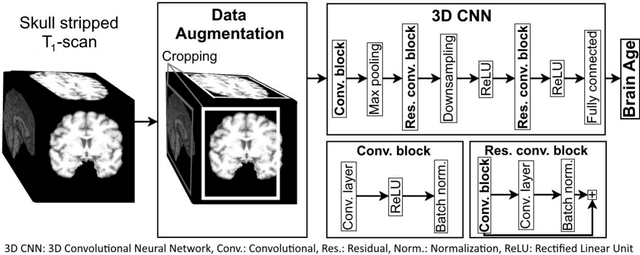
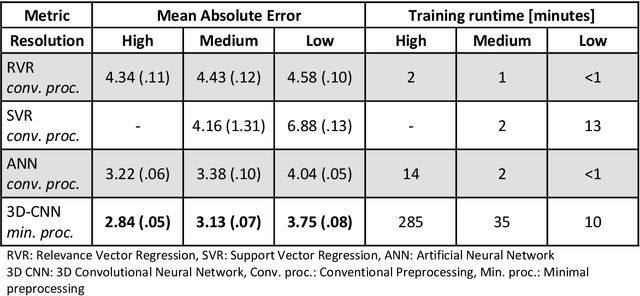
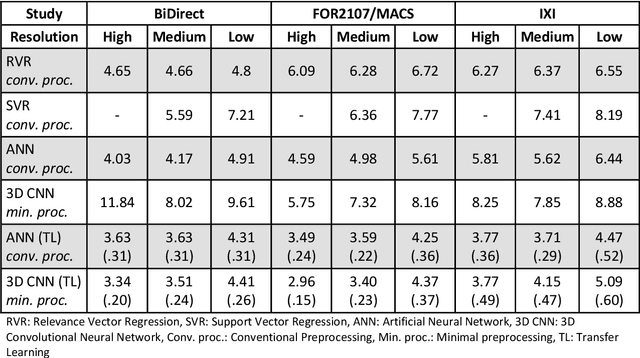
Abstract:Age prediction based on Magnetic Resonance Imaging (MRI) data of the brain is a biomarker to quantify the progress of brain diseases and aging. Current approaches rely on preparing the data with multiple preprocessing steps, such as registering voxels to a standardized brain atlas, which yields a significant computational overhead, hampers widespread usage and results in the predicted brain-age to be sensitive to preprocessing parameters. Here we describe a 3D Convolutional Neural Network (CNN) based on the ResNet architecture being trained on raw, non-registered T$_ 1$-weighted MRI data of N=10,691 samples from the German National Cohort and additionally applied and validated in N=2,173 samples from three independent studies using transfer learning. For comparison, state-of-the-art models using preprocessed neuroimaging data are trained and validated on the same samples. The 3D CNN using raw neuroimaging data predicts age with a mean average deviation of 2.84 years, outperforming the state-of-the-art brain-age models using preprocessed data. Since our approach is invariant to preprocessing software and parameter choices, it enables faster, more robust and more accurate brain-age modeling.
PHOTON -- A Python API for Rapid Machine Learning Model Development
Feb 13, 2020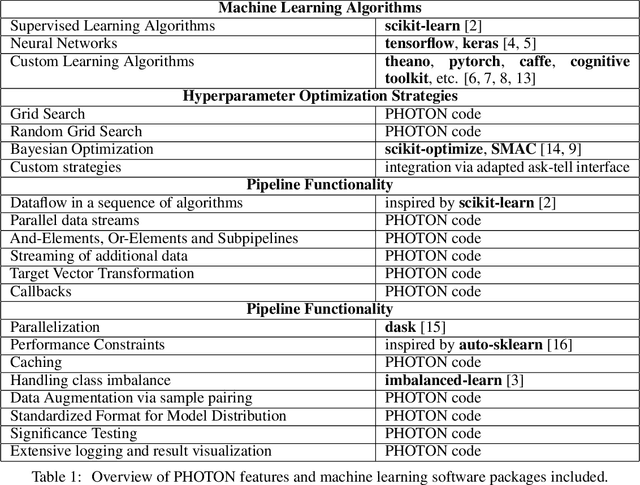
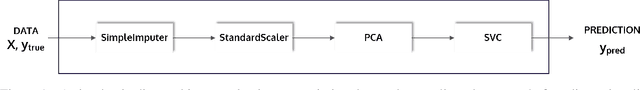
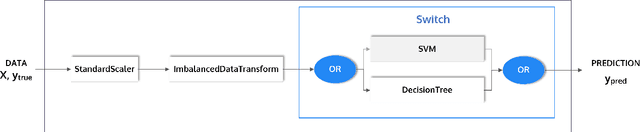
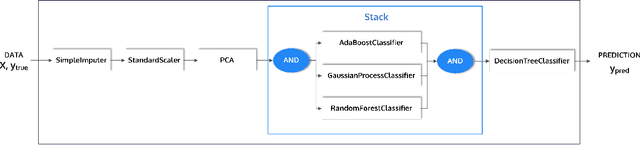
Abstract:This article describes the implementation and use of PHOTON, a high-level Python API designed to simplify and accelerate the process of machine learning model development. It enables designing both basic and advanced machine learning pipeline architectures and automatizes the repetitive training, optimization and evaluation workflow. PHOTON offers easy access to established machine learning toolboxes as well as the possibility to integrate custom algorithms and solutions for any part of the model construction and evaluation process. By adding a layer of abstraction incorporating current best practices it offers an easy-to-use, flexible approach to implementing fast, reproducible, and unbiased machine learning solutions.
Systematic Overestimation of Machine Learning Performance in Neuroimaging Studies of Depression
Dec 13, 2019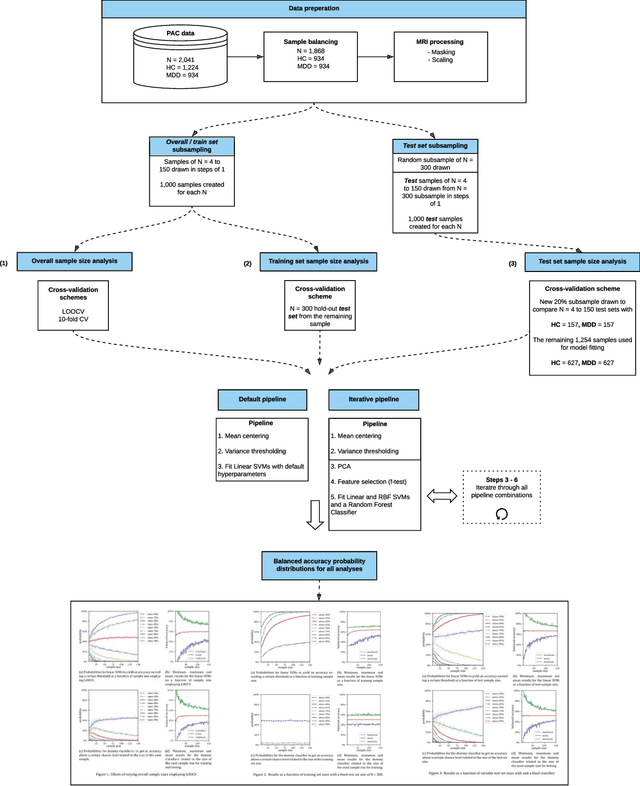
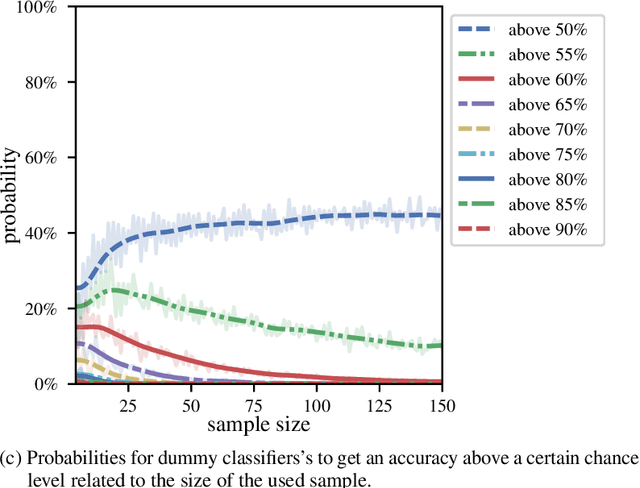
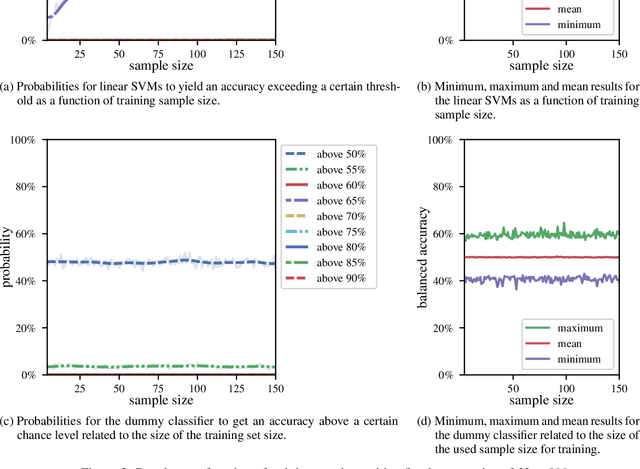
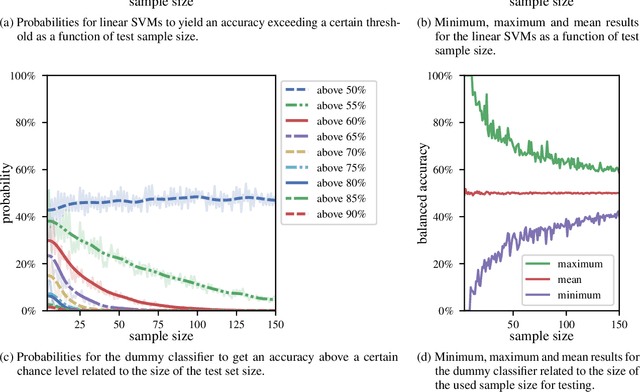
Abstract:We currently observe a disconcerting phenomenon in machine learning studies in psychiatry: While we would expect larger samples to yield better results due to the availability of more data, larger machine learning studies consistently show much weaker performance than the numerous small-scale studies. Here, we systematically investigated this effect focusing on one of the most heavily studied questions in the field, namely the classification of patients suffering from Major Depressive Disorder (MDD) and healthy controls. Drawing upon a balanced sample of $N = 1,868$ MDD patients and healthy controls from our recent international Predictive Analytics Competition (PAC), we first trained and tested a classification model on the full dataset which yielded an accuracy of 61%. Next, we mimicked the process by which researchers would draw samples of various sizes ($N=4$ to $N=150$) from the population and showed a strong risk of overestimation. Specifically, for small sample sizes ($N=20$), we observe accuracies of up to 95%. For medium sample sizes ($N=100$) accuracies up to 75% were found. Importantly, further investigation showed that sufficiently large test sets effectively protect against performance overestimation whereas larger datasets per se do not. While these results question the validity of a substantial part of the current literature, we outline the relatively low-cost remedy of larger test sets.
Biological sex classification with structural MRI data shows increased misclassification in transgender women
Nov 24, 2019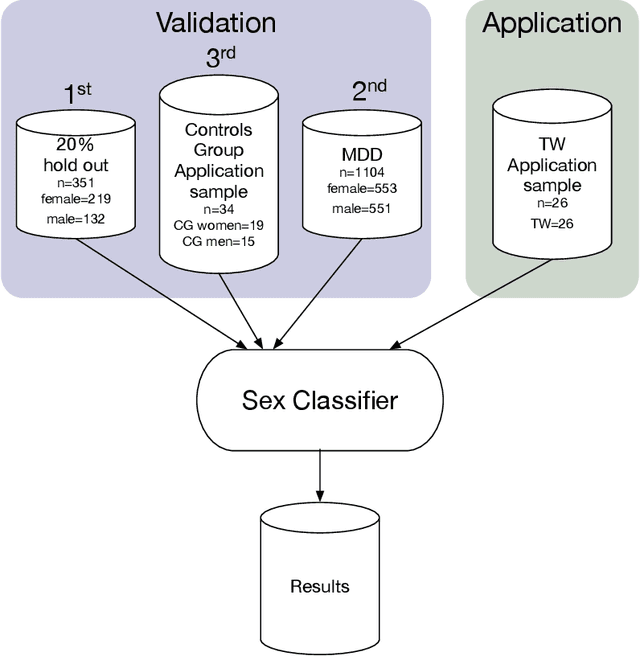
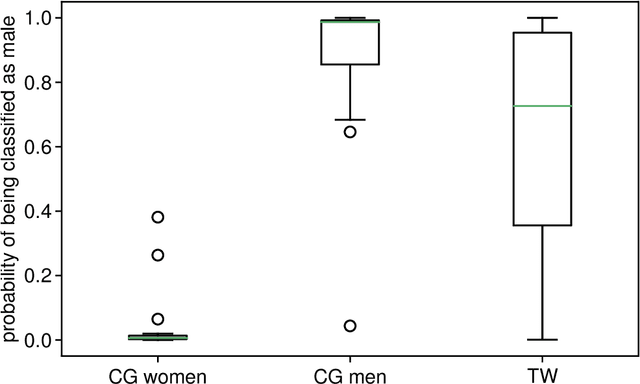
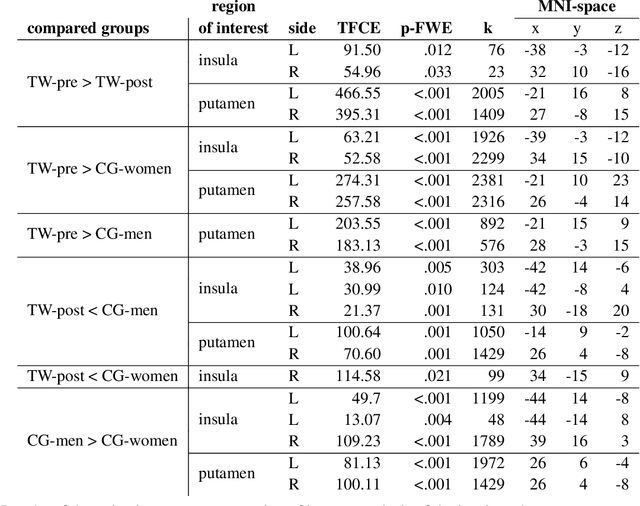
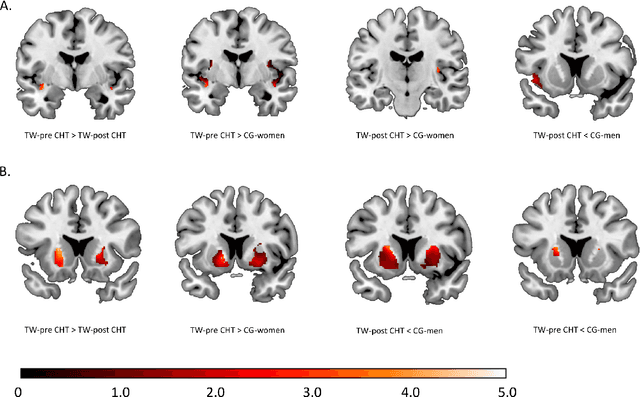
Abstract:Transgender individuals show brain structural alterations that differ from their biological sex as well as their perceived gender. To substantiate evidence that the brain structure of transgender individuals differs from male and female, we use a combined multivariate and univariate approach. Gray matter segments resulting from voxel-based morphometry preprocessing of N = 1753 cisgender (CG) healthy participants were used to train (N = 1402) and validate (20% hold-out N = 351) a support vector machine classifying the biological sex. As a second validation, we classified N = 1104 patients with depression. A third validation was performed using the matched CG sample of the transgender women (TW) application sample. Subsequently, the classifier was applied to N = 25 TW. Finally, we compared brain volumes of CG-men, women and TW pre/post treatment (CHT) in a univariate analysis controlling for sexual orientation, age and total brain volume. The application of our biological sex classifier to the transgender sample resulted in a significantly lower true positive rate (TPR-male = 56.0%). The TPR did not differ between CG-individuals with (TPR-male = 86.9%) and without depression (TPR-male = 88.5%). The univariate analysis of the transgender application sample revealed that TW pre/post treatment show brain structural differences from CG-women and CG-men in the putamen and insula, as well as the whole-brain analysis. Our results support the hypothesis that brain structure in TW differs from brain structure of their biological sex (male) as well as their perceived gender (female). This finding substantiates evidence that transgender individuals show specific brain structural alterations leading to a different pattern of brain structure than CG individuals.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge