Steven Truong
GIIM: Graph-based Learning of Inter- and Intra-view Dependencies for Multi-view Medical Image Diagnosis
Mar 10, 2026Abstract:Computer-aided diagnosis (CADx) has become vital in medical imaging, but automated systems often struggle to replicate the nuanced process of clinical interpretation. Expert diagnosis requires a comprehensive analysis of how abnormalities relate to each other across various views and time points, but current multi-view CADx methods frequently overlook these complex dependencies. Specifically, they fail to model the crucial relationships within a single view and the dynamic changes lesions exhibit across different views. This limitation, combined with the common challenge of incomplete data, greatly reduces their predictive reliability. To address these gaps, we reframe the diagnostic task as one of relationship modeling and propose GIIM, a novel graph-based approach. Our framework is uniquely designed to simultaneously capture both critical intra-view dependencies between abnormalities and inter-view dynamics. Furthermore, it ensures diagnostic robustness by incorporating specific techniques to effectively handle missing data, a common clinical issue. We demonstrate the generality of this approach through extensive evaluations on diverse imaging modalities, including CT, MRI, and mammography. The results confirm that our GIIM model significantly enhances diagnostic accuracy and robustness over existing methods, establishing a more effective framework for future CADx systems.
Improving the Performance of Radiology Report De-identification with Large-Scale Training and Benchmarking Against Cloud Vendor Methods
Nov 06, 2025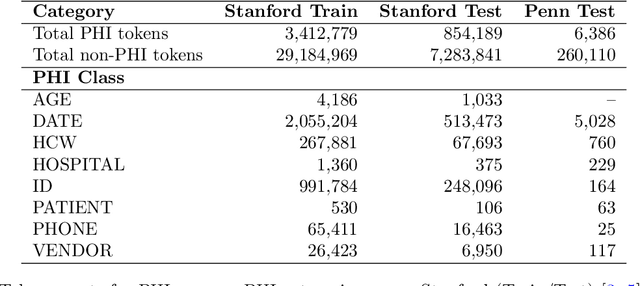

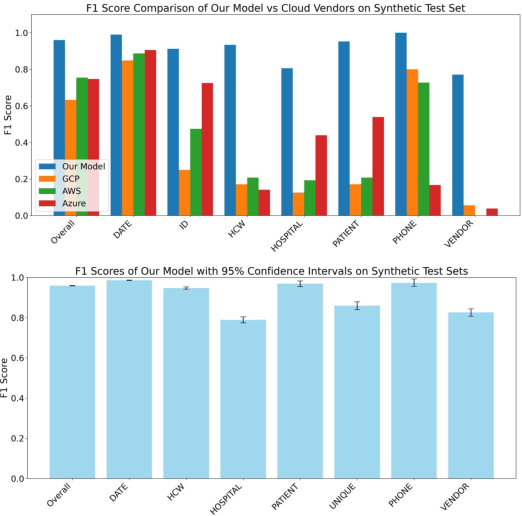
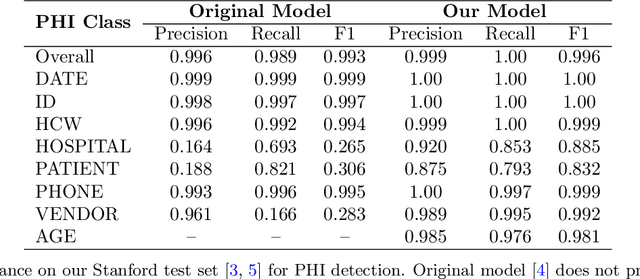
Abstract:Objective: To enhance automated de-identification of radiology reports by scaling transformer-based models through extensive training datasets and benchmarking performance against commercial cloud vendor systems for protected health information (PHI) detection. Materials and Methods: In this retrospective study, we built upon a state-of-the-art, transformer-based, PHI de-identification pipeline by fine-tuning on two large annotated radiology corpora from Stanford University, encompassing chest X-ray, chest CT, abdomen/pelvis CT, and brain MR reports and introducing an additional PHI category (AGE) into the architecture. Model performance was evaluated on test sets from Stanford and the University of Pennsylvania (Penn) for token-level PHI detection. We further assessed (1) the stability of synthetic PHI generation using a "hide-in-plain-sight" method and (2) performance against commercial systems. Precision, recall, and F1 scores were computed across all PHI categories. Results: Our model achieved overall F1 scores of 0.973 on the Penn dataset and 0.996 on the Stanford dataset, outperforming or maintaining the previous state-of-the-art model performance. Synthetic PHI evaluation showed consistent detectability (overall F1: 0.959 [0.958-0.960]) across 50 independently de-identified Penn datasets. Our model outperformed all vendor systems on synthetic Penn reports (overall F1: 0.960 vs. 0.632-0.754). Discussion: Large-scale, multimodal training improved cross-institutional generalization and robustness. Synthetic PHI generation preserved data utility while ensuring privacy. Conclusion: A transformer-based de-identification model trained on diverse radiology datasets outperforms prior academic and commercial systems in PHI detection and establishes a new benchmark for secure clinical text processing.
A Nested Attention Neural Hybrid Model for Grammatical Error Correction
Jul 10, 2017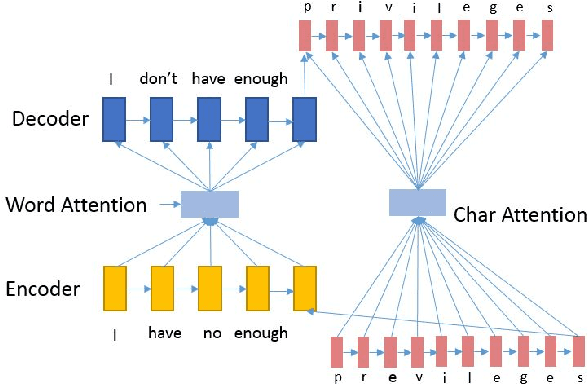
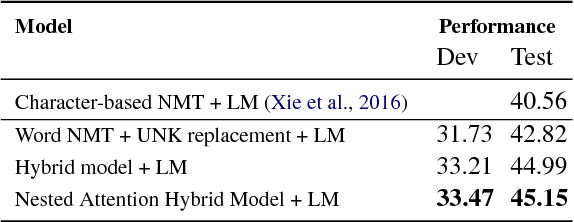
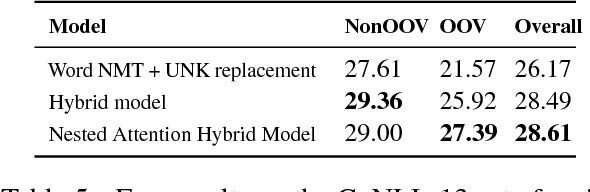



Abstract:Grammatical error correction (GEC) systems strive to correct both global errors in word order and usage, and local errors in spelling and inflection. Further developing upon recent work on neural machine translation, we propose a new hybrid neural model with nested attention layers for GEC. Experiments show that the new model can effectively correct errors of both types by incorporating word and character-level information,and that the model significantly outperforms previous neural models for GEC as measured on the standard CoNLL-14 benchmark dataset. Further analysis also shows that the superiority of the proposed model can be largely attributed to the use of the nested attention mechanism, which has proven particularly effective in correcting local errors that involve small edits in orthography.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge