Steve Jiang
Medical Artificial Intelligence and Automation Laboratory and Department of Radiation Oncology, UT Southwestern Medical Center, Dallas TX 75235, USA
Deep Dose Plugin Towards Real-time Monte Carlo Dose Calculation Through a Deep Learning based Denoising Algorithm
Dec 23, 2020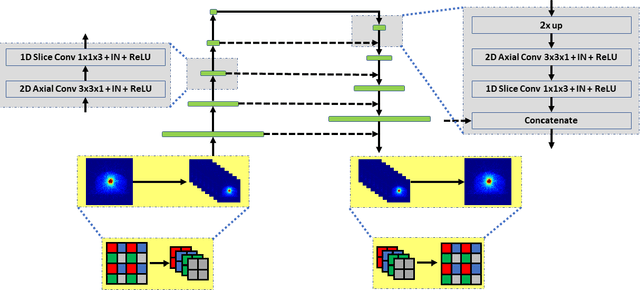
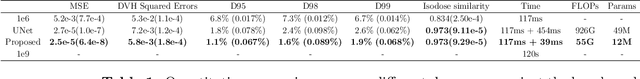
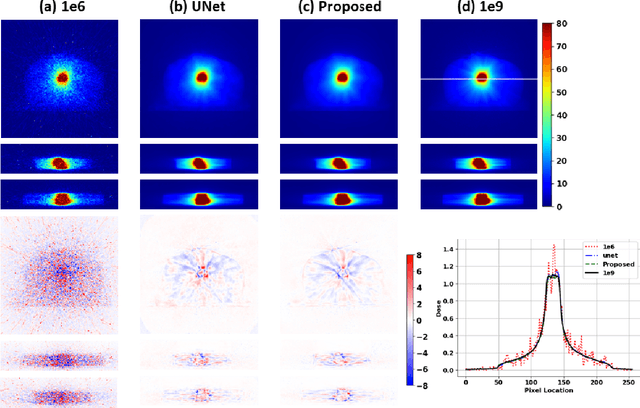
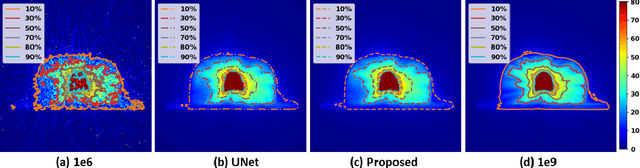
Abstract:Monte Carlo (MC) simulation is considered the gold standard method for radiotherapy dose calculation. However, achieving high precision requires a large number of simulation histories, which is time consuming. The use of computer graphics processing units (GPUs) has greatly accelerated MC simulation and allows dose calculation within a few minutes for a typical radiotherapy treatment plan. However, some clinical applications demand real time efficiency for MC dose calculation. To tackle this problem, we have developed a real time, deep learning based dose denoiser that can be plugged into a current GPU based MC dose engine to enable real time MC dose calculation. We used two different acceleration strategies to achieve this goal: 1) we applied voxel unshuffle and voxel shuffle operators to decrease the input and output sizes without any information loss, and 2) we decoupled the 3D volumetric convolution into a 2D axial convolution and a 1D slice convolution. In addition, we used a weakly supervised learning framework to train the network, which greatly reduces the size of the required training dataset and thus enables fast fine tuning based adaptation of the trained model to different radiation beams. Experimental results show that the proposed denoiser can run in as little as 39 ms, which is around 11.6 times faster than the baseline model. As a result, the whole MC dose calculation pipeline can be finished within 0.15 seconds, including both GPU MC dose calculation and deep learning based denoising, achieving the real time efficiency needed for some radiotherapy applications, such as online adaptive radiotherapy.
Deep Interactive Denoiser for X-Ray Computed Tomography
Dec 06, 2020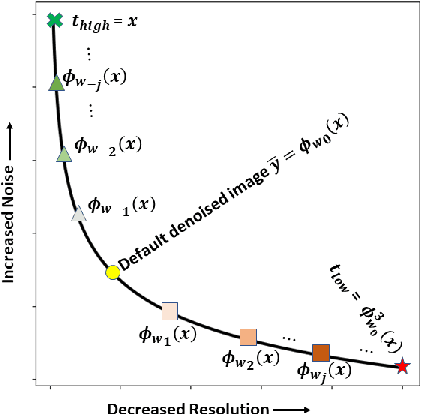
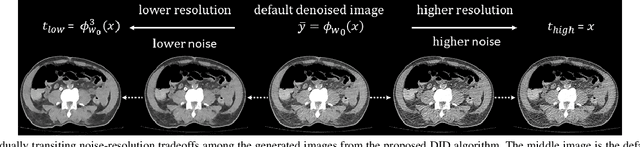
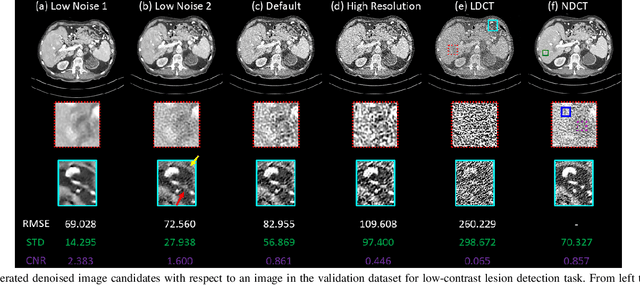
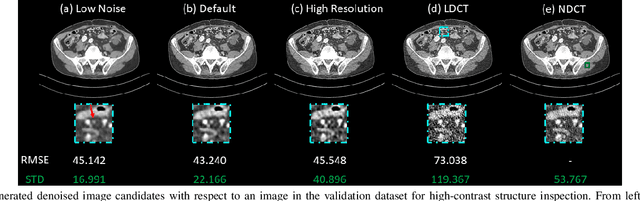
Abstract:Low dose computed tomography (LDCT) is desirable for both diagnostic imaging and image guided interventions. Denoisers are openly used to improve the quality of LDCT. Deep learning (DL)-based denoisers have shown state-of-the-art performance and are becoming one of the mainstream methods. However, there exists two challenges regarding the DL-based denoisers: 1) a trained model typically does not generate different image candidates with different noise-resolution tradeoffs which sometimes are needed for different clinical tasks; 2) the model generalizability might be an issue when the noise level in the testing images is different from that in the training dataset. To address these two challenges, in this work, we introduce a lightweight optimization process at the testing phase on top of any existing DL-based denoisers to generate multiple image candidates with different noise-resolution tradeoffs suitable for different clinical tasks in real-time. Consequently, our method allows the users to interact with the denoiser to efficiently review various image candidates and quickly pick up the desired one, and thereby was termed as deep interactive denoiser (DID). Experimental results demonstrated that DID can deliver multiple image candidates with different noise-resolution tradeoffs, and shows great generalizability regarding various network architectures, as well as training and testing datasets with various noise levels.
Using Monte Carlo dropout and bootstrap aggregation for uncertainty estimation in radiation therapy dose prediction with deep learning neural networks
Nov 01, 2020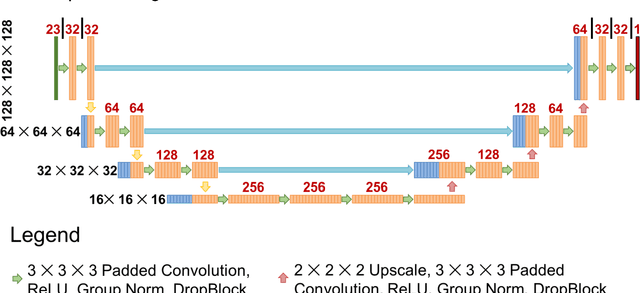
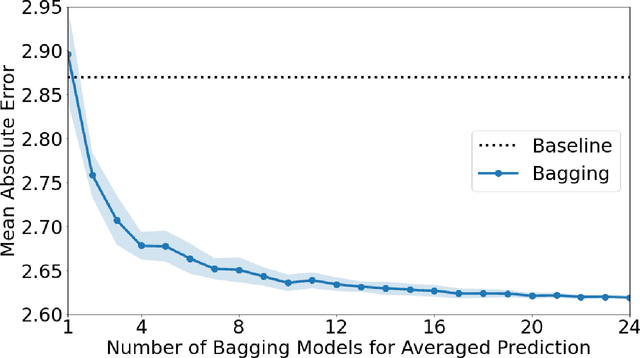
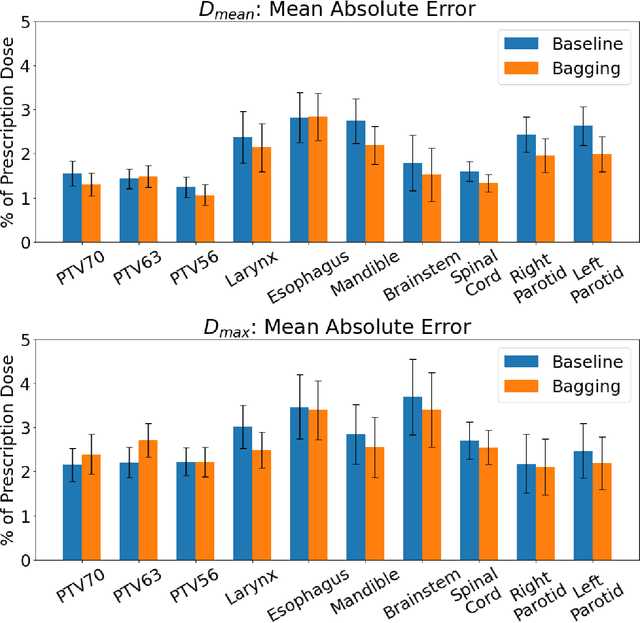
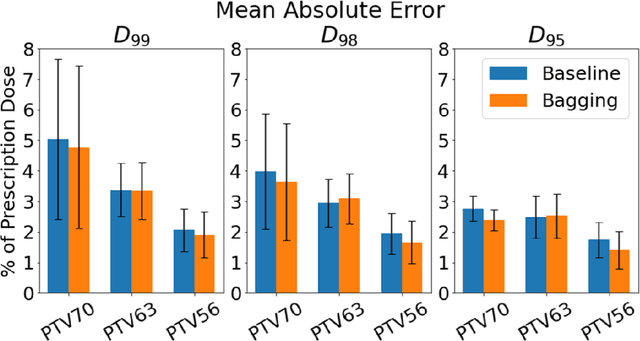
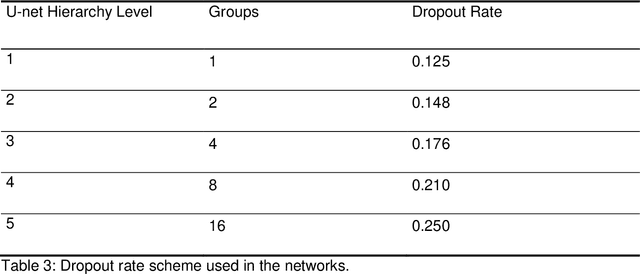
Abstract:Recently, artificial intelligence technologies and algorithms have become a major focus for advancements in treatment planning for radiation therapy. As these are starting to become incorporated into the clinical workflow, a major concern from clinicians is not whether the model is accurate, but whether the model can express to a human operator when it does not know if its answer is correct. We propose to use Monte Carlo dropout (MCDO) and the bootstrap aggregation (bagging) technique on deep learning models to produce uncertainty estimations for radiation therapy dose prediction. We show that both models are capable of generating a reasonable uncertainty map, and, with our proposed scaling technique, creating interpretable uncertainties and bounds on the prediction and any relevant metrics. Performance-wise, bagging provides statistically significant reduced loss value and errors in most of the metrics investigated in this study. The addition of bagging was able to further reduce errors by another 0.34% for Dmean and 0.19% for Dmax, on average, when compared to the baseline framework. Overall, the bagging framework provided significantly lower MAE of 2.62, as opposed to the baseline framework's MAE of 2.87. The usefulness of bagging, from solely a performance standpoint, does highly depend on the problem and the acceptable predictive error, and its high upfront computational cost during training should be factored in to deciding whether it is advantageous to use it. In terms of deployment with uncertainty estimations turned on, both frameworks offer the same performance time of about 12 seconds. As an ensemble-based metaheuristic, bagging can be used with existing machine learning architectures to improve stability and performance, and MCDO can be applied to any deep learning models that have dropout as part of their architecture.
Dose Prediction with Deep Learning for Prostate Cancer Radiation Therapy: Model Adaptation to Different Treatment Planning Practices
Jun 30, 2020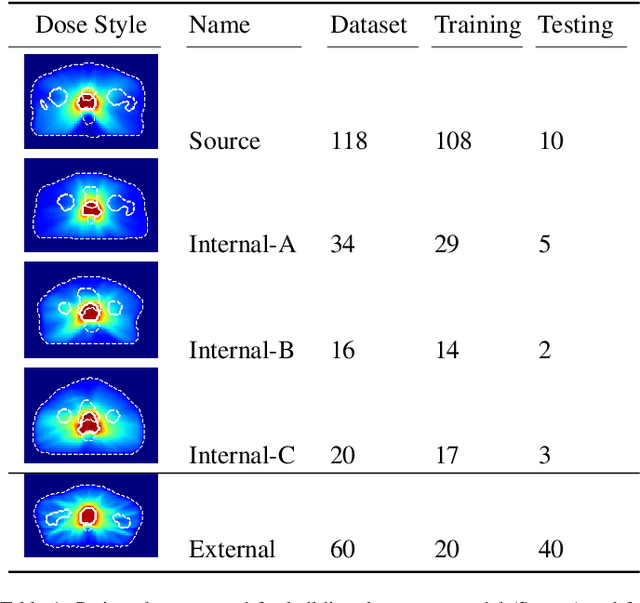
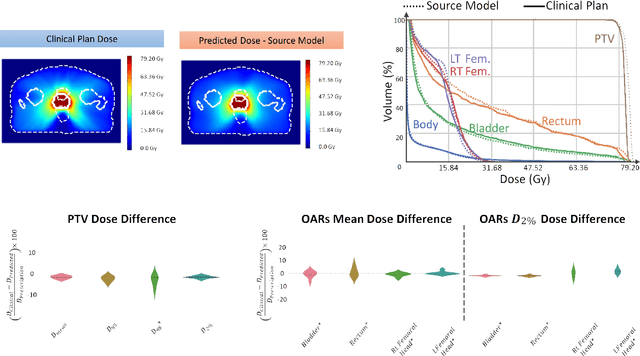
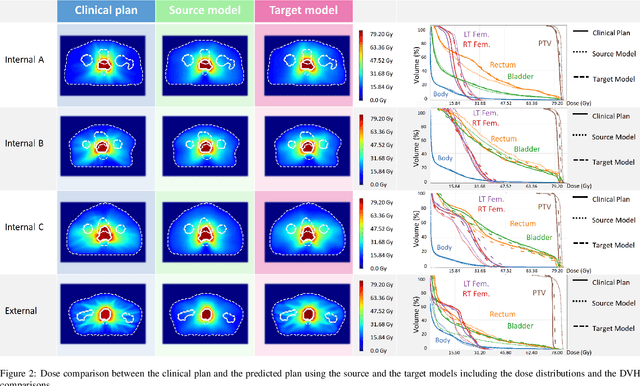
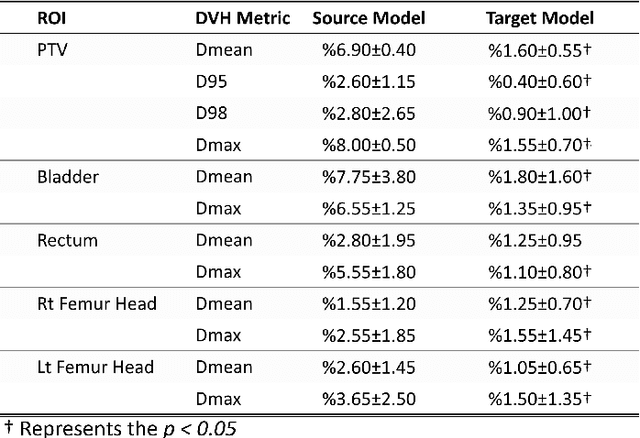
Abstract:This work aims to study the generalizability of a pre-developed deep learning (DL) dose prediction model for volumetric modulated arc therapy (VMAT) for prostate cancer and to adapt the model to three different internal treatment planning styles and one external institution planning style. We built the source model with planning data from 108 patients previously treated with VMAT for prostate cancer. For the transfer learning, we selected patient cases planned with three different styles from the same institution and one style from a different institution to adapt the source model to four target models. We compared the dose distributions predicted by the source model and the target models with the clinical dose predictions and quantified the improvement in the prediction quality for the target models over the source model using the Dice similarity coefficients (DSC) of 10% to 100% isodose volumes and the dose-volume-histogram (DVH) parameters of the planning target volume and the organs-at-risk. The source model accurately predicts dose distributions for plans generated in the same source style but performs sub-optimally for the three internal and one external target styles, with the mean DSC ranging between 0.81-0.94 and 0.82-0.91 for the internal and the external styles, respectively. With transfer learning, the target model predictions improved the mean DSC to 0.88-0.95 and 0.92-0.96 for the internal and the external styles, respectively. Target model predictions significantly improved the accuracy of the DVH parameter predictions to within 1.6%. We demonstrated model generalizability for DL-based dose prediction and the feasibility of using transfer learning to solve this problem. With 14-29 cases per style, we successfully adapted the source model into several different practice styles. This indicates a realistic way to widespread clinical implementation of DL-based dose prediction.
Using Deep Learning to Predict Beam-Tunable Pareto Optimal Dose Distribution for Intensity Modulated Radiation Therapy
Jun 19, 2020

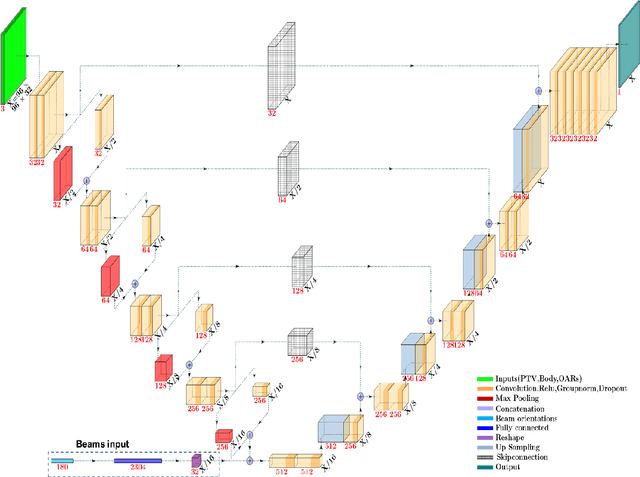
Abstract:We propose to develop deep learning models that can predict Pareto optimal dose distributions by using any given set of beam angles, along with patient anatomy, as input to train the deep neural networks. We implement and compare two deep learning networks that predict with two different beam configuration modalities. We generated Pareto optimal plans for 70 patients with prostate cancer. We used fluence map optimization to generate 500 IMRT plans that sampled the Pareto surface for each patient, for a total of 35,000 plans. We studied and compared two different models, Model I and Model II. Model I directly uses beam angles as a second input to the network as a binary vector. Model II converts the beam angles into beam doses that are conformal to the PTV. Our deep learning models predicted voxel-level dose distributions that precisely matched the ground truth dose distributions. Quantitatively, Model I prediction error of 0.043 (confirmation), 0.043 (homogeneity), 0.327 (R50), 2.80% (D95), 3.90% (D98), 0.6% (D50), 1.10% (D2) was lower than that of Model II, which obtained 0.076 (confirmation), 0.058 (homogeneity), 0.626 (R50), 7.10% (D95), 6.50% (D98), 8.40% (D50), 6.30% (D2). Treatment planners who use our models will be able to use deep learning to control the tradeoffs between the PTV and OAR weights, as well as the beam number and configurations in real time. Our dose prediction methods provide a stepping stone to building automatic IMRT treatment planning.
Probabilistic self-learning framework for Low-dose CT Denoising
May 30, 2020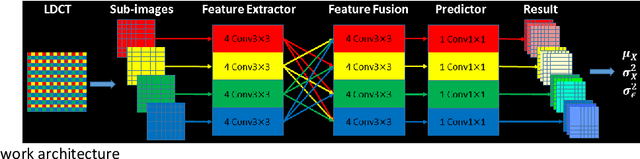
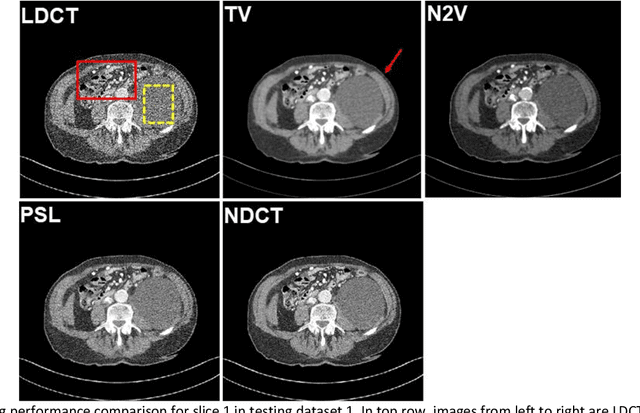
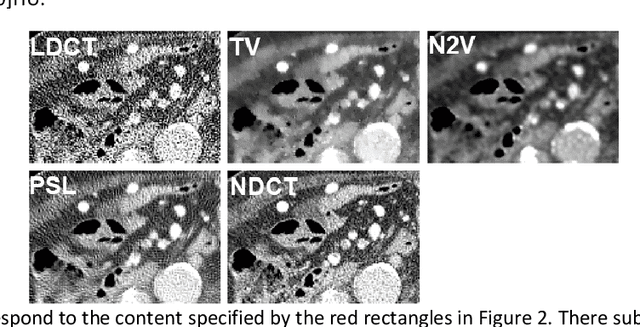
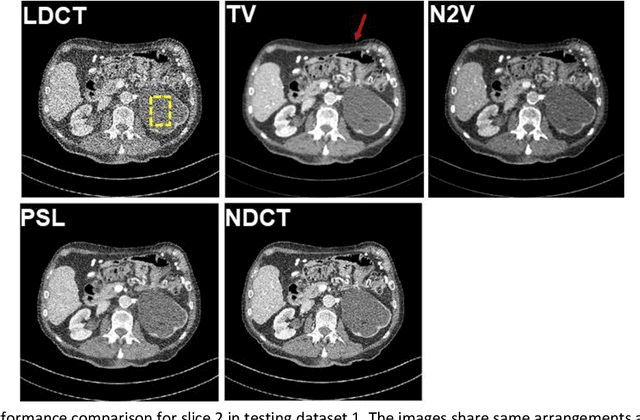
Abstract:Despite the indispensable role of X-ray computed tomography (CT) in diagnostic medicine field, the associated ionizing radiation is still a major concern considering that it may cause genetic and cancerous diseases. Decreasing the exposure can reduce the dose and hence the radiation-related risk, but will also induce higher quantum noise. Supervised deep learning can be used to train a neural network to denoise the low-dose CT (LDCT). However, its success requires massive pixel-wise paired LDCT and normal-dose CT (NDCT) images, which are rarely available in real practice. To alleviate this problem, in this paper, a shift-invariant property based neural network was devised to learn the inherent pixel correlations and also the noise distribution by only using the LDCT images, shaping into our probabilistic self-learning framework. Experimental results demonstrated that the proposed method outperformed the competitors, producing an enhanced LDCT image that has similar image style as the routine NDCT which is highly-preferable in clinic practice.
A deep learning-based framework for segmenting invisible clinical target volumes with estimated uncertainties for post-operative prostate cancer radiotherapy
Apr 28, 2020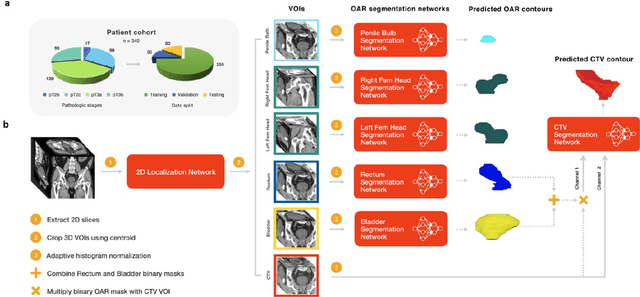
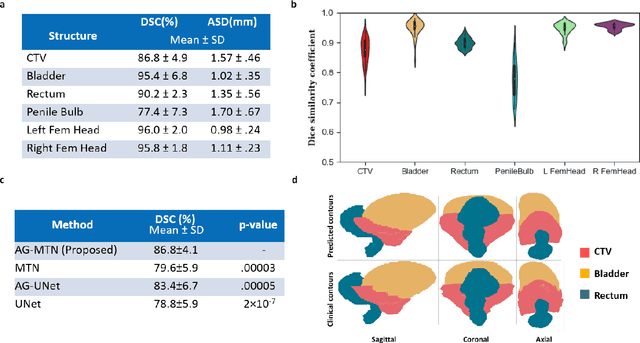
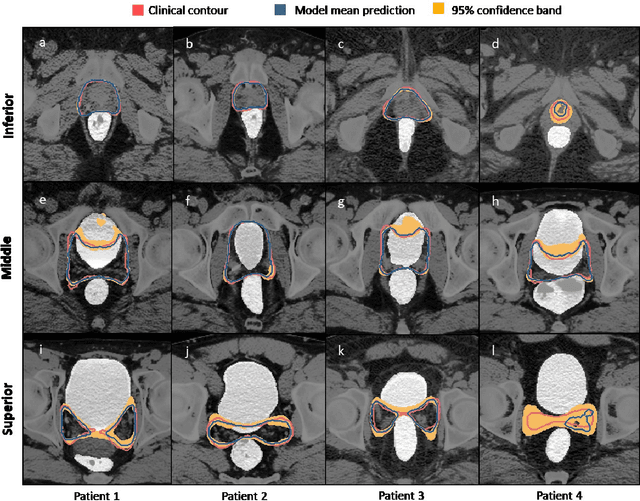
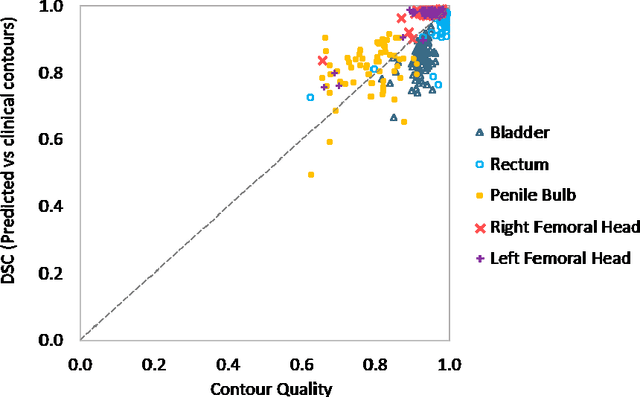
Abstract:In post-operative radiotherapy for prostate cancer, the cancerous prostate gland has been surgically removed, so the clinical target volume (CTV) to be irradiated encompasses the microscopic spread of tumor cells, which cannot be visualized in typical clinical images such as computed tomography or magnetic resonance imaging. In current clinical practice, physicians segment CTVs manually based on their relationship with nearby organs and other clinical information, per clinical guidelines. Automating post-operative prostate CTV segmentation with traditional image segmentation methods has been a major challenge. Here, we propose a deep learning model to overcome this problem by segmenting nearby organs first, then using their relationship with the CTV to assist CTV segmentation. The model proposed is trained using labels clinically approved and used for patient treatment, which are subject to relatively large inter-physician variations due to the absence of a visual ground truth. The model achieves an average Dice similarity coefficient (DSC) of 0.87 on a holdout dataset of 50 patients, much better than established methods, such as atlas-based methods (DSC<0.7). The uncertainties associated with automatically segmented CTV contours are also estimated to help physicians inspect and revise the contours, especially in areas with large inter-physician variations. We also use a 4-point grading system to show that the clinical quality of the automatically segmented CTV contours is equal to that of approved clinical contours manually drawn by physicians.
Generalizability issues with deep learning models in medicine and their potential solutions: illustrated with Cone-Beam Computed Tomography (CBCT) to Computed Tomography (CT) image conversion
Apr 17, 2020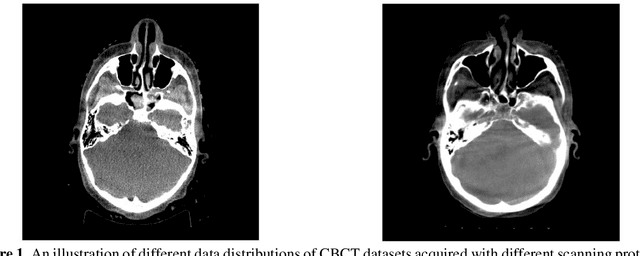
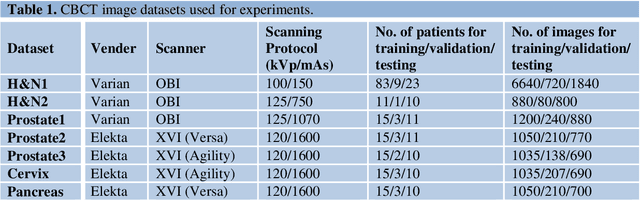
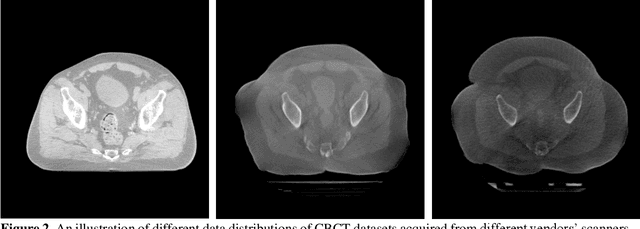
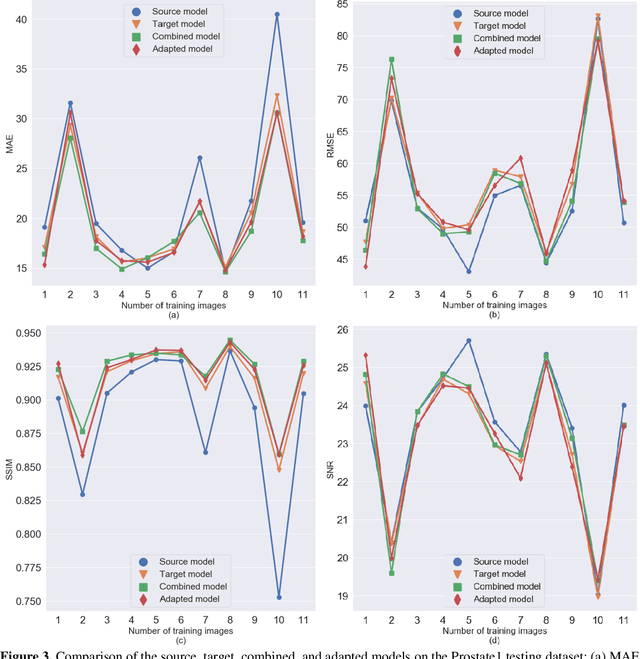
Abstract:Generalizability is a concern when applying a deep learning (DL) model trained on one dataset to other datasets. Training a universal model that works anywhere, anytime, for anybody is unrealistic. In this work, we demonstrate the generalizability problem, then explore potential solutions based on transfer learning (TL) by using the cone-beam computed tomography (CBCT) to computed tomography (CT) image conversion task as the testbed. Previous works have converted CBCT to CT-like images. However, all of those works studied only one or two anatomical sites and used images from the same vendor's scanners. Here, we investigated how a model trained for one machine and one anatomical site works on other machines and other sites. We trained a model on CBCT images acquired from one vendor's scanners for head and neck cancer patients and applied it to images from another vendor's scanners and for other disease sites. We found that generalizability could be a significant problem for this particular application when applying a trained DL model to datasets from another vendor's scanners. We then explored three practical solutions based on TL to solve this generalization problem: the target model, which is trained on a target domain from scratch; the combined model, which is trained on both source and target domain datasets from scratch; and the adapted model, which fine-tunes the trained source model to a target domain. We found that when there are sufficient data in the target domain, all three models can achieve good performance. When the target dataset is limited, the adapted model works the best, which indicates that using the fine-tuning strategy to adapt the trained model to an unseen target domain dataset is a viable and easy way to implement DL models in the clinic.
A reinforcement learning application of guided Monte Carlo Tree Search algorithm for beam orientation selection in radiation therapy
Apr 14, 2020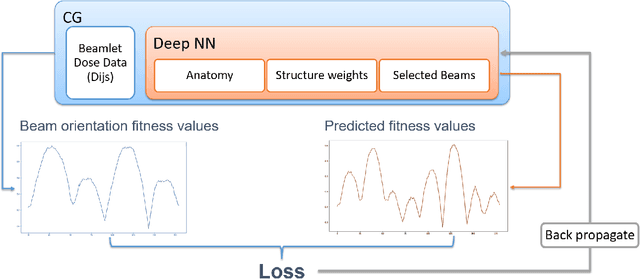
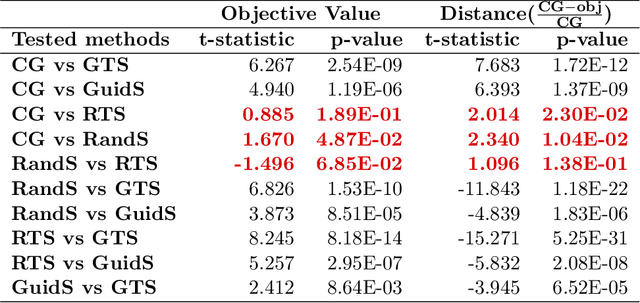
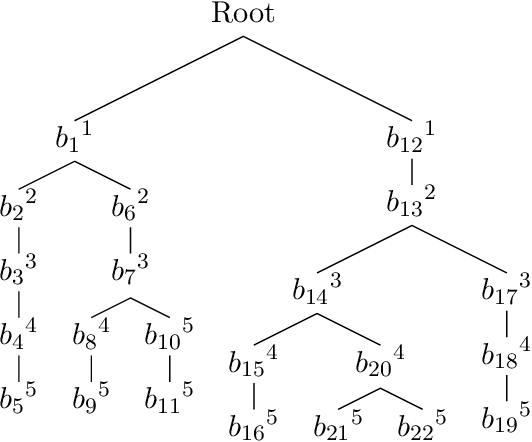
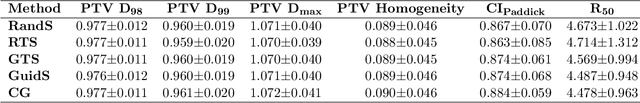
Abstract:Due to the large combinatorial problem, current beam orientation optimization algorithms for radiotherapy, such as column generation (CG), are typically heuristic or greedy in nature, leading to suboptimal solutions. We propose a reinforcement learning strategy using Monte Carlo Tree Search capable of finding a superior beam orientation set and in less time than CG.We utilized a reinforcement learning structure involving a supervised learning network to guide Monte Carlo tree search (GTS) to explore the decision space of beam orientation selection problem. We have previously trained a deep neural network (DNN) that takes in the patient anatomy, organ weights, and current beams, and then approximates beam fitness values, indicating the next best beam to add. This DNN is used to probabilistically guide the traversal of the branches of the Monte Carlo decision tree to add a new beam to the plan. To test the feasibility of the algorithm, we solved for 5-beam plans, using 13 test prostate cancer patients, different from the 57 training and validation patients originally trained the DNN. To show the strength of GTS to other search methods, performances of three other search methods including a guided search, uniform tree search and random search algorithms are also provided. On average GTS outperforms all other methods, it find a solution better than CG in 237 seconds on average, compared to CG which takes 360 seconds, and outperforms all other methods in finding a solution with lower objective function value in less than 1000 seconds. Using our guided tree search (GTS) method we were able to maintain a similar planning target volume (PTV) coverage within 1% error, and reduce the organ at risk (OAR) mean dose for body, rectum, left and right femoral heads, but a slight increase of 1% in bladder mean dose.
Mining Domain Knowledge: Improved Framework towards Automatically Standardizing Anatomical Structure Nomenclature in Radiotherapy
Dec 04, 2019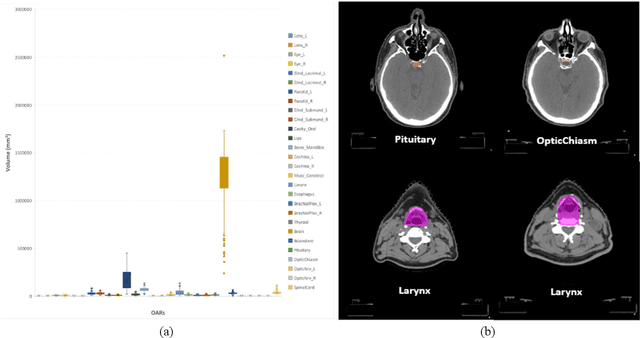
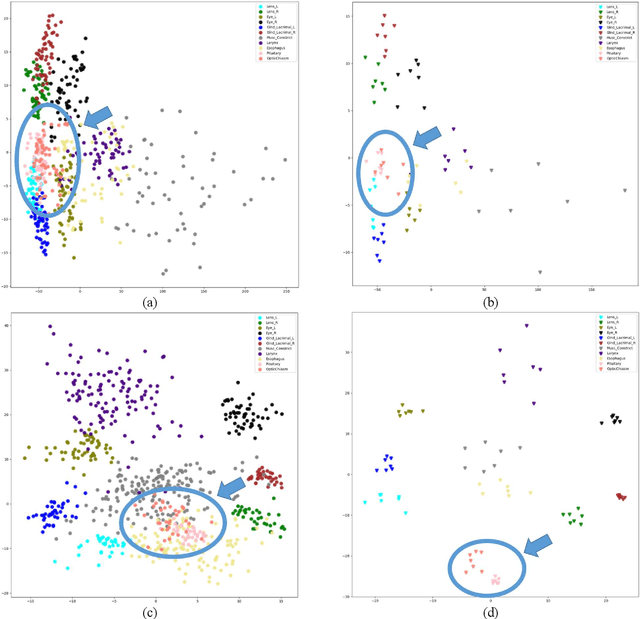
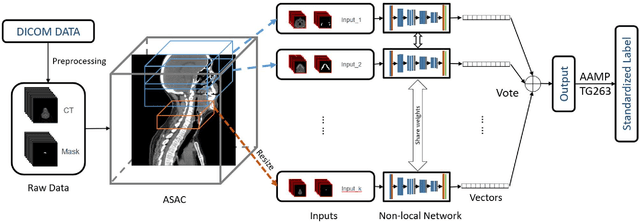
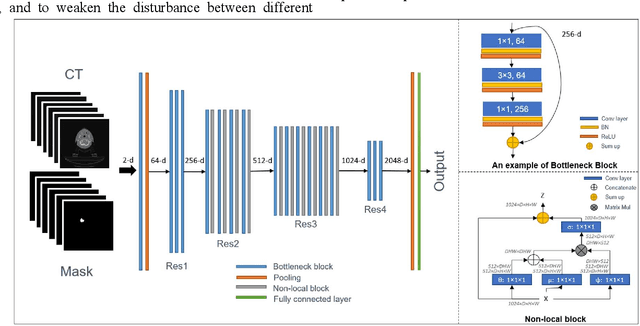
Abstract:Automatically standardizing nomenclature for anatomical structures in radiotherapy (RT) clinical data is an unmet urgent need in the era of big data and artificial intelligence. Existing methods either can hardly handle cross-institutional datasets or suffer from heavy imbalance and poor-quality delineation in clinical RT datasets. To solve these problems, we propose an automated structure nomenclature standardization framework, 3DNNV, which consists of an improved data processing strategy (ASAC/Voting) and an optimized feature extraction module to simulate clinicians' domain knowledge and recognition mechanisms to identify heavily imbalanced small-volume organs at risk (OARs) better than other methods. We used partial data from an open-source head-and-neck cancer dataset (HN_PETCT) to train the model, then tested the model on three cross-institutional datasets to demonstrate its generalizability. 3DNNV outperformed the baseline model (ResNet50), achieving a significantly higher average true positive rate (TPR) on the three test datasets (+8.27%, +2.39%, +5.53%). More importantly, the 3DNNV outperformed the baseline, 28.63% to 91.17%, on the F1 score of a small-volume OAR with only 9 training samples, when tested on the HN_UTSW dataset. The developed framework can be used to help standardizing structure nomenclature to facilitate data-driven clinical research in cancer radiotherapy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge