Simon Kelow
Tokenizing Loops of Antibodies
Sep 10, 2025



Abstract:The complementarity-determining regions of antibodies are loop structures that are key to their interactions with antigens, and of high importance to the design of novel biologics. Since the 1980s, categorizing the diversity of CDR structures into canonical clusters has enabled the identification of key structural motifs of antibodies. However, existing approaches have limited coverage and cannot be readily incorporated into protein foundation models. Here we introduce ImmunoGlobulin LOOp Tokenizer, Igloo, a multimodal antibody loop tokenizer that encodes backbone dihedral angles and sequence. Igloo is trained using a contrastive learning objective to map loops with similar backbone dihedral angles closer together in latent space. Igloo can efficiently retrieve the closest matching loop structures from a structural antibody database, outperforming existing methods on identifying similar H3 loops by 5.9\%. Igloo assigns tokens to all loops, addressing the limited coverage issue of canonical clusters, while retaining the ability to recover canonical loop conformations. To demonstrate the versatility of Igloo tokens, we show that they can be incorporated into protein language models with IglooLM and IglooALM. On predicting binding affinity of heavy chain variants, IglooLM outperforms the base protein language model on 8 out of 10 antibody-antigen targets. Additionally, it is on par with existing state-of-the-art sequence-based and multimodal protein language models, performing comparably to models with $7\times$ more parameters. IglooALM samples antibody loops which are diverse in sequence and more consistent in structure than state-of-the-art antibody inverse folding models. Igloo demonstrates the benefit of introducing multimodal tokens for antibody loops for encoding the diverse landscape of antibody loops, improving protein foundation models, and for antibody CDR design.
Multi-segment preserving sampling for deep manifold sampler
May 09, 2022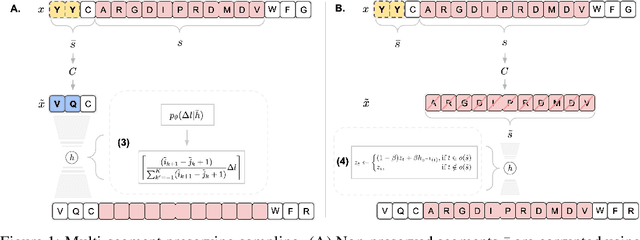
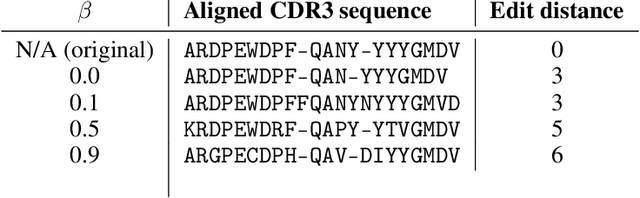

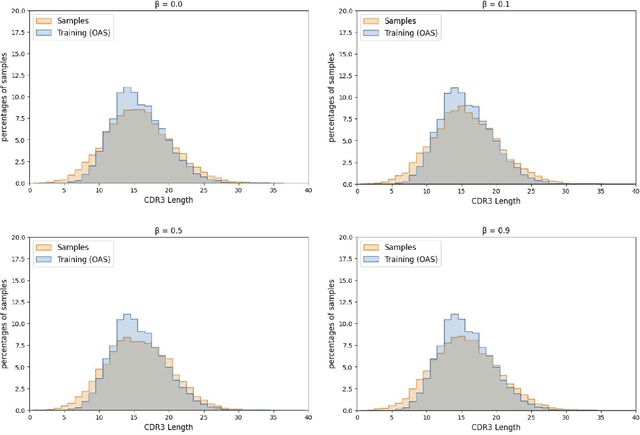
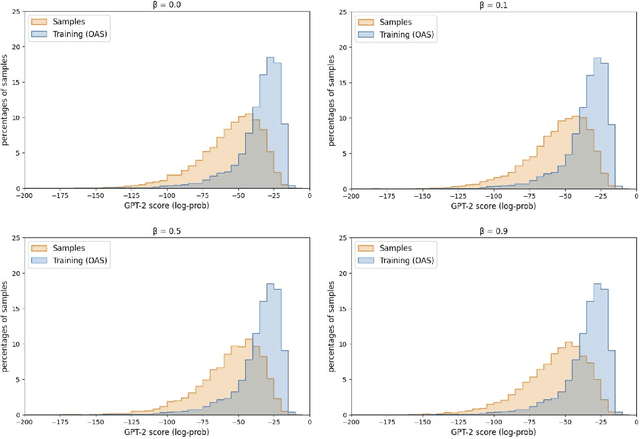
Abstract:Deep generative modeling for biological sequences presents a unique challenge in reconciling the bias-variance trade-off between explicit biological insight and model flexibility. The deep manifold sampler was recently proposed as a means to iteratively sample variable-length protein sequences by exploiting the gradients from a function predictor. We introduce an alternative approach to this guided sampling procedure, multi-segment preserving sampling, that enables the direct inclusion of domain-specific knowledge by designating preserved and non-preserved segments along the input sequence, thereby restricting variation to only select regions. We present its effectiveness in the context of antibody design by training two models: a deep manifold sampler and a GPT-2 language model on nearly six million heavy chain sequences annotated with the IGHV1-18 gene. During sampling, we restrict variation to only the complementarity-determining region 3 (CDR3) of the input. We obtain log probability scores from a GPT-2 model for each sampled CDR3 and demonstrate that multi-segment preserving sampling generates reasonable designs while maintaining the desired, preserved regions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge