Shinjae Yoo
A Mixture of Experts Foundation Model for Scanning Electron Microscopy Image Analysis
Apr 07, 2026Abstract:Scanning Electron Microscopy (SEM) is indispensable in modern materials science, enabling high-resolution imaging across a wide range of structural, chemical, and functional investigations. However, SEM imaging remains constrained by task-specific models and labor-intensive acquisition processes that limit its scalability across diverse applications. Here, we introduce the first foundation model for SEM images, pretrained on a large corpus of multi-instrument, multi-condition scientific micrographs, enabling generalization across diverse material systems and imaging conditions. Leveraging a self-supervised transformer architecture, our model learns rich and transferable representations that can be fine-tuned or adapted to a wide range of downstream tasks. As a compelling demonstration, we focus on defocus-to-focus image translation-an essential yet underexplored challenge in automated microscopy pipelines. Our method not only restores focused detail from defocused inputs without paired supervision but also outperforms state-of-the-art techniques across multiple evaluation metrics. This work lays the groundwork for a new class of adaptable SEM models, accelerating materials discovery by bridging foundational representation learning with real-world imaging needs.
Hybrid Quantum Temporal Convolutional Networks
Feb 27, 2026Abstract:Quantum machine learning models for sequential data face scalability challenges with complex multivariate signals. We introduce the Hybrid Quantum Temporal Convolutional Network (HQTCN), which combines classical temporal windowing with a quantum convolutional neural network core. By applying a shared quantum circuit across temporal windows, HQTCN captures long-range dependencies while achieving significant parameter reduction. Evaluated on synthetic NARMA sequences and high-dimensional EEG time-series, HQTCN performs competitively with classical baselines on univariate data and outperforms all baselines on multivariate tasks. The model demonstrates particular strength under data-limited conditions, maintaining high performance with substantially fewer parameters than conventional approaches. These results establish HQTCN as a parameter-efficient approach for multivariate time-series analysis.
DANCE: Doubly Adaptive Neighborhood Conformal Estimation
Feb 24, 2026Abstract:The recent developments of complex deep learning models have led to unprecedented ability to accurately predict across multiple data representation types. Conformal prediction for uncertainty quantification of these models has risen in popularity, providing adaptive, statistically-valid prediction sets. For classification tasks, conformal methods have typically focused on utilizing logit scores. For pre-trained models, however, this can result in inefficient, overly conservative set sizes when not calibrated towards the target task. We propose DANCE, a doubly locally adaptive nearest-neighbor based conformal algorithm combining two novel nonconformity scores directly using the data's embedded representation. DANCE first fits a task-adaptive kernel regression model from the embedding layer before using the learned kernel space to produce the final prediction sets for uncertainty quantification. We test against state-of-the-art local, task-adapted and zero-shot conformal baselines, demonstrating DANCE's superior blend of set size efficiency and robustness across various datasets.
Quantum Super-resolution by Adaptive Non-local Observables
Jan 20, 2026Abstract:Super-resolution (SR) seeks to reconstruct high-resolution (HR) data from low-resolution (LR) observations. Classical deep learning methods have advanced SR substantially, but require increasingly deeper networks, large datasets, and heavy computation to capture fine-grained correlations. In this work, we present the \emph{first study} to investigate quantum circuits for SR. We propose a framework based on Variational Quantum Circuits (VQCs) with \emph{Adaptive Non-Local Observable} (ANO) measurements. Unlike conventional VQCs with fixed Pauli readouts, ANO introduces trainable multi-qubit Hermitian observables, allowing the measurement process to adapt during training. This design leverages the high-dimensional Hilbert space of quantum systems and the representational structure provided by entanglement and superposition. Experiments demonstrate that ANO-VQCs achieve up to five-fold higher resolution with a relatively small model size, suggesting a promising new direction at the intersection of quantum machine learning and super-resolution.
DIVER-1 : Deep Integration of Vast Electrophysiological Recordings at Scale
Dec 22, 2025



Abstract:Electrophysiology signals such as EEG and iEEG are central to neuroscience, brain-computer interfaces, and clinical applications, yet existing foundation models remain limited in scale despite clear evidence that scaling improves performance. We introduce DIVER-1, a family of EEG and iEEG foundation models trained on the largest and most diverse corpus to date-5.3k hours of iEEG and 54k hours of EEG (1.6M channel-hours from over 17.7k subjects)-and scaled up to 1.82B parameters. We present the first systematic scaling law analysis for this domain, showing that they follow data-constrained scaling laws: for a given amount of data and compute, smaller models trained for extended epochs consistently outperform larger models trained briefly. This behavior contrasts with prior electrophysiology foundation models that emphasized model size over training duration. To achieve strong performance, we also design architectural innovations including any-variate attention, sliding temporal conditional positional encoding, and multi-domain reconstruction. DIVER-1 iEEG and EEG models each achieve state-of-the-art performance on their respective benchmarks, establishing a concrete guidelines for efficient scaling and resource allocation in electrophysiology foundation model development.
Maximal Update Parametrization and Zero-Shot Hyperparameter Transfer for Fourier Neural Operators
Jun 24, 2025Abstract:Fourier Neural Operators (FNOs) offer a principled approach for solving complex partial differential equations (PDEs). However, scaling them to handle more complex PDEs requires increasing the number of Fourier modes, which significantly expands the number of model parameters and makes hyperparameter tuning computationally impractical. To address this, we introduce $\mu$Transfer-FNO, a zero-shot hyperparameter transfer technique that enables optimal configurations, tuned on smaller FNOs, to be directly applied to billion-parameter FNOs without additional tuning. Building on the Maximal Update Parametrization ($\mu$P) framework, we mathematically derive a parametrization scheme that facilitates the transfer of optimal hyperparameters across models with different numbers of Fourier modes in FNOs, which is validated through extensive experiments on various PDEs. Our empirical study shows that Transfer-FNO reduces computational cost for tuning hyperparameters on large FNOs while maintaining or improving accuracy.
Towards an Introspective Dynamic Model of Globally Distributed Computing Infrastructures
Jun 24, 2025Abstract:Large-scale scientific collaborations like ATLAS, Belle II, CMS, DUNE, and others involve hundreds of research institutes and thousands of researchers spread across the globe. These experiments generate petabytes of data, with volumes soon expected to reach exabytes. Consequently, there is a growing need for computation, including structured data processing from raw data to consumer-ready derived data, extensive Monte Carlo simulation campaigns, and a wide range of end-user analysis. To manage these computational and storage demands, centralized workflow and data management systems are implemented. However, decisions regarding data placement and payload allocation are often made disjointly and via heuristic means. A significant obstacle in adopting more effective heuristic or AI-driven solutions is the absence of a quick and reliable introspective dynamic model to evaluate and refine alternative approaches. In this study, we aim to develop such an interactive system using real-world data. By examining job execution records from the PanDA workflow management system, we have pinpointed key performance indicators such as queuing time, error rate, and the extent of remote data access. The dataset includes five months of activity. Additionally, we are creating a generative AI model to simulate time series of payloads, which incorporate visible features like category, event count, and submitting group, as well as hidden features like the total computational load-derived from existing PanDA records and computing site capabilities. These hidden features, which are not visible to job allocators, whether heuristic or AI-driven, influence factors such as queuing times and data movement.
STACI: Spatio-Temporal Aleatoric Conformal Inference
May 27, 2025Abstract:Fitting Gaussian Processes (GPs) provides interpretable aleatoric uncertainty quantification for estimation of spatio-temporal fields. Spatio-temporal deep learning models, while scalable, typically assume a simplistic independent covariance matrix for the response, failing to capture the underlying correlation structure. However, spatio-temporal GPs suffer from issues of scalability and various forms of approximation bias resulting from restrictive assumptions of the covariance kernel function. We propose STACI, a novel framework consisting of a variational Bayesian neural network approximation of non-stationary spatio-temporal GP along with a novel spatio-temporal conformal inference algorithm. STACI is highly scalable, taking advantage of GPU training capabilities for neural network models, and provides statistically valid prediction intervals for uncertainty quantification. STACI outperforms competing GPs and deep methods in accurately approximating spatio-temporal processes and we show it easily scales to datasets with millions of observations.
Addressing the Current Challenges of Quantum Machine Learning through Multi-Chip Ensembles
May 13, 2025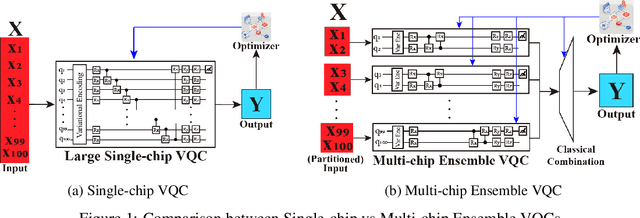
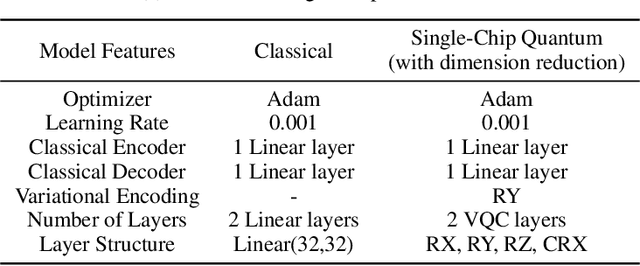
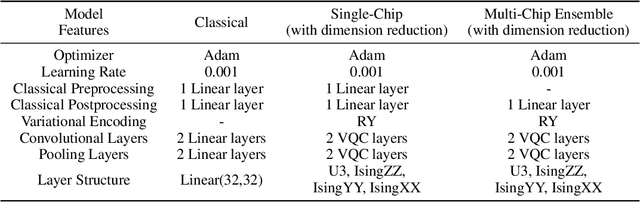

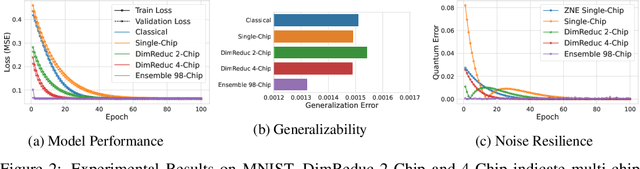

Abstract:Quantum Machine Learning (QML) holds significant promise for solving computational challenges across diverse domains. However, its practical deployment is constrained by the limitations of noisy intermediate-scale quantum (NISQ) devices, including noise, limited scalability, and trainability issues in variational quantum circuits (VQCs). We introduce the multi-chip ensemble VQC framework, which partitions high-dimensional computations across smaller quantum chips to enhance scalability, trainability, and noise resilience. We show that this approach mitigates barren plateaus, reduces quantum error bias and variance, and maintains robust generalization through controlled entanglement. Designed to align with current and emerging quantum hardware, the framework demonstrates strong potential for enabling scalable QML on near-term devices, as validated by experiments on standard benchmark datasets (MNIST, FashionMNIST, CIFAR-10) and real world dataset (PhysioNet EEG).
Adaptive Non-local Observable on Quantum Neural Networks
Apr 18, 2025Abstract:Conventional Variational Quantum Circuits (VQCs) for Quantum Machine Learning typically rely on a fixed Hermitian observable, often built from Pauli operators. Inspired by the Heisenberg picture, we propose an adaptive non-local measurement framework that substantially increases the model complexity of the quantum circuits. Our introduction of dynamical Hermitian observables with evolving parameters shows that optimizing VQC rotations corresponds to tracing a trajectory in the observable space. This viewpoint reveals that standard VQCs are merely a special case of the Heisenberg representation. Furthermore, we show that properly incorporating variational rotations with non-local observables enhances qubit interaction and information mixture, admitting flexible circuit designs. Two non-local measurement schemes are introduced, and numerical simulations on classification tasks confirm that our approach outperforms conventional VQCs, yielding a more powerful and resource-efficient approach as a Quantum Neural Network.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge