Shijia Wang
Climber-Pilot: A Non-Myopic Generative Recommendation Model Towards Better Instruction-Following
Feb 14, 2026Abstract:Generative retrieval has emerged as a promising paradigm in recommender systems, offering superior sequence modeling capabilities over traditional dual-tower architectures. However, in large-scale industrial scenarios, such models often suffer from inherent myopia: due to single-step inference and strict latency constraints, they tend to collapse diverse user intents into locally optimal predictions, failing to capture long-horizon and multi-item consumption patterns. Moreover, real-world retrieval systems must follow explicit retrieval instructions, such as category-level control and policy constraints. Incorporating such instruction-following behavior into generative retrieval remains challenging, as existing conditioning or post-hoc filtering approaches often compromise relevance or efficiency. In this work, we present Climber-Pilot, a unified generative retrieval framework to address both limitations. First, we introduce Time-Aware Multi-Item Prediction (TAMIP), a novel training paradigm designed to mitigate inherent myopia in generative retrieval. By distilling long-horizon, multi-item foresight into model parameters through time-aware masking, TAMIP alleviates locally optimal predictions while preserving efficient single-step inference. Second, to support flexible instruction-following retrieval, we propose Condition-Guided Sparse Attention (CGSA), which incorporates business constraints directly into the generative process via sparse attention, without introducing additional inference steps. Extensive offline experiments and online A/B testing at NetEase Cloud Music, one of the largest music streaming platforms, demonstrate that Climber-Pilot significantly outperforms state-of-the-art baselines, achieving a 4.24\% lift of the core business metric.
Shape Modeling with Spline Partitions
Aug 05, 2021
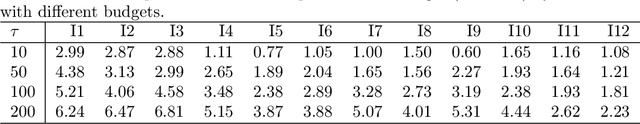
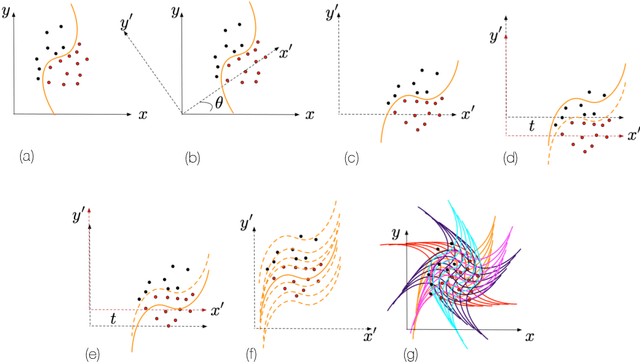
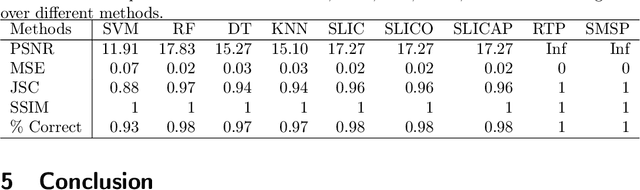
Abstract:Shape modelling (with methods that output shapes) is a new and important task in Bayesian nonparametrics and bioinformatics. In this work, we focus on Bayesian nonparametric methods for capturing shapes by partitioning a space using curves. In related work, the classical Mondrian process is used to partition spaces recursively with axis-aligned cuts, and is widely applied in multi-dimensional and relational data. The Mondrian process outputs hyper-rectangles. Recently, the random tessellation process was introduced as a generalization of the Mondrian process, partitioning a domain with non-axis aligned cuts in an arbitrary dimensional space, and outputting polytopes. Motivated by these processes, in this work, we propose a novel parallelized Bayesian nonparametric approach to partition a domain with curves, enabling complex data-shapes to be acquired. We apply our method to HIV-1-infected human macrophage image dataset, and also simulated datasets sets to illustrate our approach. We compare to support vector machines, random forests and state-of-the-art computer vision methods such as simple linear iterative clustering super pixel image segmentation. We develop an R package that is available at \url{https://github.com/ShufeiGe/Shape-Modeling-with-Spline-Partitions}.
Random Tessellation Forests
Jun 13, 2019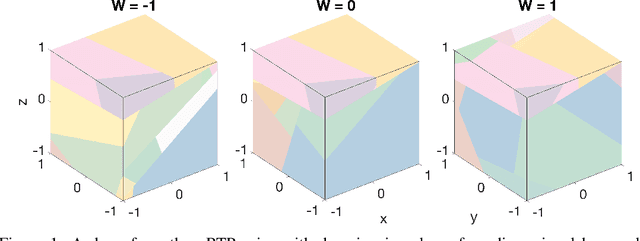
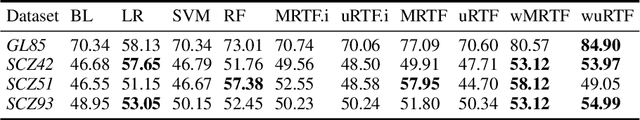

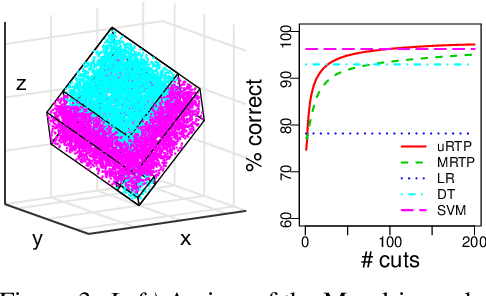
Abstract:Space partitioning methods such as random forests and the Mondrian process are powerful machine learning methods for multi-dimensional and relational data, and are based on recursively cutting a domain. The flexibility of these methods is often limited by the requirement that the cuts be axis aligned. The Ostomachion process and the self-consistent binary space partitioning-tree process were recently introduced as generalizations of the Mondrian process for space partitioning with non-axis aligned cuts in the two dimensional plane. Motivated by the need for a multi-dimensional partitioning tree with non-axis aligned cuts, we propose the Random Tessellation Process (RTP), a framework that includes the Mondrian process and the binary space partitioning-tree process as special cases. We derive a sequential Monte Carlo algorithm for inference, and provide random forest methods. Our process is self-consistent and can relax axis-aligned constraints, allowing complex inter-dimensional dependence to be captured. We present a simulation study, and analyse gene expression data of brain tissue, showing improved accuracies over other methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge