Shadi Albarqouni
University Hospital Bonn, Venusberg-Campus 1, D-53127, Bonn, Germany, Helmholtz Munich, Ingolstädter Landstraße 1, D-85764, Neuherberg, Germany, Technical University of Munich, Boltzmannstr. 3, D-85748 Garching, Germany
Semi-Supervised Few-Shot Learning with Local and Global Consistency
Mar 06, 2019
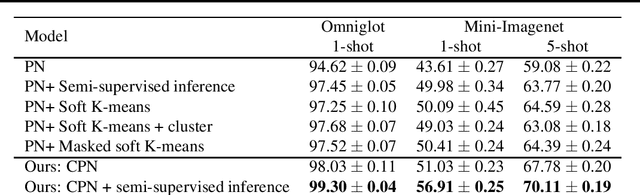
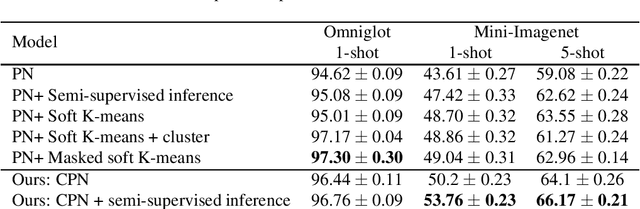
Abstract:Learning from a few examples is a key characteristic of human intelligence that AI researchers have been excited about modeling. With the web-scale data being mostly unlabeled, few recent works showed that few-shot learning performance can be significantly improved with access to unlabeled data, known as semi-supervised few shot learning (SS-FSL). We introduce a SS-FSL approach that we denote as Consistent Prototypical Networks (CPN), which builds on top of Prototypical Networks. We propose new loss terms to leverage unlabelled data, by enforcing notions of local and global consistency. Our work shows the effectiveness of our consistency losses in semi-supervised few shot setting. Our model outperforms the state-of-the-art in most benchmarks, showing large improvements in some cases. For example, in one mini-Imagenet 5-shot classification task, we obtain 70.1% accuracy to the 64.59% state-of-the-art. Moreover, our semi-supervised model, trained with 40% of the labels, compares well against the vanilla prototypical network trained on 100% of the labels, even outperforming it in the 1-shot mini-Imagenet case with 51.03% to 49.4% accuracy. For reproducibility, we make our code publicly available.
Towards an Interactive and Interpretable CAD System to Support Proximal Femur Fracture Classification
Feb 04, 2019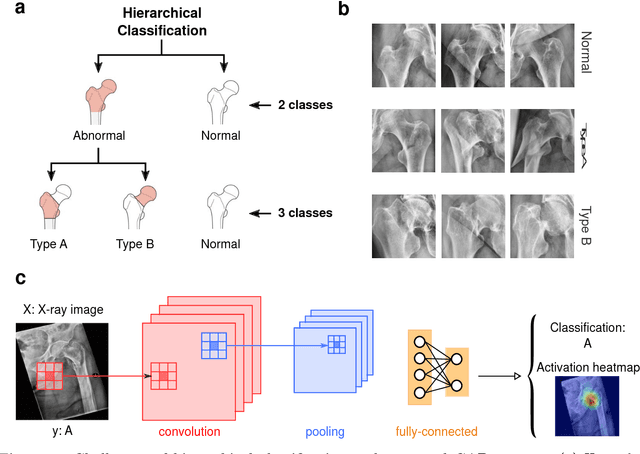
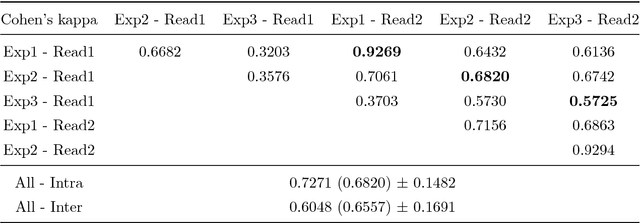
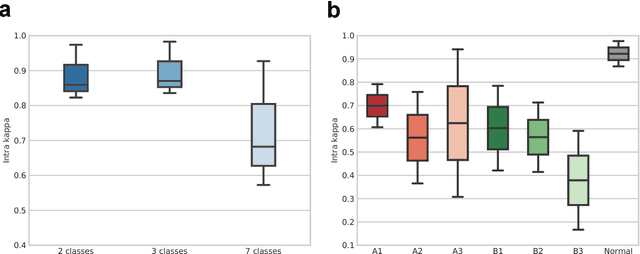
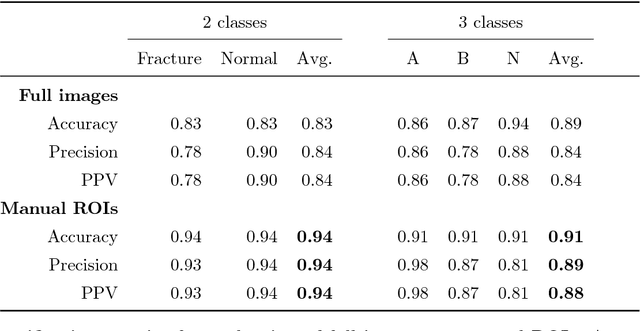
Abstract:Fractures of the proximal femur represent a critical entity in the western world, particularly with the growing elderly population. Such fractures result in high morbidity and mortality, reflecting a significant health and economic impact on our society. Different treatment strategies are recommended for different fracture types, with surgical treatment still being the gold standard in most of the cases. The success of the treatment and prognosis after surgery strongly depends on an accurate classification of the fracture among standard types, such as those defined by the AO system. However, the classification of fracture types based on x-ray images is difficult as confirmed by low intra- and inter-expert agreement rates of our in-house study and also in the previous literature. The presented work proposes a fully automatic computer-aided diagnosis (CAD) tool, based on current deep learning techniques, able to identify, localize and finally classify proximal femur fractures on x-rays images according to the AO classification. Results of our experimental evaluation show that the performance achieved by the proposed CAD tool is comparable to the average expert for the classification of x-ray images into types ''A'', ''B'' and ''normal'' (precision of 89%), while the performance is even superior when classifying fractures versus ''normal'' cases (precision of 94%). In addition, the integration of the proposed CAD tool into daily clinical routine is extensively discussed, towards improving the interface between humans and AI-powered machines in supporting medical decisions.
MRI to CT Translation with GANs
Jan 16, 2019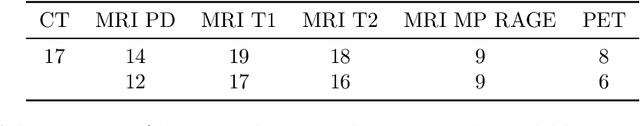
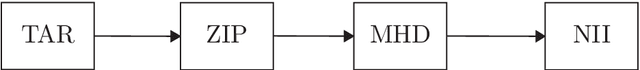

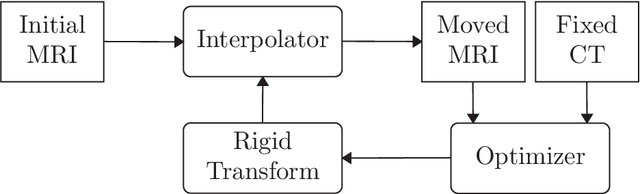
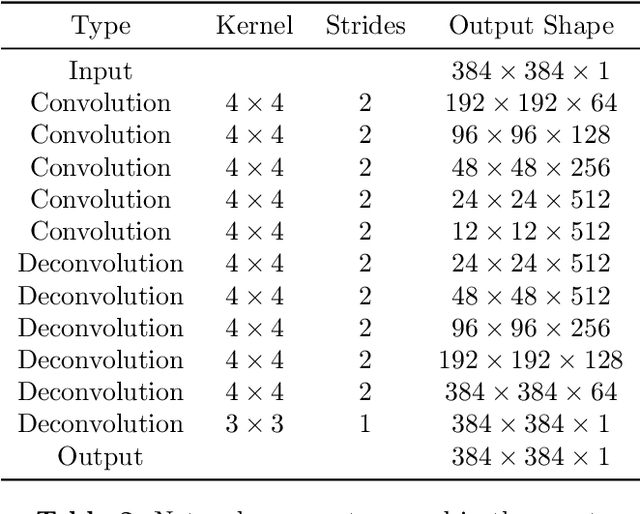
Abstract:We present a detailed description and reference implementation of preprocessing steps necessary to prepare the public Retrospective Image Registration Evaluation (RIRE) dataset for the task of magnetic resonance imaging (MRI) to X-ray computed tomography (CT) translation. Furthermore we describe and implement three state of the art convolutional neural network (CNN) and generative adversarial network (GAN) models where we report statistics and visual results of two of them.
Self-Attention Equipped Graph Convolutions for Disease Prediction
Dec 24, 2018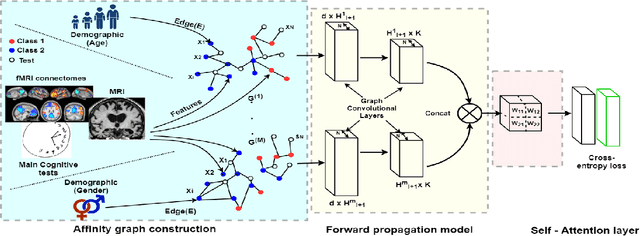
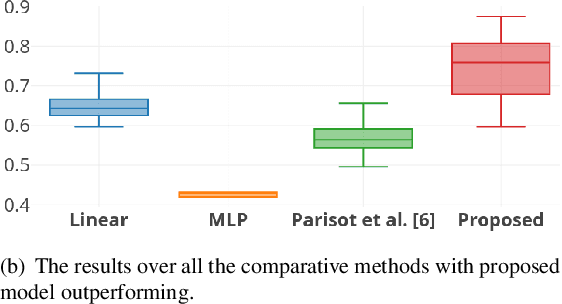
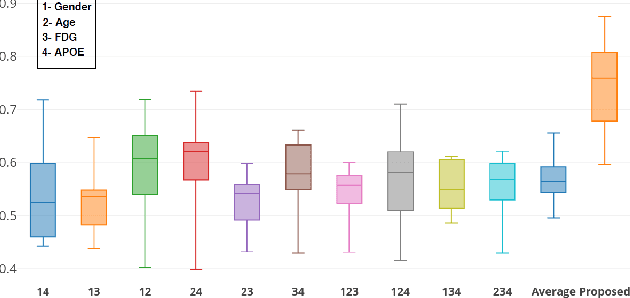
Abstract:Multi-modal data comprising imaging (MRI, fMRI, PET, etc.) and non-imaging (clinical test, demographics, etc.) data can be collected together and used for disease prediction. Such diverse data gives complementary information about the patient\'s condition to make an informed diagnosis. A model capable of leveraging the individuality of each multi-modal data is required for better disease prediction. We propose a graph convolution based deep model which takes into account the distinctiveness of each element of the multi-modal data. We incorporate a novel self-attention layer, which weights every element of the demographic data by exploring its relation to the underlying disease. We demonstrate the superiority of our developed technique in terms of computational speed and performance when compared to state-of-the-art methods. Our method outperforms other methods with a significant margin.
Virtualization of tissue staining in digital pathology using an unsupervised deep learning approach
Oct 15, 2018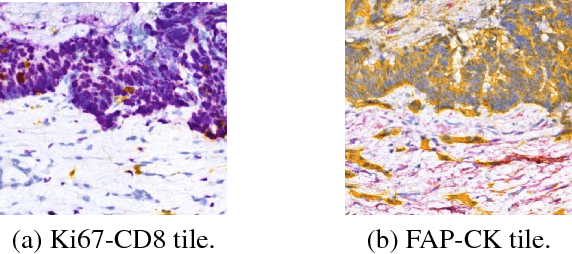
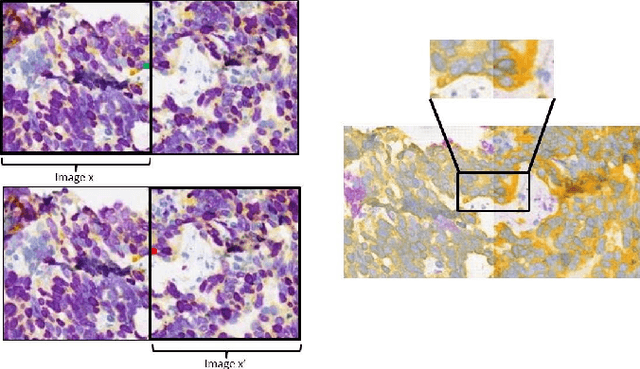

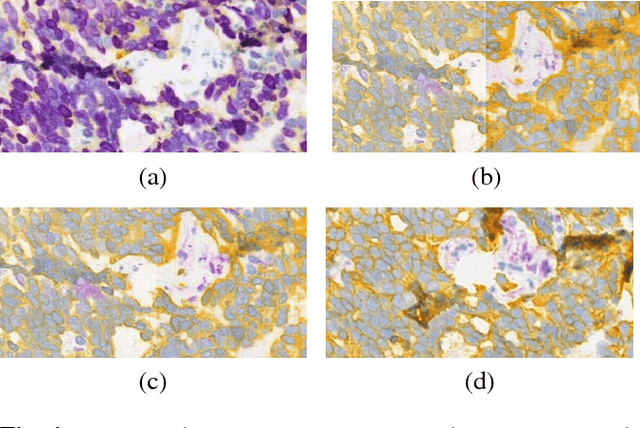
Abstract:Histopathological evaluation of tissue samples is a key practice in patient diagnosis and drug development, especially in oncology. Historically, Hematoxylin and Eosin (H&E) has been used by pathologists as a gold standard staining. However, in many cases, various target specific stains, including immunohistochemistry (IHC), are needed in order to highlight specific structures in the tissue. As tissue is scarce and staining procedures are tedious, it would be beneficial to generate images of stained tissue virtually. Virtual staining could also generate in-silico multiplexing of different stains on the same tissue segment. In this paper, we present a sample application that generates FAP-CK virtual IHC images from Ki67-CD8 real IHC images using an unsupervised deep learning approach based on CycleGAN. We also propose a method to deal with tiling artifacts caused by normalization layers and we validate our approach by comparing the results of tissue analysis algorithms for virtual and real images.
Weakly-Supervised Localization and Classification of Proximal Femur Fractures
Sep 27, 2018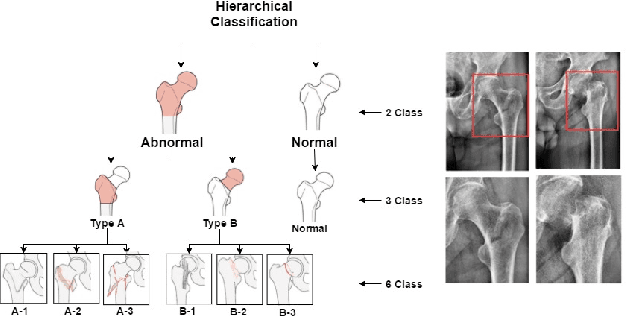
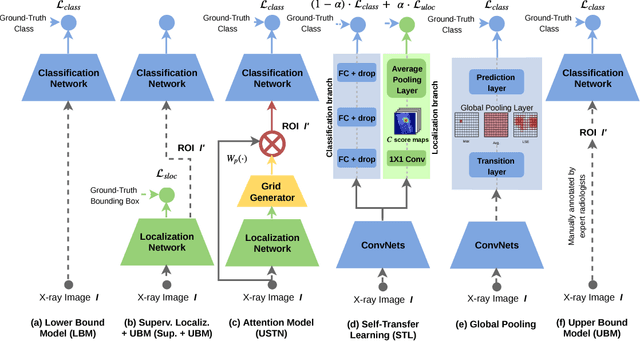
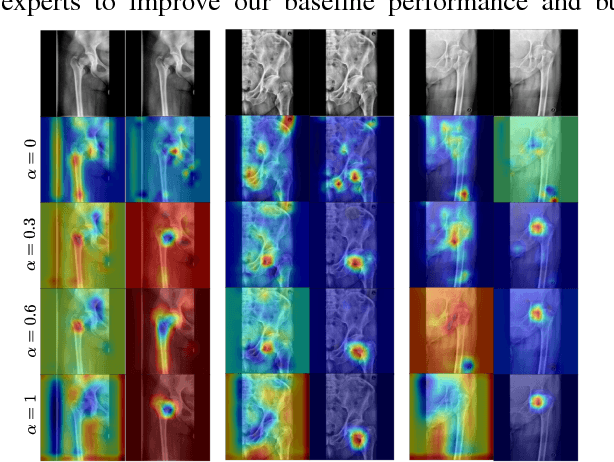
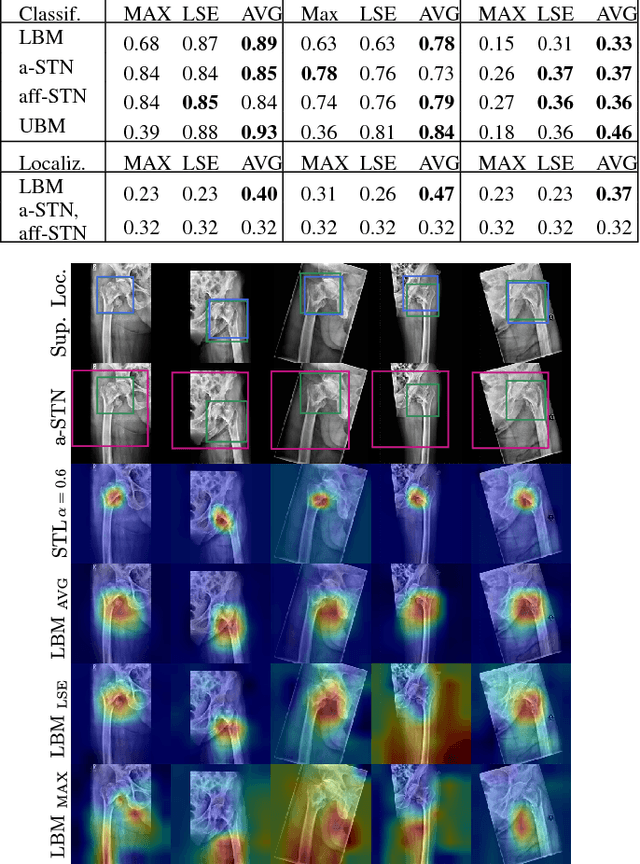
Abstract:In this paper, we target the problem of fracture classification from clinical X-Ray images towards an automated Computer Aided Diagnosis (CAD) system. Although primarily dealing with an image classification problem, we argue that localizing the fracture in the image is crucial to make good class predictions. Therefore, we propose and thoroughly analyze several schemes for simultaneous fracture localization and classification. We show that using an auxiliary localization task, in general, improves the classification performance. Moreover, it is possible to avoid the need for additional localization annotations thanks to recent advancements in weakly-supervised deep learning approaches. Among such approaches, we investigate and adapt Spatial Transformers (ST), Self-Transfer Learning (STL), and localization from global pooling layers. We provide a detailed quantitative and qualitative validation on a dataset of 1347 femur fractures images and report high accuracy with regard to inter-expert correlation values reported in the literature. Our investigations show that i) lesion localization improves the classification outcome, ii) weakly-supervised methods improve baseline classification without any additional cost, iii) STL guides feature activations and boost performance. We plan to make both the dataset and code available.
GANs for Medical Image Analysis
Sep 13, 2018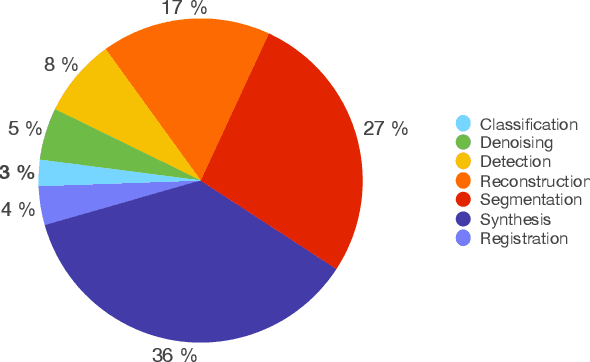

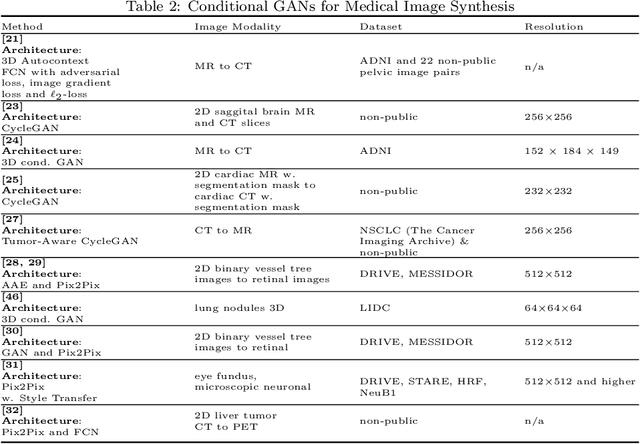
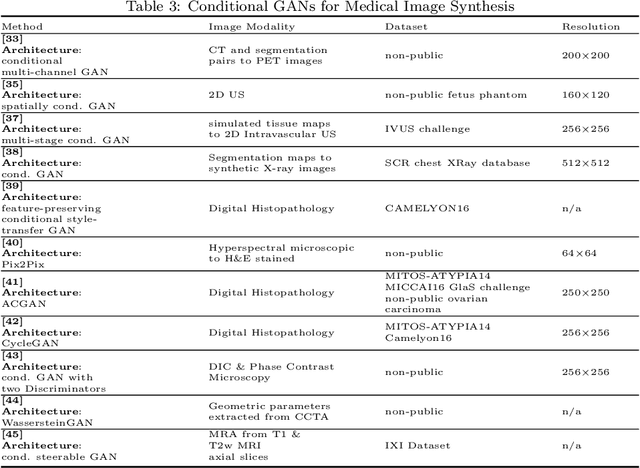
Abstract:Generative Adversarial Networks (GANs) and their extensions have carved open many exciting ways to tackle well known and challenging medical image analysis problems such as medical image denoising, reconstruction, segmentation, data simulation, detection or classification. Furthermore, their ability to synthesize images at unprecedented levels of realism also gives hope that the chronic scarcity of labeled data in the medical field can be resolved with the help of these generative models. In this review paper, a broad overview of recent literature on GANs for medical applications is given, the shortcomings and opportunities of the proposed methods are thoroughly discussed and potential future work is elaborated. A total of 63 papers published until end of July 2018 are reviewed. For quick access, the papers and important details such as the underlying method, datasets and performance are summarized in tables.
Generating Highly Realistic Images of Skin Lesions with GANs
Sep 06, 2018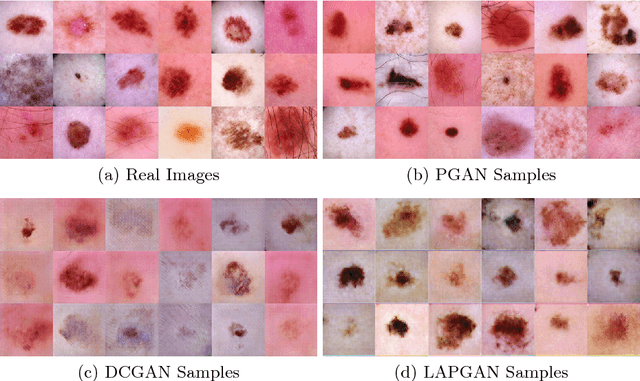
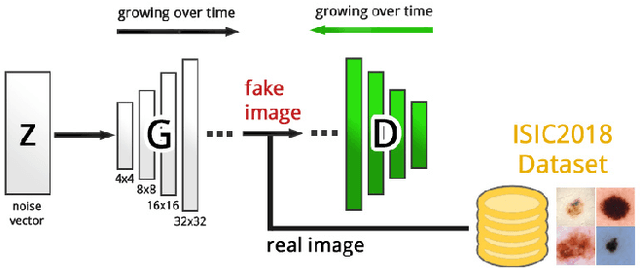
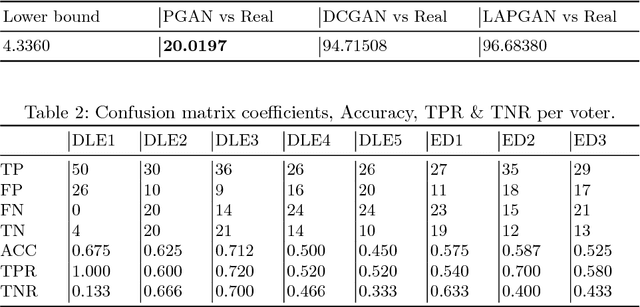
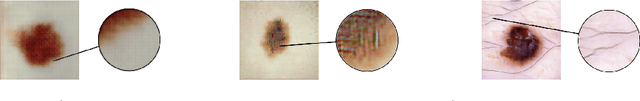
Abstract:As many other machine learning driven medical image analysis tasks, skin image analysis suffers from a chronic lack of labeled data and skewed class distributions, which poses problems for the training of robust and well-generalizing models. The ability to synthesize realistic looking images of skin lesions could act as a reliever for the aforementioned problems. Generative Adversarial Networks (GANs) have been successfully used to synthesize realistically looking medical images, however limited to low resolution, whereas machine learning models for challenging tasks such as skin lesion segmentation or classification benefit from much higher resolution data. In this work, we successfully synthesize realistically looking images of skin lesions with GANs at such high resolution. Therefore, we utilize the concept of progressive growing, which we both quantitatively and qualitatively compare to other GAN architectures such as the DCGAN and the LAPGAN. Our results show that with the help of progressive growing, we can synthesize highly realistic dermoscopic images of skin lesions that even expert dermatologists find hard to distinguish from real ones.
Scene Coordinate and Correspondence Learning for Image-Based Localization
Jul 26, 2018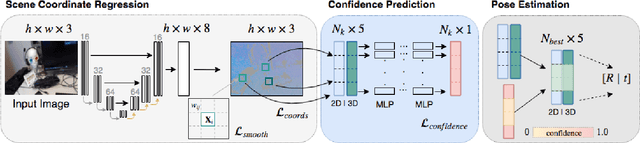
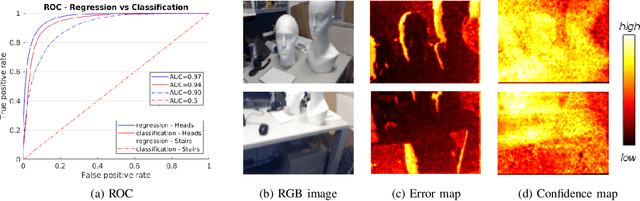
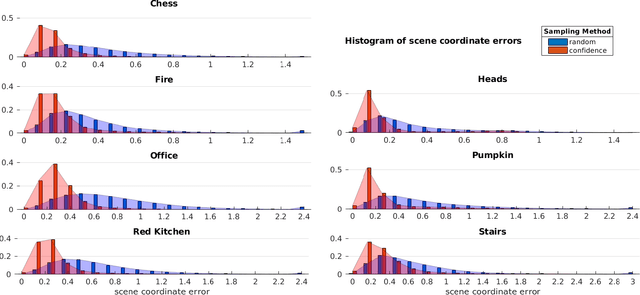

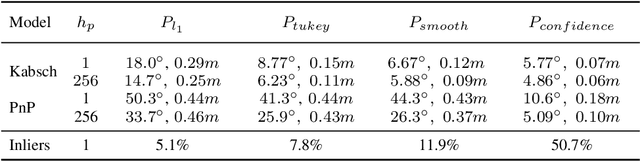
Abstract:Scene coordinate regression has become an essential part of current camera re-localization methods. Different versions, such as regression forests and deep learning methods, have been successfully applied to estimate the corresponding camera pose given a single input image. In this work, we propose to regress the scene coordinates pixel-wise for a given RGB image by using deep learning. Compared to the recent methods, which usually employ RANSAC to obtain a robust pose estimate from the established point correspondences, we propose to regress confidences of these correspondences, which allows us to immediately discard erroneous predictions and improve the initial pose estimates. Finally, the resulting confidences can be used to score initial pose hypothesis and aid in pose refinement, offering a generalized solution to solve this task.
Capsule Networks against Medical Imaging Data Challenges
Jul 19, 2018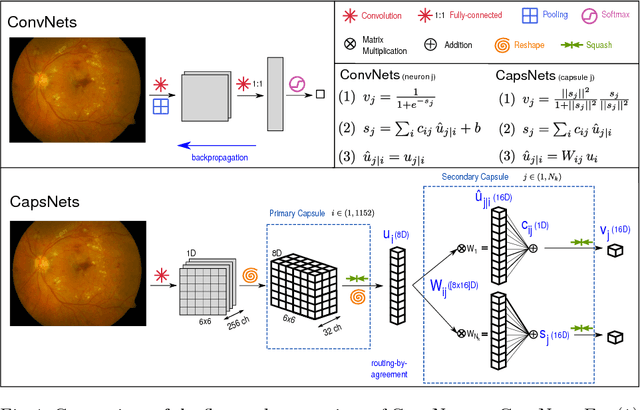
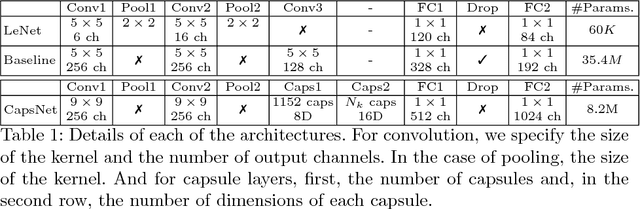
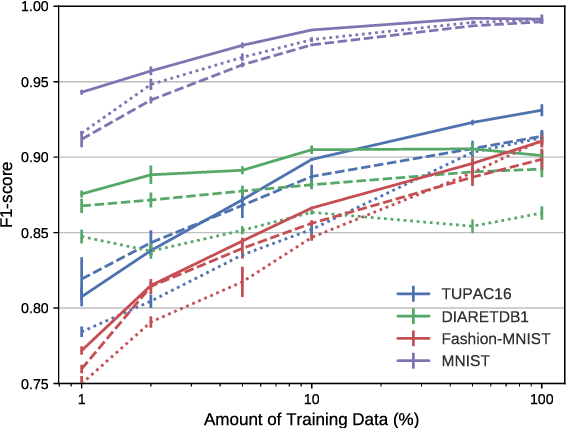
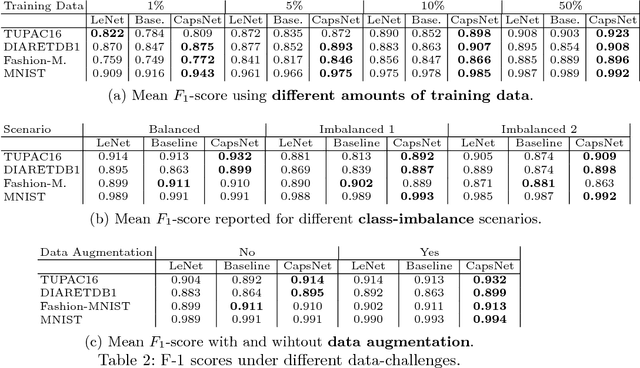
Abstract:A key component to the success of deep learning is the availability of massive amounts of training data. Building and annotating large datasets for solving medical image classification problems is today a bottleneck for many applications. Recently, capsule networks were proposed to deal with shortcomings of Convolutional Neural Networks (ConvNets). In this work, we compare the behavior of capsule networks against ConvNets under typical datasets constraints of medical image analysis, namely, small amounts of annotated data and class-imbalance. We evaluate our experiments on MNIST, Fashion-MNIST and medical (histological and retina images) publicly available datasets. Our results suggest that capsule networks can be trained with less amount of data for the same or better performance and are more robust to an imbalanced class distribution, which makes our approach very promising for the medical imaging community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge