Sandeep Konam
ASR Error Detection via Audio-Transcript entailment
Jul 22, 2022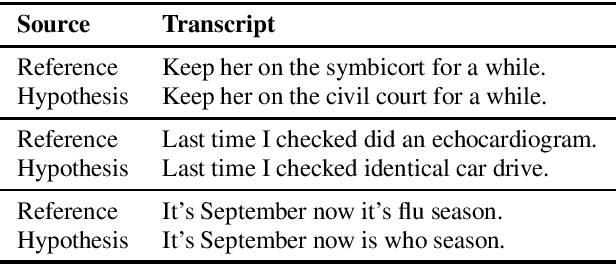

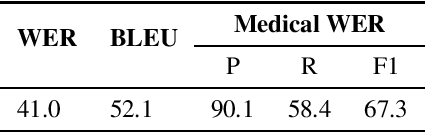
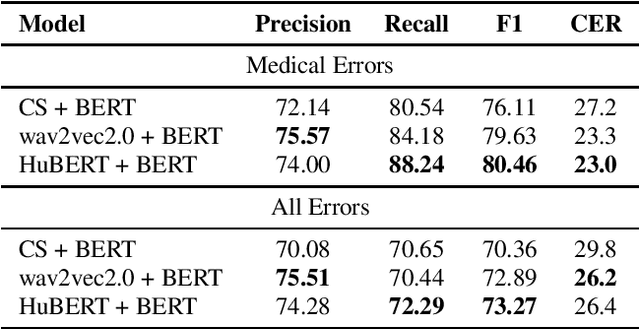
Abstract:Despite improved performances of the latest Automatic Speech Recognition (ASR) systems, transcription errors are still unavoidable. These errors can have a considerable impact in critical domains such as healthcare, when used to help with clinical documentation. Therefore, detecting ASR errors is a critical first step in preventing further error propagation to downstream applications. To this end, we propose a novel end-to-end approach for ASR error detection using audio-transcript entailment. To the best of our knowledge, we are the first to frame this problem as an end-to-end entailment task between the audio segment and its corresponding transcript segment. Our intuition is that there should be a bidirectional entailment between audio and transcript when there is no recognition error and vice versa. The proposed model utilizes an acoustic encoder and a linguistic encoder to model the speech and transcript respectively. The encoded representations of both modalities are fused to predict the entailment. Since doctor-patient conversations are used in our experiments, a particular emphasis is placed on medical terms. Our proposed model achieves classification error rates (CER) of 26.2% on all transcription errors and 23% on medical errors specifically, leading to improvements upon a strong baseline by 12% and 15.4%, respectively.
Towards Fairness in Classifying Medical Conversations into SOAP Sections
Dec 02, 2020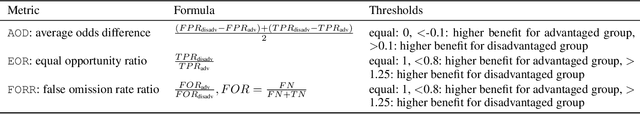

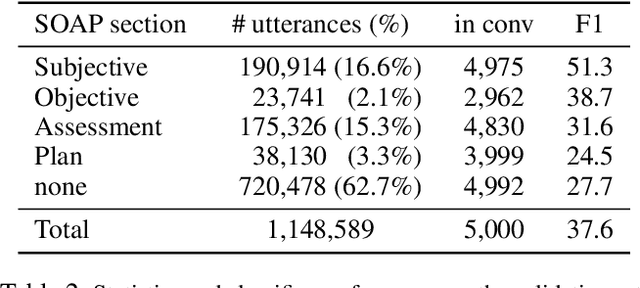
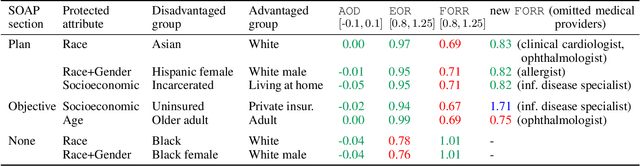
Abstract:As machine learning algorithms are more widely deployed in healthcare, the question of algorithmic fairness becomes more critical to examine. Our work seeks to identify and understand disparities in a deployed model that classifies doctor-patient conversations into sections of a medical SOAP note. We employ several metrics to measure disparities in the classifier performance, and find small differences in a portion of the disadvantaged groups. A deeper analysis of the language in these conversations and further stratifying the groups suggests these differences are related to and often attributable to the type of medical appointment (e.g., psychiatric vs. internist). Our findings stress the importance of understanding the disparities that may exist in the data itself and how that affects a model's ability to equally distribute benefits.
Weakly Supervised Medication Regimen Extraction from Medical Conversations
Oct 11, 2020
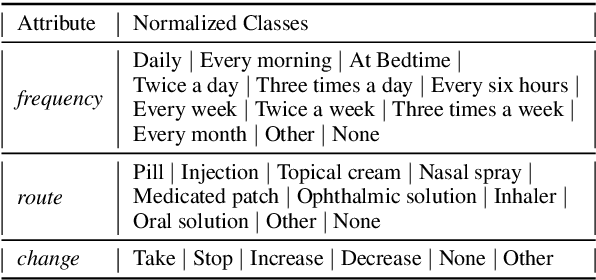

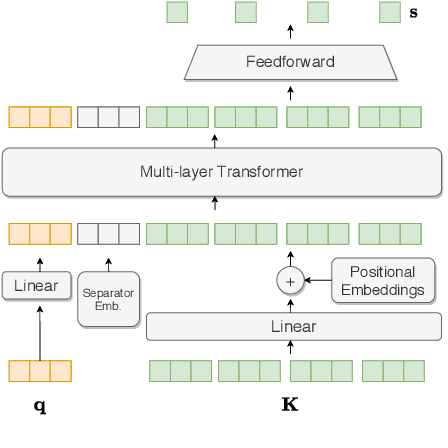
Abstract:Automated Medication Regimen (MR) extraction from medical conversations can not only improve recall and help patients follow through with their care plan, but also reduce the documentation burden for doctors. In this paper, we focus on extracting spans for frequency, route and change, corresponding to medications discussed in the conversation. We first describe a unique dataset of annotated doctor-patient conversations and then present a weakly supervised model architecture that can perform span extraction using noisy classification data. The model utilizes an attention bottleneck inside a classification model to perform the extraction. We experiment with several variants of attention scoring and projection functions and propose a novel transformer-based attention scoring function (TAScore). The proposed combination of TAScore and Fusedmax projection achieves a 10 point increase in Longest Common Substring F1 compared to the baseline of additive scoring plus softmax projection.
Towards an Automated SOAP Note: Classifying Utterances from Medical Conversations
Jul 27, 2020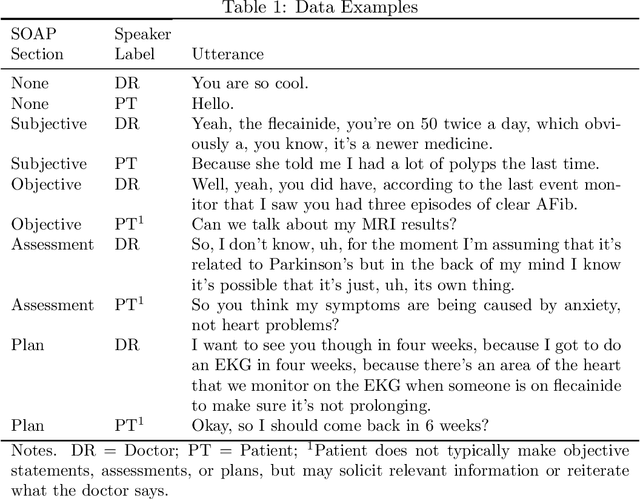
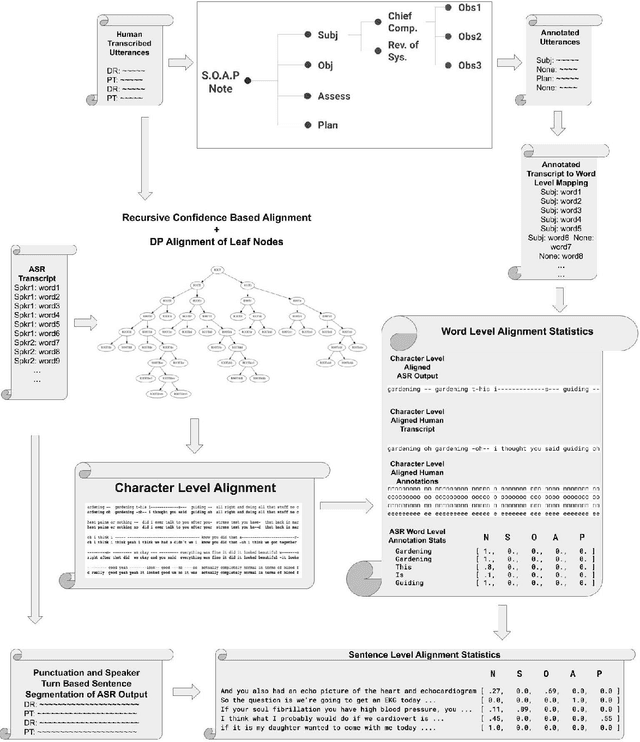

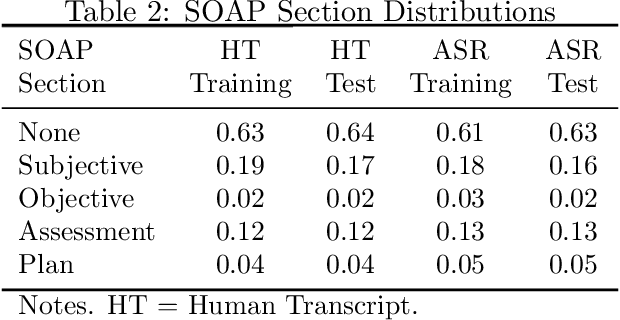
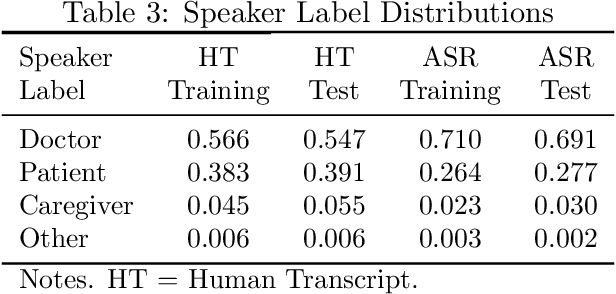
Abstract:Summaries generated from medical conversations can improve recall and understanding of care plans for patients and reduce documentation burden for doctors. Recent advancements in automatic speech recognition (ASR) and natural language understanding (NLU) offer potential solutions to generate these summaries automatically, but rigorous quantitative baselines for benchmarking research in this domain are lacking. In this paper, we bridge this gap for two tasks: classifying utterances from medical conversations according to (i) the SOAP section and (ii) the speaker role. Both are fundamental building blocks along the path towards an end-to-end, automated SOAP note for medical conversations. We provide details on a dataset that contains human and ASR transcriptions of medical conversations and corresponding machine learning optimized SOAP notes. We then present a systematic analysis in which we adapt an existing deep learning architecture to the two aforementioned tasks. The results suggest that modelling context in a hierarchical manner, which captures both word and utterance level context, yields substantial improvements on both classification tasks. Additionally, we develop and analyze a modular method for adapting our model to ASR output.
ASR Error Correction and Domain Adaptation Using Machine Translation
Mar 13, 2020
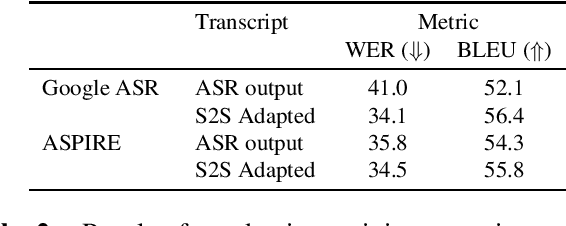
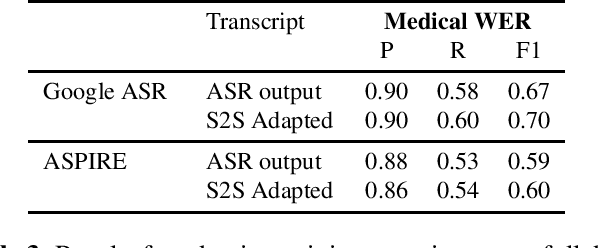
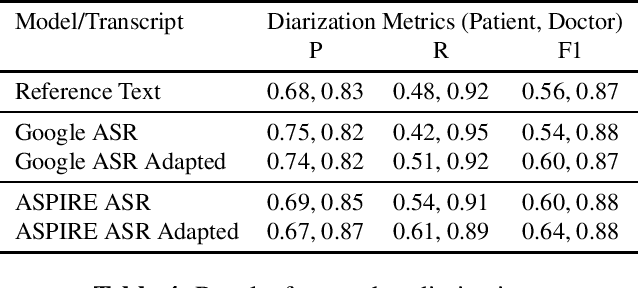
Abstract:Off-the-shelf pre-trained Automatic Speech Recognition (ASR) systems are an increasingly viable service for companies of any size building speech-based products. While these ASR systems are trained on large amounts of data, domain mismatch is still an issue for many such parties that want to use this service as-is leading to not so optimal results for their task. We propose a simple technique to perform domain adaptation for ASR error correction via machine translation. The machine translation model is a strong candidate to learn a mapping from out-of-domain ASR errors to in-domain terms in the corresponding reference files. We use two off-the-shelf ASR systems in this work: Google ASR (commercial) and the ASPIRE model (open-source). We observe 7% absolute improvement in word error rate and 4 point absolute improvement in BLEU score in Google ASR output via our proposed method. We also evaluate ASR error correction via a downstream task of Speaker Diarization that captures speaker style, syntax, structure and semantic improvements we obtain via ASR correction.
Medication Regimen Extraction From Medical Conversations
Jan 03, 2020
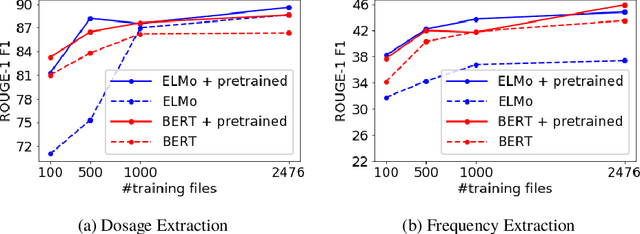
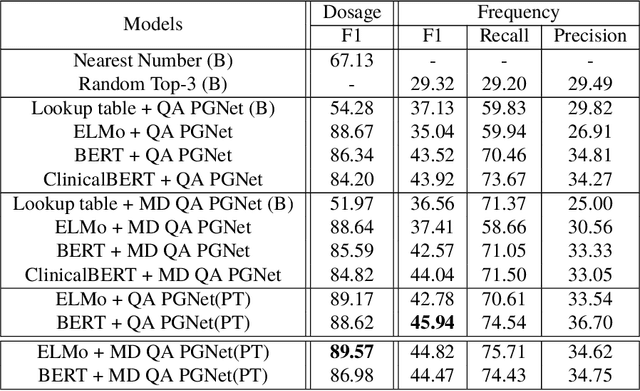
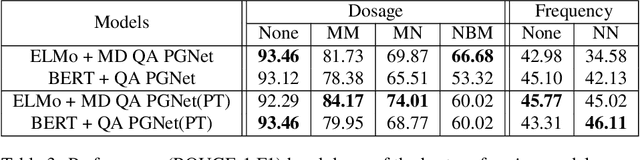
Abstract:Extracting relevant information from medical conversations and providing it to doctors and patients might help in addressing doctor burnout and patient forgetfulness. In this paper, we focus on extracting the Medication Regimen (dosage and frequency for medications) discussed in a medical conversation. We frame the problem as a Question Answering (QA) task and perform comparative analysis over: a QA approach, a new combined QA and Information Extraction approach, and other baselines. We use a small corpus of 6,692 annotated doctor-patient conversations for the task. Clinical conversation corpora are costly to create, difficult to handle (because of data privacy concerns), and thus scarce. We address this data scarcity challenge through data augmentation methods, using publicly available embeddings and pretrain part of the network on a related task (summarization) to improve the model's performance. Compared to the baseline, our best-performing models improve the dosage and frequency extractions' ROUGE-1 F1 scores from 54.28 and 37.13 to 89.57 and 45.94, respectively. Using our best-performing model, we present the first fully automated system that can extract Medication Regimen tags from spontaneous doctor-patient conversations with about ~71% accuracy.
Understanding Convolutional Networks with APPLE : Automatic Patch Pattern Labeling for Explanation
Feb 11, 2018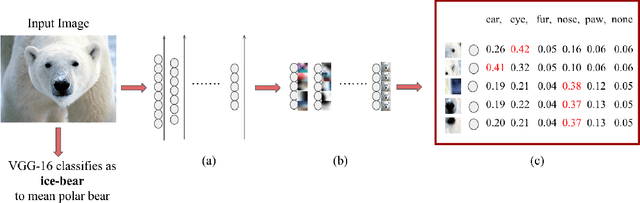
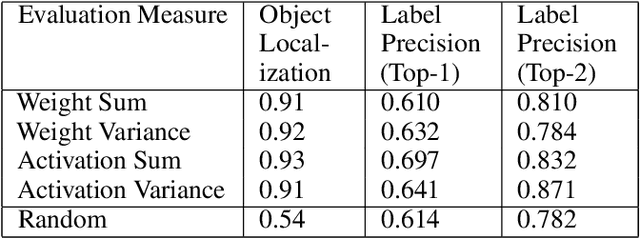


Abstract:With the success of deep learning, recent efforts have been focused on analyzing how learned networks make their classifications. We are interested in analyzing the network output based on the network structure and information flow through the network layers. We contribute an algorithm for 1) analyzing a deep network to find neurons that are 'important' in terms of the network classification outcome, and 2)automatically labeling the patches of the input image that activate these important neurons. We propose several measures of importance for neurons and demonstrate that our technique can be used to gain insight into, and explain how a network decomposes an image to make its final classification.
UAV and Service Robot Coordination for Indoor Object Search Tasks
Sep 26, 2017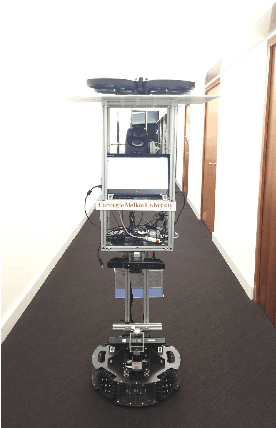
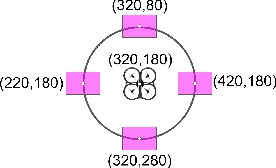
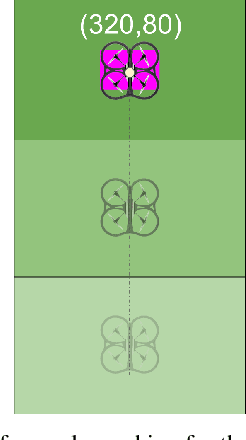
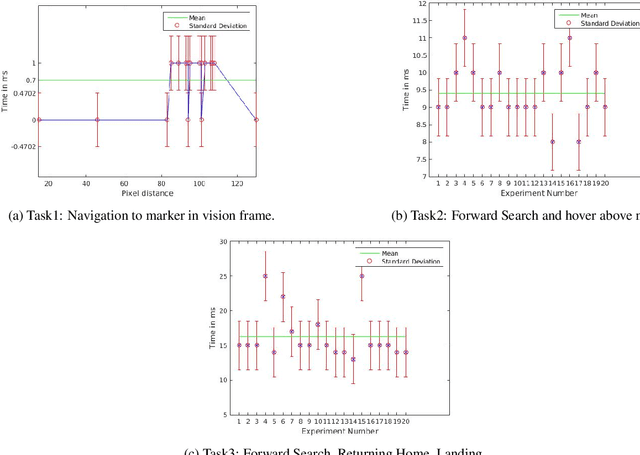
Abstract:Our CoBot robots have successfully performed a variety of service tasks in our multi-building environment including accompanying people to meetings and delivering objects to offices due to its navigation and localization capabilities. However, they lack the capability to visually search over desks and other confined locations for an object of interest. Conversely, an inexpensive GPS-denied quadcopter platform such as the Parrot ARDrone 2.0 could perform this object search task if it had access to reasonable localization. In this paper, we propose the concept of coordination between CoBot and the Parrot ARDrone 2.0 to perform service-based object search tasks, in which CoBot localizes and navigates to the general search areas carrying the ARDrone and the ARDrone searches locally for objects. We propose a vision-based moving target navigation algorithm that enables the ARDrone to localize with respect to CoBot, search for objects, and return to the CoBot for future searches. We demonstrate our algorithm in indoor environments on several search trajectories.
Towards Visual Explanations for Convolutional Neural Networks via Input Resampling
Aug 16, 2017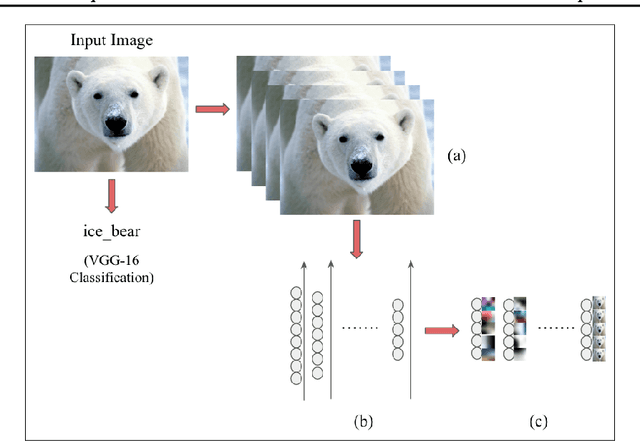
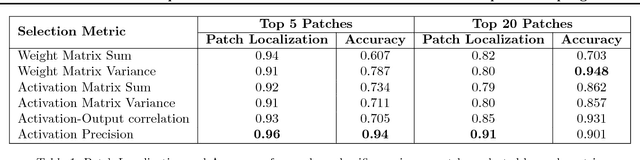
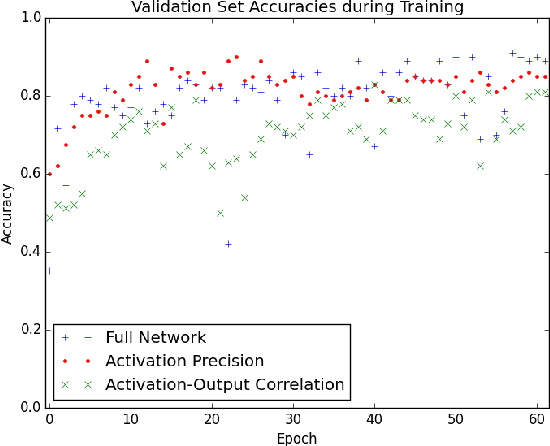
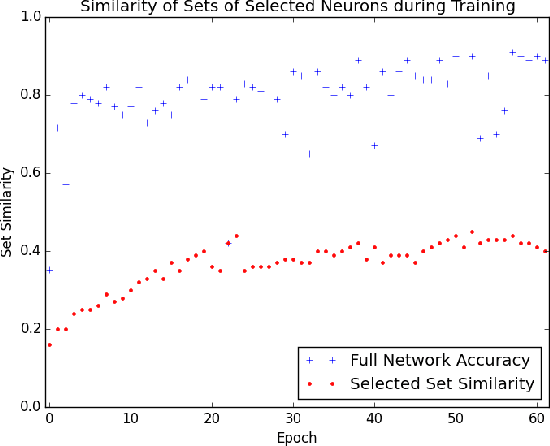
Abstract:The predictive power of neural networks often costs model interpretability. Several techniques have been developed for explaining model outputs in terms of input features; however, it is difficult to translate such interpretations into actionable insight. Here, we propose a framework to analyze predictions in terms of the model's internal features by inspecting information flow through the network. Given a trained network and a test image, we select neurons by two metrics, both measured over a set of images created by perturbations to the input image: (1) magnitude of the correlation between the neuron activation and the network output and (2) precision of the neuron activation. We show that the former metric selects neurons that exert large influence over the network output while the latter metric selects neurons that activate on generalizable features. By comparing the sets of neurons selected by these two metrics, our framework suggests a way to investigate the internal attention mechanisms of convolutional neural networks.
Obstacle Avoidance through Deep Networks based Intermediate Perception
Apr 27, 2017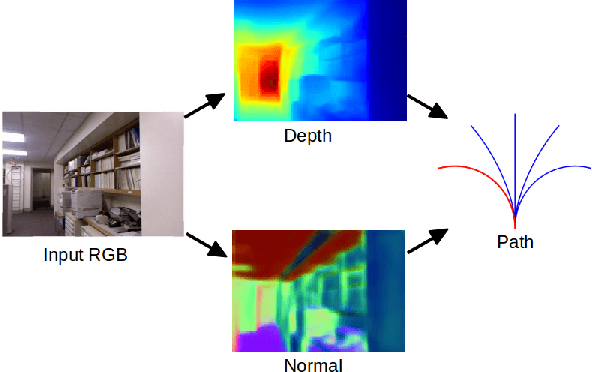
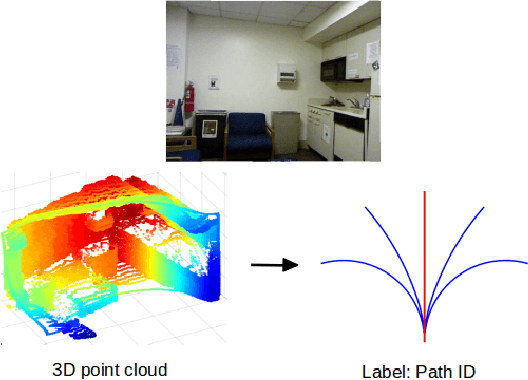
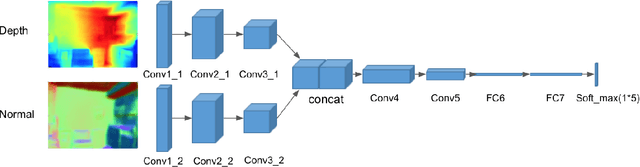
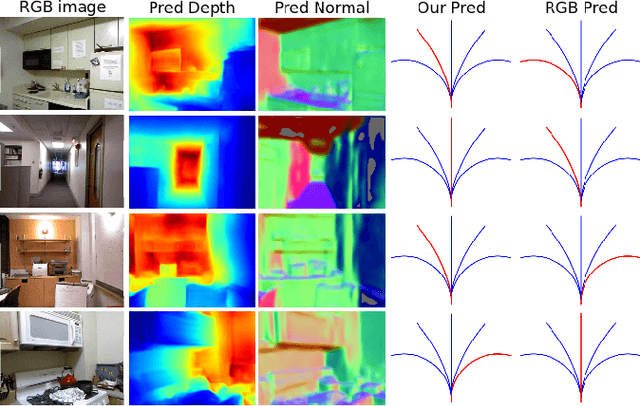
Abstract:Obstacle avoidance from monocular images is a challenging problem for robots. Though multi-view structure-from-motion could build 3D maps, it is not robust in textureless environments. Some learning based methods exploit human demonstration to predict a steering command directly from a single image. However, this method is usually biased towards certain tasks or demonstration scenarios and also biased by human understanding. In this paper, we propose a new method to predict a trajectory from images. We train our system on more diverse NYUv2 dataset. The ground truth trajectory is computed from the designed cost functions automatically. The Convolutional Neural Network perception is divided into two stages: first, predict depth map and surface normal from RGB images, which are two important geometric properties related to 3D obstacle representation. Second, predict the trajectory from the depth and normal. Results show that our intermediate perception increases the accuracy by 20% than the direct prediction. Our model generalizes well to other public indoor datasets and is also demonstrated for robot flights in simulation and experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge