Sabrina Maniscalco
Quantum computing and artificial intelligence: status and perspectives
May 29, 2025


Abstract:This white paper discusses and explores the various points of intersection between quantum computing and artificial intelligence (AI). It describes how quantum computing could support the development of innovative AI solutions. It also examines use cases of classical AI that can empower research and development in quantum technologies, with a focus on quantum computing and quantum sensing. The purpose of this white paper is to provide a long-term research agenda aimed at addressing foundational questions about how AI and quantum computing interact and benefit one another. It concludes with a set of recommendations and challenges, including how to orchestrate the proposed theoretical work, align quantum AI developments with quantum hardware roadmaps, estimate both classical and quantum resources - especially with the goal of mitigating and optimizing energy consumption - advance this emerging hybrid software engineering discipline, and enhance European industrial competitiveness while considering societal implications.
Disease Gene Prioritization With Quantum Walks
Nov 09, 2023



Abstract:Disease gene prioritization assigns scores to genes or proteins according to their likely relevance for a given disease based on a provided set of seed genes. Here, we describe a new algorithm for disease gene prioritization based on continuous-time quantum walks using the adjacency matrix of a protein-protein interaction (PPI) network. Our algorithm can be seen as a quantum version of a previous method known as the diffusion kernel, but, importantly, has higher performance in predicting disease genes, and also permits the encoding of seed node self-loops into the underlying Hamiltonian, which offers yet another boost in performance. We demonstrate the success of our proposed method by comparing it to several well-known gene prioritization methods on three disease sets, across seven different PPI networks. In order to compare these methods, we use cross-validation and examine the mean reciprocal ranks and recall values. We further validate our method by performing an enrichment analysis of the predicted genes for coronary artery disease. We also investigate the impact of adding self-loops to the seeds, and argue that they allow the quantum walker to remain more local to low-degree seed nodes.
Link prediction with continuous-time classical and quantum walks
Aug 23, 2022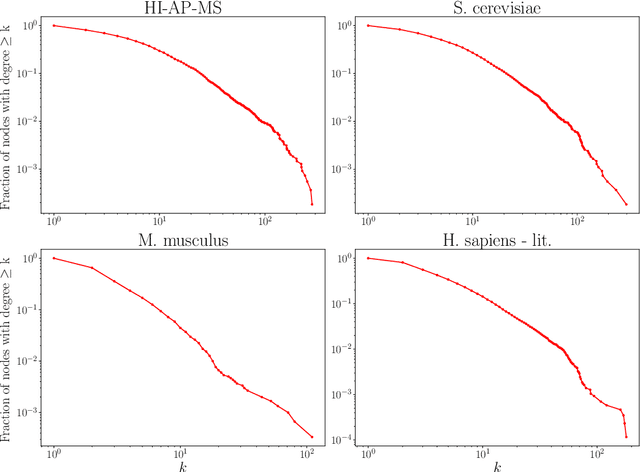
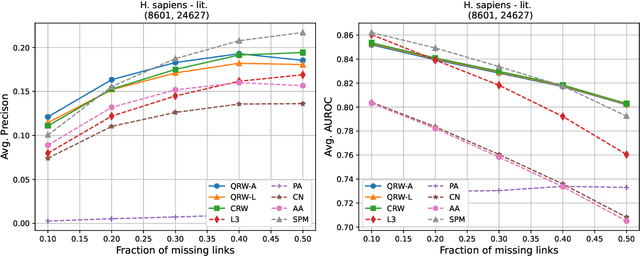
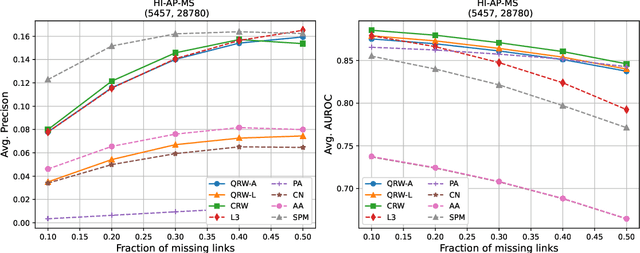
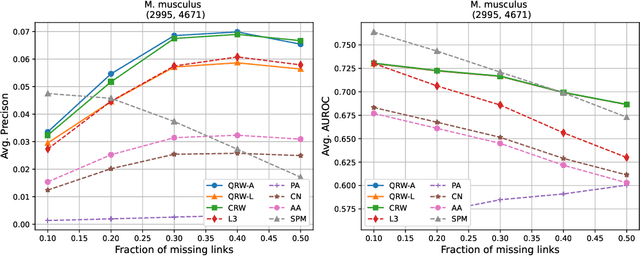
Abstract:Protein-protein interaction (PPI) networks consist of the physical and/or functional interactions between the proteins of an organism. Since the biophysical and high-throughput methods used to form PPI networks are expensive, time-consuming, and often contain inaccuracies, the resulting networks are usually incomplete. In order to infer missing interactions in these networks, we propose a novel class of link prediction methods based on continuous-time classical and quantum random walks. In the case of quantum walks, we examine the usage of both the network adjacency and Laplacian matrices for controlling the walk dynamics. We define a score function based on the corresponding transition probabilities and perform tests on four real-world PPI datasets. Our results show that continuous-time classical random walks and quantum walks using the network adjacency matrix can successfully predict missing protein-protein interactions, with performance rivalling the state of the art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge