Sabbir M. Rashid
Knowledge Graphs
Mar 28, 2020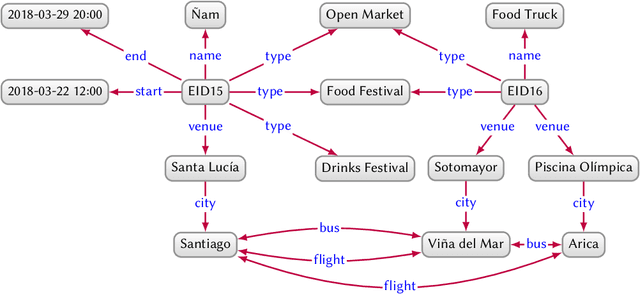
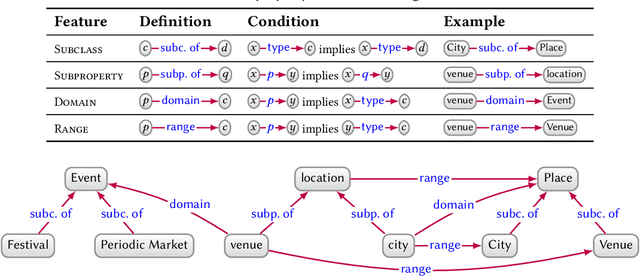
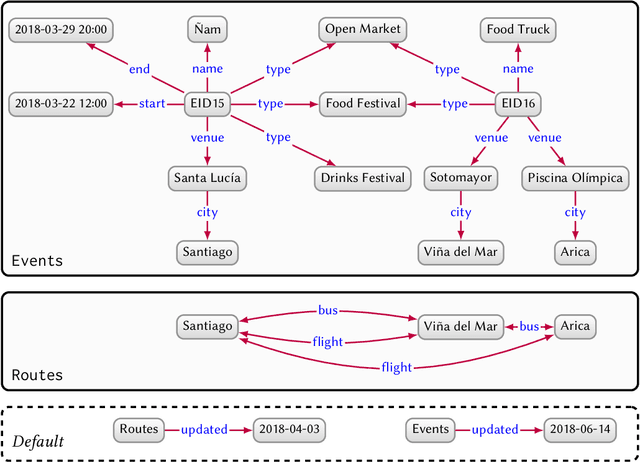
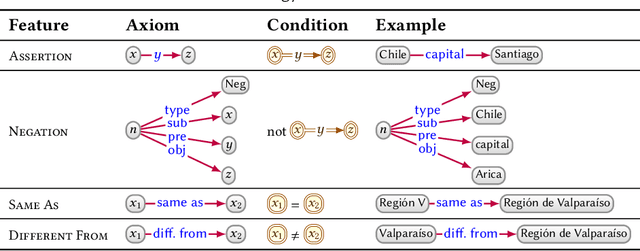
Abstract:In this paper we provide a comprehensive introduction to knowledge graphs, which have recently garnered significant attention from both industry and academia in scenarios that require exploiting diverse, dynamic, large-scale collections of data. After a general introduction, we motivate and contrast various graph-based data models and query languages that are used for knowledge graphs. We discuss the roles of schema, identity, and context in knowledge graphs. We explain how knowledge can be represented and extracted using a combination of deductive and inductive techniques. We summarise methods for the creation, enrichment, quality assessment, refinement, and publication of knowledge graphs. We provide an overview of prominent open knowledge graphs and enterprise knowledge graphs, their applications, and how they use the aforementioned techniques. We conclude with high-level future research directions for knowledge graphs.
Semantically-aware population health risk analyses
Nov 27, 2018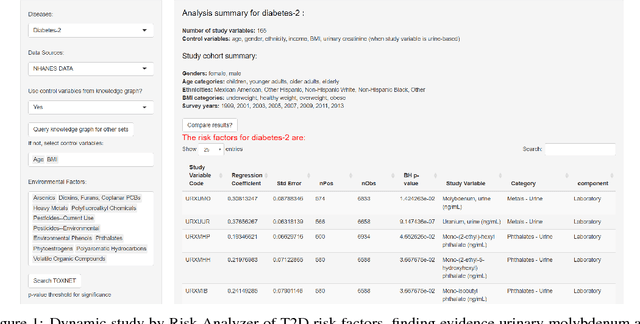

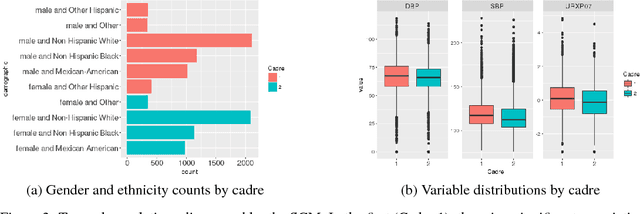
Abstract:One primary task of population health analysis is the identification of risk factors that, for some subpopulation, have a significant association with some health condition. Examples include finding lifestyle factors associated with chronic diseases and finding genetic mutations associated with diseases in precision health. We develop a combined semantic and machine learning system that uses a health risk ontology and knowledge graph (KG) to dynamically discover risk factors and their associated subpopulations. Semantics and the novel supervised cadre model make our system explainable. Future population health studies are easily performed and documented with provenance by specifying additional input and output KG cartridges.
Knowledge Integration for Disease Characterization: A Breast Cancer Example
Jul 20, 2018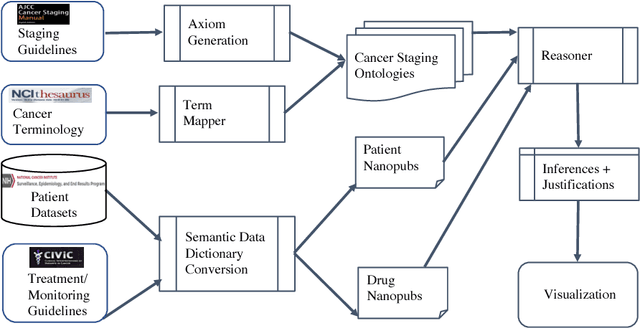
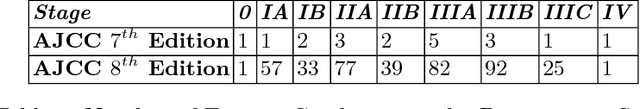
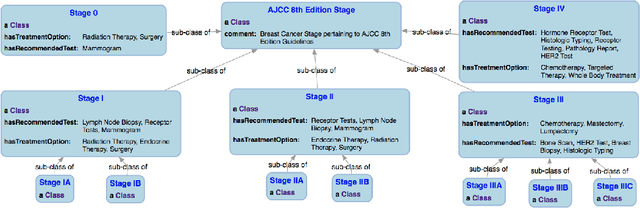
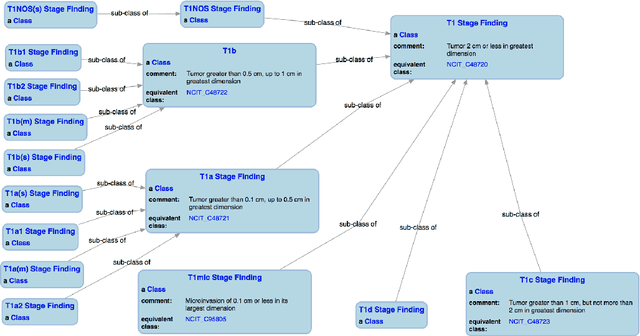
Abstract:With the rapid advancements in cancer research, the information that is useful for characterizing disease, staging tumors, and creating treatment and survivorship plans has been changing at a pace that creates challenges when physicians try to remain current. One example involves increasing usage of biomarkers when characterizing the pathologic prognostic stage of a breast tumor. We present our semantic technology approach to support cancer characterization and demonstrate it in our end-to-end prototype system that collects the newest breast cancer staging criteria from authoritative oncology manuals to construct an ontology for breast cancer. Using a tool we developed that utilizes this ontology, physician-facing applications can be used to quickly stage a new patient to support identifying risks, treatment options, and monitoring plans based on authoritative and best practice guidelines. Physicians can also re-stage existing patients or patient populations, allowing them to find patients whose stage has changed in a given patient cohort. As new guidelines emerge, using our proposed mechanism, which is grounded by semantic technologies for ingesting new data from staging manuals, we have created an enriched cancer staging ontology that integrates relevant data from several sources with very little human intervention.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge