Ryan P. Adams
Patterns of Scalable Bayesian Inference
Mar 22, 2016
Abstract:Datasets are growing not just in size but in complexity, creating a demand for rich models and quantification of uncertainty. Bayesian methods are an excellent fit for this demand, but scaling Bayesian inference is a challenge. In response to this challenge, there has been considerable recent work based on varying assumptions about model structure, underlying computational resources, and the importance of asymptotic correctness. As a result, there is a zoo of ideas with few clear overarching principles. In this paper, we seek to identify unifying principles, patterns, and intuitions for scaling Bayesian inference. We review existing work on utilizing modern computing resources with both MCMC and variational approximation techniques. From this taxonomy of ideas, we characterize the general principles that have proven successful for designing scalable inference procedures and comment on the path forward.
Predictive Entropy Search for Multi-objective Bayesian Optimization
Feb 21, 2016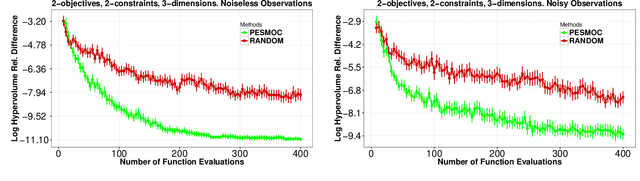
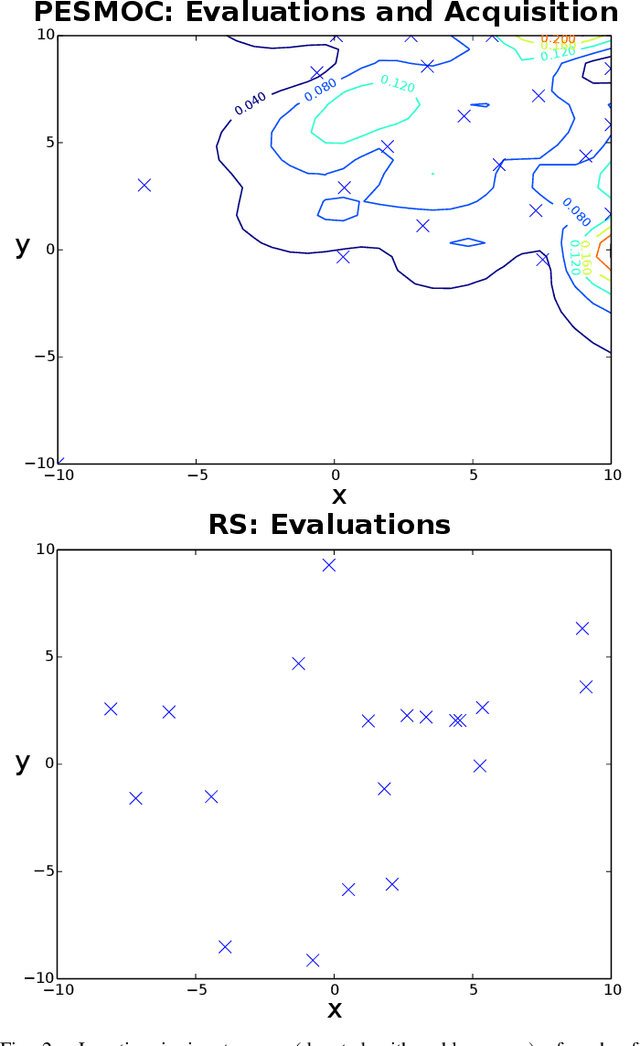
Abstract:We present PESMO, a Bayesian method for identifying the Pareto set of multi-objective optimization problems, when the functions are expensive to evaluate. The central idea of PESMO is to choose evaluation points so as to maximally reduce the entropy of the posterior distribution over the Pareto set. Critically, the PESMO multi-objective acquisition function can be decomposed as a sum of objective-specific acquisition functions, which enables the algorithm to be used in \emph{decoupled} scenarios in which the objectives can be evaluated separately and perhaps with different costs. This decoupling capability also makes it possible to identify difficult objectives that require more evaluations. PESMO also offers gains in efficiency, as its cost scales linearly with the number of objectives, in comparison to the exponential cost of other methods. We compare PESMO with other related methods for multi-objective Bayesian optimization on synthetic and real-world problems. The results show that PESMO produces better recommendations with a smaller number of evaluations of the objectives, and that a decoupled evaluation can lead to improvements in performance, particularly when the number of objectives is large.
Sandwiching the marginal likelihood using bidirectional Monte Carlo
Nov 08, 2015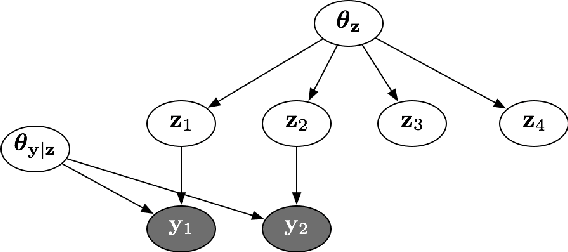

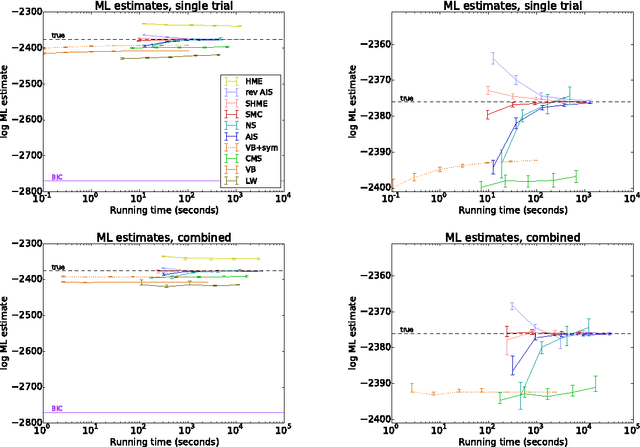
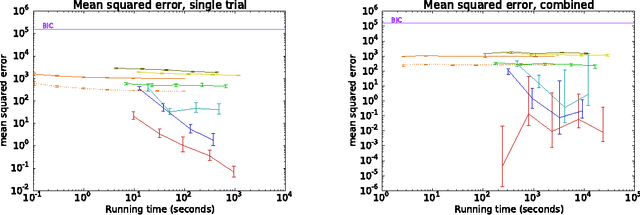
Abstract:Computing the marginal likelihood (ML) of a model requires marginalizing out all of the parameters and latent variables, a difficult high-dimensional summation or integration problem. To make matters worse, it is often hard to measure the accuracy of one's ML estimates. We present bidirectional Monte Carlo, a technique for obtaining accurate log-ML estimates on data simulated from a model. This method obtains stochastic lower bounds on the log-ML using annealed importance sampling or sequential Monte Carlo, and obtains stochastic upper bounds by running these same algorithms in reverse starting from an exact posterior sample. The true value can be sandwiched between these two stochastic bounds with high probability. Using the ground truth log-ML estimates obtained from our method, we quantitatively evaluate a wide variety of existing ML estimators on several latent variable models: clustering, a low rank approximation, and a binary attributes model. These experiments yield insights into how to accurately estimate marginal likelihoods.
Convolutional Networks on Graphs for Learning Molecular Fingerprints
Nov 03, 2015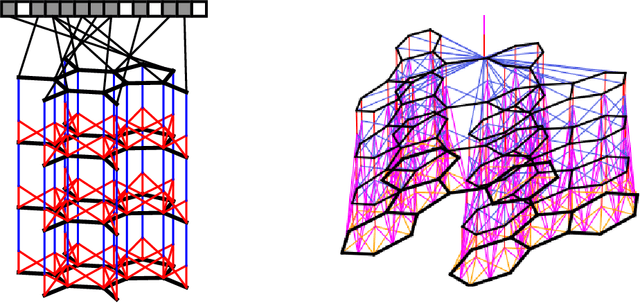
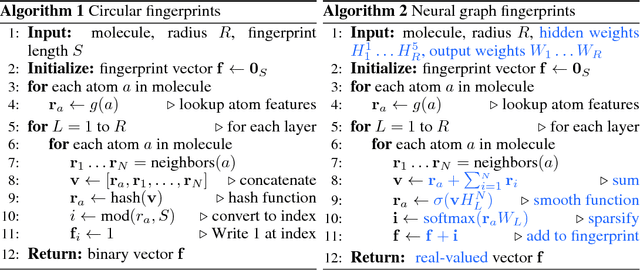
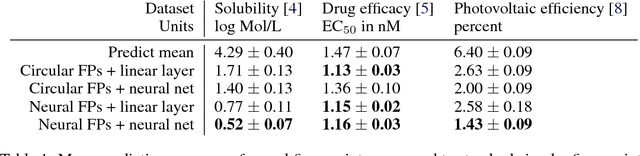

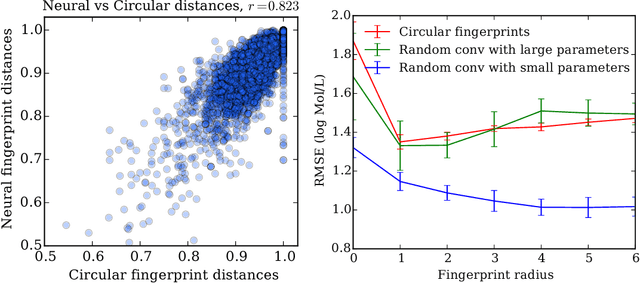
Abstract:We introduce a convolutional neural network that operates directly on graphs. These networks allow end-to-end learning of prediction pipelines whose inputs are graphs of arbitrary size and shape. The architecture we present generalizes standard molecular feature extraction methods based on circular fingerprints. We show that these data-driven features are more interpretable, and have better predictive performance on a variety of tasks.
Predictive Entropy Search for Bayesian Optimization with Unknown Constraints
Jul 15, 2015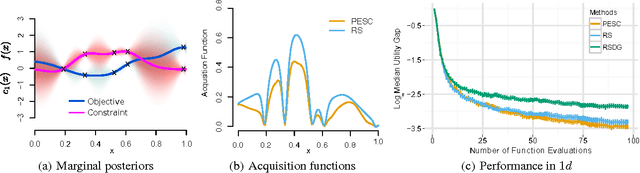
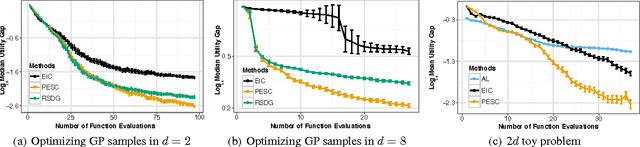
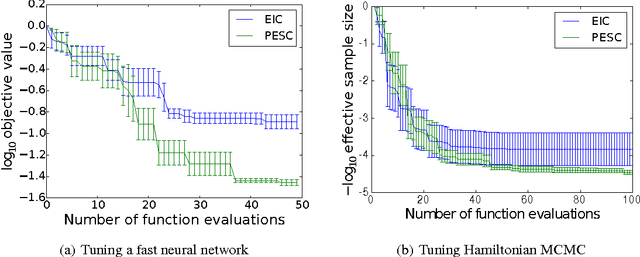
Abstract:Unknown constraints arise in many types of expensive black-box optimization problems. Several methods have been proposed recently for performing Bayesian optimization with constraints, based on the expected improvement (EI) heuristic. However, EI can lead to pathologies when used with constraints. For example, in the case of decoupled constraints---i.e., when one can independently evaluate the objective or the constraints---EI can encounter a pathology that prevents exploration. Additionally, computing EI requires a current best solution, which may not exist if none of the data collected so far satisfy the constraints. By contrast, information-based approaches do not suffer from these failure modes. In this paper, we present a new information-based method called Predictive Entropy Search with Constraints (PESC). We analyze the performance of PESC and show that it compares favorably to EI-based approaches on synthetic and benchmark problems, as well as several real-world examples. We demonstrate that PESC is an effective algorithm that provides a promising direction towards a unified solution for constrained Bayesian optimization.
Probabilistic Backpropagation for Scalable Learning of Bayesian Neural Networks
Jul 15, 2015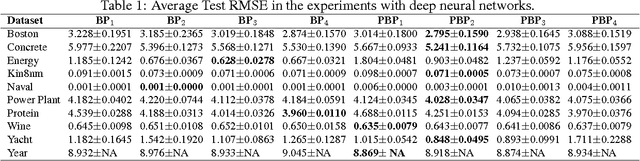
Abstract:Large multilayer neural networks trained with backpropagation have recently achieved state-of-the-art results in a wide range of problems. However, using backprop for neural net learning still has some disadvantages, e.g., having to tune a large number of hyperparameters to the data, lack of calibrated probabilistic predictions, and a tendency to overfit the training data. In principle, the Bayesian approach to learning neural networks does not have these problems. However, existing Bayesian techniques lack scalability to large dataset and network sizes. In this work we present a novel scalable method for learning Bayesian neural networks, called probabilistic backpropagation (PBP). Similar to classical backpropagation, PBP works by computing a forward propagation of probabilities through the network and then doing a backward computation of gradients. A series of experiments on ten real-world datasets show that PBP is significantly faster than other techniques, while offering competitive predictive abilities. Our experiments also show that PBP provides accurate estimates of the posterior variance on the network weights.
Scalable Bayesian Optimization Using Deep Neural Networks
Jul 13, 2015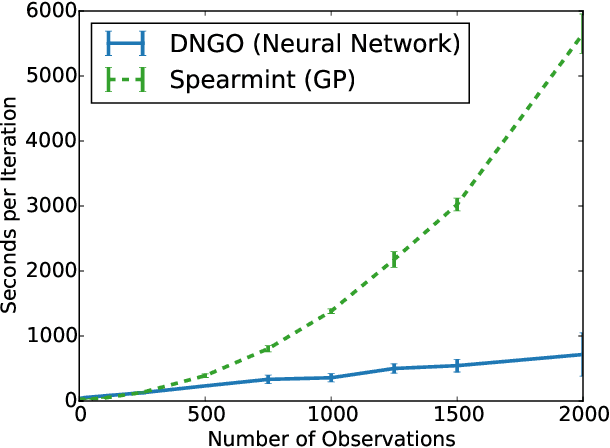
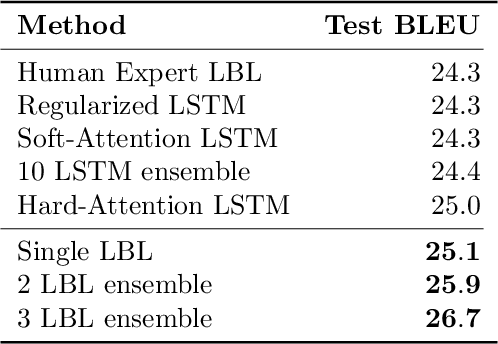
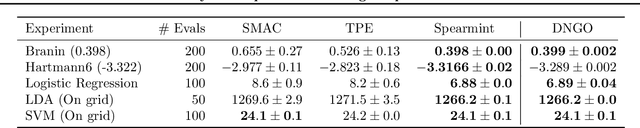
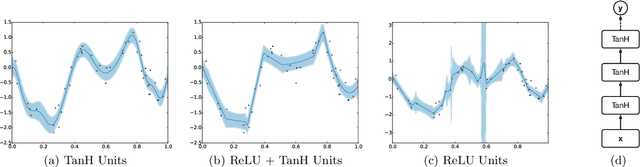

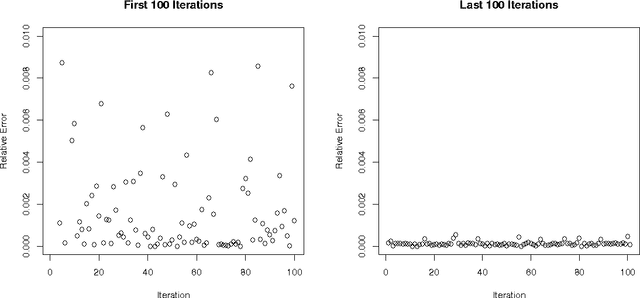
Abstract:Bayesian optimization is an effective methodology for the global optimization of functions with expensive evaluations. It relies on querying a distribution over functions defined by a relatively cheap surrogate model. An accurate model for this distribution over functions is critical to the effectiveness of the approach, and is typically fit using Gaussian processes (GPs). However, since GPs scale cubically with the number of observations, it has been challenging to handle objectives whose optimization requires many evaluations, and as such, massively parallelizing the optimization. In this work, we explore the use of neural networks as an alternative to GPs to model distributions over functions. We show that performing adaptive basis function regression with a neural network as the parametric form performs competitively with state-of-the-art GP-based approaches, but scales linearly with the number of data rather than cubically. This allows us to achieve a previously intractable degree of parallelism, which we apply to large scale hyperparameter optimization, rapidly finding competitive models on benchmark object recognition tasks using convolutional networks, and image caption generation using neural language models.
Scalable Bayesian Inference for Excitatory Point Process Networks
Jul 12, 2015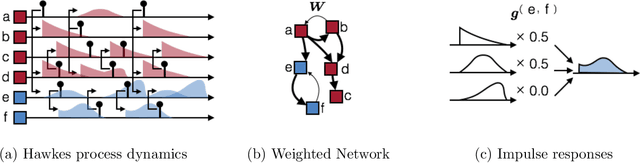
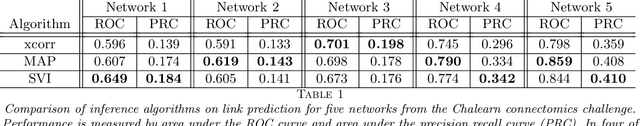
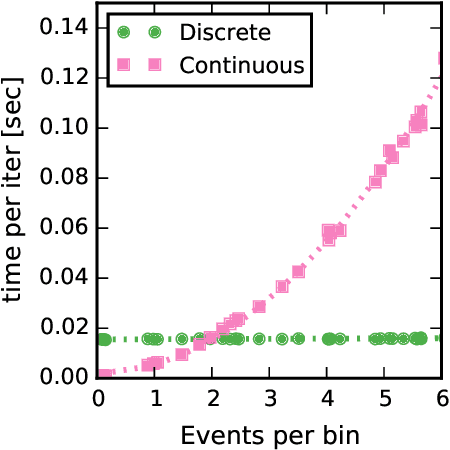
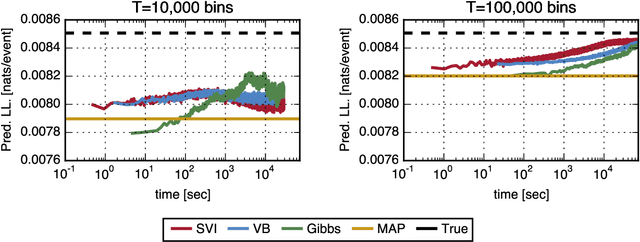
Abstract:Networks capture our intuition about relationships in the world. They describe the friendships between Facebook users, interactions in financial markets, and synapses connecting neurons in the brain. These networks are richly structured with cliques of friends, sectors of stocks, and a smorgasbord of cell types that govern how neurons connect. Some networks, like social network friendships, can be directly observed, but in many cases we only have an indirect view of the network through the actions of its constituents and an understanding of how the network mediates that activity. In this work, we focus on the problem of latent network discovery in the case where the observable activity takes the form of a mutually-excitatory point process known as a Hawkes process. We build on previous work that has taken a Bayesian approach to this problem, specifying prior distributions over the latent network structure and a likelihood of observed activity given this network. We extend this work by proposing a discrete-time formulation and developing a computationally efficient stochastic variational inference (SVI) algorithm that allows us to scale the approach to long sequences of observations. We demonstrate our algorithm on the calcium imaging data used in the Chalearn neural connectomics challenge.
Dependent Multinomial Models Made Easy: Stick Breaking with the Pólya-Gamma Augmentation
Jun 18, 2015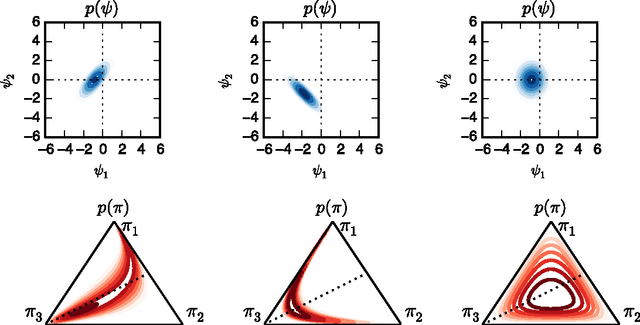
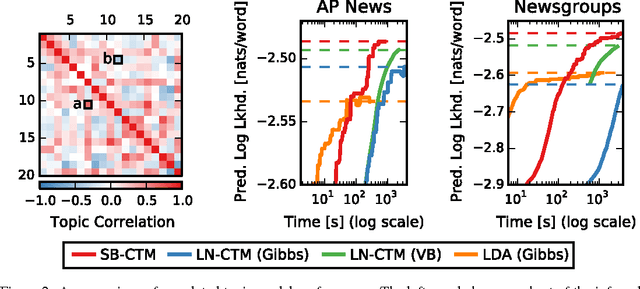
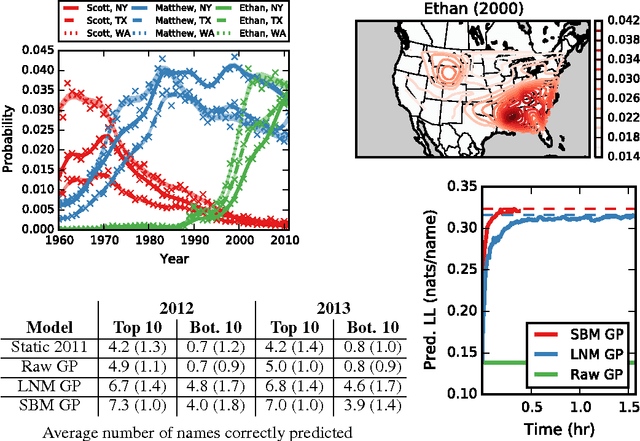
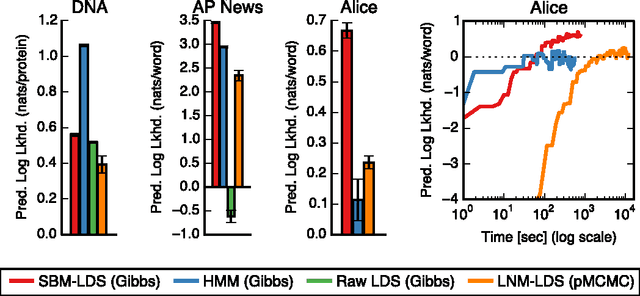
Abstract:Many practical modeling problems involve discrete data that are best represented as draws from multinomial or categorical distributions. For example, nucleotides in a DNA sequence, children's names in a given state and year, and text documents are all commonly modeled with multinomial distributions. In all of these cases, we expect some form of dependency between the draws: the nucleotide at one position in the DNA strand may depend on the preceding nucleotides, children's names are highly correlated from year to year, and topics in text may be correlated and dynamic. These dependencies are not naturally captured by the typical Dirichlet-multinomial formulation. Here, we leverage a logistic stick-breaking representation and recent innovations in P\'olya-gamma augmentation to reformulate the multinomial distribution in terms of latent variables with jointly Gaussian likelihoods, enabling us to take advantage of a host of Bayesian inference techniques for Gaussian models with minimal overhead.
Spectral Representations for Convolutional Neural Networks
Jun 11, 2015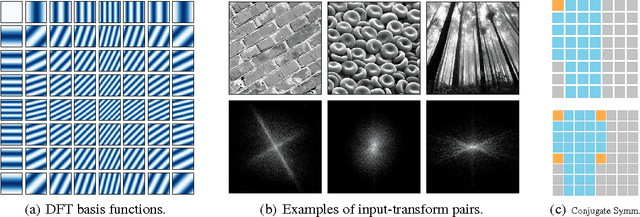
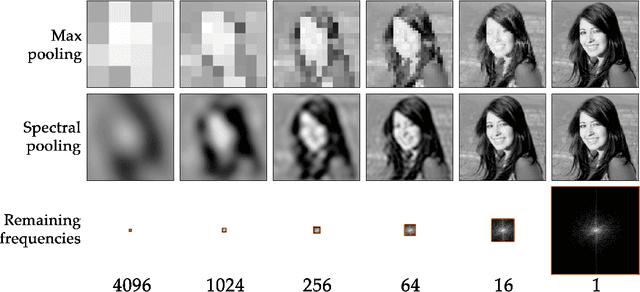

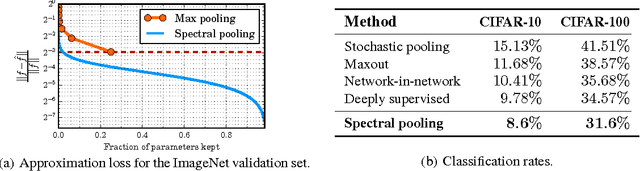
Abstract:Discrete Fourier transforms provide a significant speedup in the computation of convolutions in deep learning. In this work, we demonstrate that, beyond its advantages for efficient computation, the spectral domain also provides a powerful representation in which to model and train convolutional neural networks (CNNs). We employ spectral representations to introduce a number of innovations to CNN design. First, we propose spectral pooling, which performs dimensionality reduction by truncating the representation in the frequency domain. This approach preserves considerably more information per parameter than other pooling strategies and enables flexibility in the choice of pooling output dimensionality. This representation also enables a new form of stochastic regularization by randomized modification of resolution. We show that these methods achieve competitive results on classification and approximation tasks, without using any dropout or max-pooling. Finally, we demonstrate the effectiveness of complex-coefficient spectral parameterization of convolutional filters. While this leaves the underlying model unchanged, it results in a representation that greatly facilitates optimization. We observe on a variety of popular CNN configurations that this leads to significantly faster convergence during training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge