Rajendra Singh
PathoScribe: Transforming Pathology Data into a Living Library with a Unified LLM-Driven Framework for Semantic Retrieval and Clinical Integration
Mar 11, 2026Abstract:Pathology underpins modern diagnosis and cancer care, yet its most valuable asset, the accumulated experience encoded in millions of narrative reports, remains largely inaccessible. Although institutions are rapidly digitizing pathology workflows, storing data without effective mechanisms for retrieval and reasoning risks transforming archives into a passive data repository, where institutional knowledge exists but cannot meaningfully inform patient care. True progress requires not only digitization, but the ability for pathologists to interrogate prior similar cases in real time while evaluating a new diagnostic dilemma. We present PathoScribe, a unified retrieval-augmented large language model (LLM) framework designed to transform static pathology archives into a searchable, reasoning-enabled living library. PathoScribe enables natural language case exploration, automated cohort construction, clinical question answering, immunohistochemistry (IHC) panel recommendation, and prompt-controlled report transformation within a single architecture. Evaluated on 70,000 multi-institutional surgical pathology reports, PathoScribe achieved perfect Recall@10 for natural language case retrieval and demonstrated high-quality retrieval-grounded reasoning (mean reviewer score 4.56/5). Critically, the system operationalized automated cohort construction from free-text eligibility criteria, assembling research-ready cohorts in minutes (mean 9.2 minutes) with 91.3% agreement to human reviewers and no eligible cases incorrectly excluded, representing orders-of-magnitude reductions in time and cost compared to traditional manual chart review. This work establishes a scalable foundation for converting digital pathology archives from passive storage systems into active clinical intelligence platforms.
Federated Learning for Computational Pathology on Gigapixel Whole Slide Images
Sep 23, 2020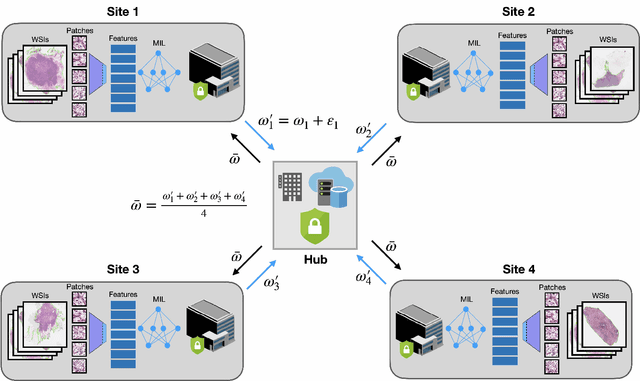
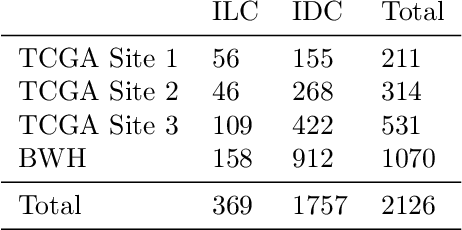
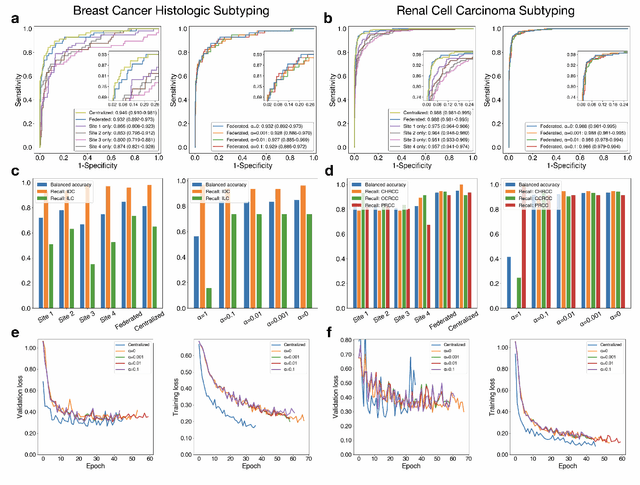
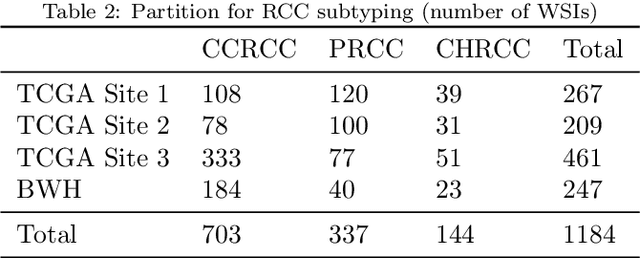
Abstract:Deep Learning-based computational pathology algorithms have demonstrated profound ability to excel in a wide array of tasks that range from characterization of well known morphological phenotypes to predicting non-human-identifiable features from histology such as molecular alterations. However, the development of robust, adaptable, and accurate deep learning-based models often rely on the collection and time-costly curation large high-quality annotated training data that should ideally come from diverse sources and patient populations to cater for the heterogeneity that exists in such datasets. Multi-centric and collaborative integration of medical data across multiple institutions can naturally help overcome this challenge and boost the model performance but is limited by privacy concerns amongst other difficulties that may arise in the complex data sharing process as models scale towards using hundreds of thousands of gigapixel whole slide images. In this paper, we introduce privacy-preserving federated learning for gigapixel whole slide images in computational pathology using weakly-supervised attention multiple instance learning and differential privacy. We evaluated our approach on two different diagnostic problems using thousands of histology whole slide images with only slide-level labels. Additionally, we present a weakly-supervised learning framework for survival prediction and patient stratification from whole slide images and demonstrate its effectiveness in a federated setting. Our results show that using federated learning, we can effectively develop accurate weakly supervised deep learning models from distributed data silos without direct data sharing and its associated complexities, while also preserving differential privacy using randomized noise generation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge