Raizy Kellerman
JEPA-DNA: Grounding Genomic Foundation Models through Joint-Embedding Predictive Architectures
Feb 19, 2026Abstract:Genomic Foundation Models (GFMs) have largely relied on Masked Language Modeling (MLM) or Next Token Prediction (NTP) to learn the language of life. While these paradigms excel at capturing local genomic syntax and fine-grained motif patterns, they often fail to capture the broader functional context, resulting in representations that lack a global biological perspective. We introduce JEPA-DNA, a novel pre-training framework that integrates the Joint-Embedding Predictive Architecture (JEPA) with traditional generative objectives. JEPA-DNA introduces latent grounding by coupling token-level recovery with a predictive objective in the latent space by supervising a CLS token. This forces the model to predict the high-level functional embeddings of masked genomic segments rather than focusing solely on individual nucleotides. JEPA-DNA extends both NTP and MLM paradigms and can be deployed either as a standalone from-scratch objective or as a continual pre-training enhancement for existing GFMs. Our evaluations across a diverse suite of genomic benchmarks demonstrate that JEPA-DNA consistently yields superior performance in supervised and zero-shot tasks compared to generative-only baselines. By providing a more robust and biologically grounded representation, JEPA-DNA offers a scalable path toward foundation models that understand not only the genomic alphabet, but also the underlying functional logic of the sequence.
Multi-Loss Sub-Ensembles for Accurate Classification with Uncertainty Estimation
Oct 05, 2020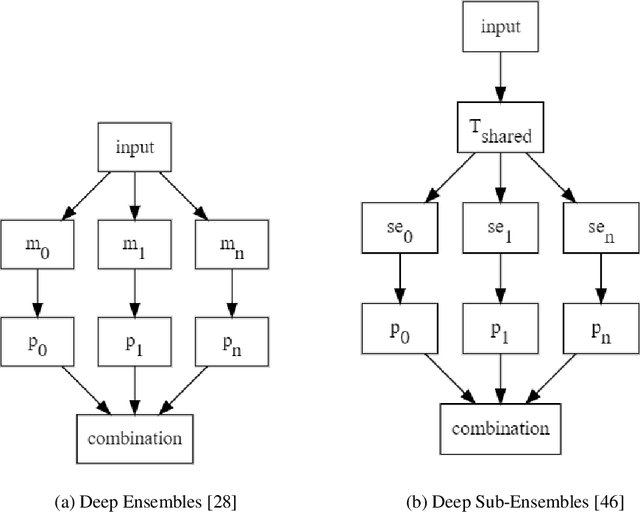
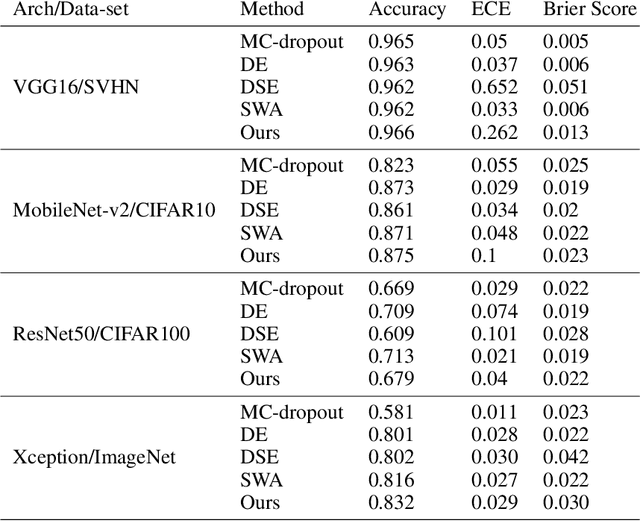
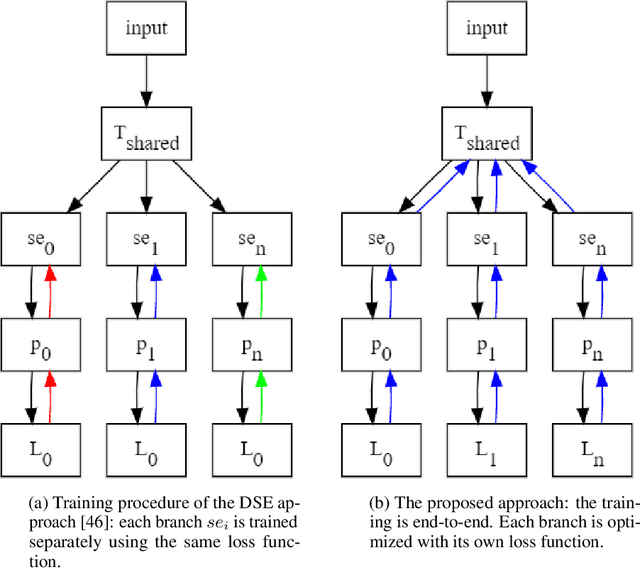
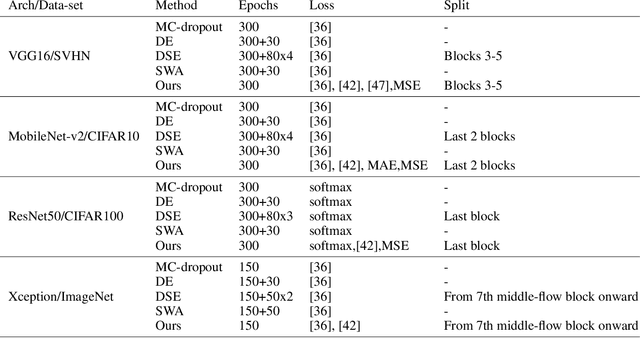
Abstract:Deep neural networks (DNNs) have made a revolution in numerous fields during the last decade. However, in tasks with high safety requirements, such as medical or autonomous driving applications, providing an assessment of the models reliability can be vital. Uncertainty estimation for DNNs has been addressed using Bayesian methods, providing mathematically founded models for reliability assessment. These model are computationally expensive and generally impractical for many real-time use cases. Recently, non-Bayesian methods were proposed to tackle uncertainty estimation more efficiently. We propose an efficient method for uncertainty estimation in DNNs achieving high accuracy. We simulate the notion of multi-task learning on single-task problems by producing parallel predictions from similar models differing by their loss. This multi-loss approach allows one-phase training for single-task learning with uncertainty estimation. We keep our inference time relatively low by leveraging the advantage proposed by the Deep-Sub-Ensembles method. The novelty of this work resides in the proposed accurate variational inference with a simple and convenient training procedure, while remaining competitive in terms of computational time. We conduct experiments on SVHN, CIFAR10, CIFAR100 as well as Image-Net using different architectures. Our results show improved accuracy on the classification task and competitive results on several uncertainty measures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge