Quande Liu
ConceptMaster: Multi-Concept Video Customization on Diffusion Transformer Models Without Test-Time Tuning
Jan 08, 2025

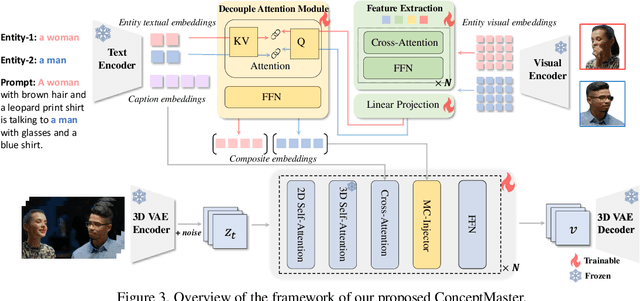
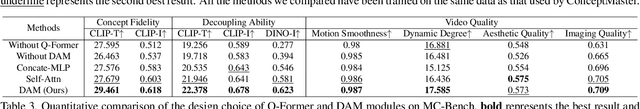
Abstract:Text-to-video generation has made remarkable advancements through diffusion models. However, Multi-Concept Video Customization (MCVC) remains a significant challenge. We identify two key challenges in this task: 1) the identity decoupling problem, where directly adopting existing customization methods inevitably mix attributes when handling multiple concepts simultaneously, and 2) the scarcity of high-quality video-entity pairs, which is crucial for training such a model that represents and decouples various concepts well. To address these challenges, we introduce ConceptMaster, an innovative framework that effectively tackles the critical issues of identity decoupling while maintaining concept fidelity in customized videos. Specifically, we introduce a novel strategy of learning decoupled multi-concept embeddings that are injected into the diffusion models in a standalone manner, which effectively guarantees the quality of customized videos with multiple identities, even for highly similar visual concepts. To further overcome the scarcity of high-quality MCVC data, we carefully establish a data construction pipeline, which enables systematic collection of precise multi-concept video-entity data across diverse concepts. A comprehensive benchmark is designed to validate the effectiveness of our model from three critical dimensions: concept fidelity, identity decoupling ability, and video generation quality across six different concept composition scenarios. Extensive experiments demonstrate that our ConceptMaster significantly outperforms previous approaches for this task, paving the way for generating personalized and semantically accurate videos across multiple concepts.
GameGen-X: Interactive Open-world Game Video Generation
Nov 01, 2024



Abstract:We introduce GameGen-X, the first diffusion transformer model specifically designed for both generating and interactively controlling open-world game videos. This model facilitates high-quality, open-domain generation by simulating an extensive array of game engine features, such as innovative characters, dynamic environments, complex actions, and diverse events. Additionally, it provides interactive controllability, predicting and altering future content based on the current clip, thus allowing for gameplay simulation. To realize this vision, we first collected and built an Open-World Video Game Dataset from scratch. It is the first and largest dataset for open-world game video generation and control, which comprises over a million diverse gameplay video clips sampling from over 150 games with informative captions from GPT-4o. GameGen-X undergoes a two-stage training process, consisting of foundation model pre-training and instruction tuning. Firstly, the model was pre-trained via text-to-video generation and video continuation, endowing it with the capability for long-sequence, high-quality open-domain game video generation. Further, to achieve interactive controllability, we designed InstructNet to incorporate game-related multi-modal control signal experts. This allows the model to adjust latent representations based on user inputs, unifying character interaction and scene content control for the first time in video generation. During instruction tuning, only the InstructNet is updated while the pre-trained foundation model is frozen, enabling the integration of interactive controllability without loss of diversity and quality of generated video content.
Text-Animator: Controllable Visual Text Video Generation
Jun 25, 2024Abstract:Video generation is a challenging yet pivotal task in various industries, such as gaming, e-commerce, and advertising. One significant unresolved aspect within T2V is the effective visualization of text within generated videos. Despite the progress achieved in Text-to-Video~(T2V) generation, current methods still cannot effectively visualize texts in videos directly, as they mainly focus on summarizing semantic scene information, understanding, and depicting actions. While recent advances in image-level visual text generation show promise, transitioning these techniques into the video domain faces problems, notably in preserving textual fidelity and motion coherence. In this paper, we propose an innovative approach termed Text-Animator for visual text video generation. Text-Animator contains a text embedding injection module to precisely depict the structures of visual text in generated videos. Besides, we develop a camera control module and a text refinement module to improve the stability of generated visual text by controlling the camera movement as well as the motion of visualized text. Quantitative and qualitative experimental results demonstrate the superiority of our approach to the accuracy of generated visual text over state-of-the-art video generation methods. The project page can be found at https://laulampaul.github.io/text-animator.html.
ID-Animator: Zero-Shot Identity-Preserving Human Video Generation
Apr 23, 2024
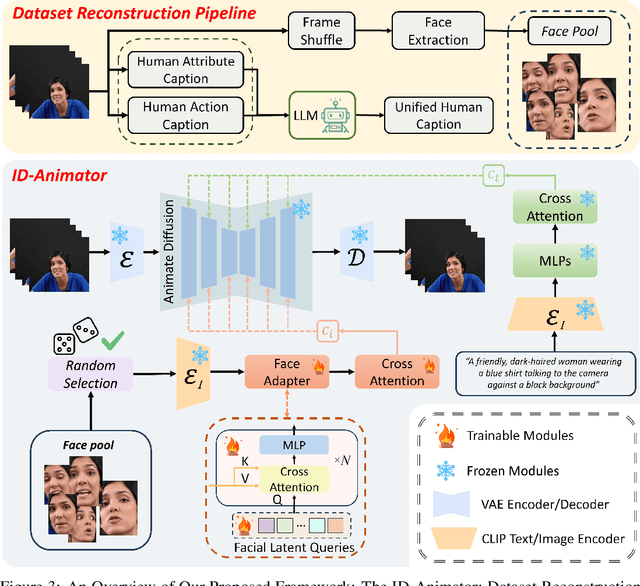


Abstract:Generating high fidelity human video with specified identities has attracted significant attention in the content generation community. However, existing techniques struggle to strike a balance between training efficiency and identity preservation, either requiring tedious case-by-case finetuning or usually missing the identity details in video generation process. In this study, we present ID-Animator, a zero-shot human-video generation approach that can perform personalized video generation given single reference facial image without further training. ID-Animator inherits existing diffusion-based video generation backbones with a face adapter to encode the ID-relevant embeddings from learnable facial latent queries. To facilitate the extraction of identity information in video generation, we introduce an ID-oriented dataset construction pipeline, which incorporates decoupled human attribute and action captioning technique from a constructed facial image pool. Based on this pipeline, a random face reference training method is further devised to precisely capture the ID-relevant embeddings from reference images, thus improving the fidelity and generalization capacity of our model for ID-specific video generation. Extensive experiments demonstrate the superiority of ID-Animator to generate personalized human videos over previous models. Moreover, our method is highly compatible with popular pre-trained T2V models like animatediff and various community backbone models, showing high extendability in real-world applications for video generation where identity preservation is highly desired. Our codes and checkpoints will be released at https://github.com/ID-Animator/ID-Animator.
Open-world Semantic Segmentation via Contrasting and Clustering Vision-Language Embedding
Jul 19, 2022
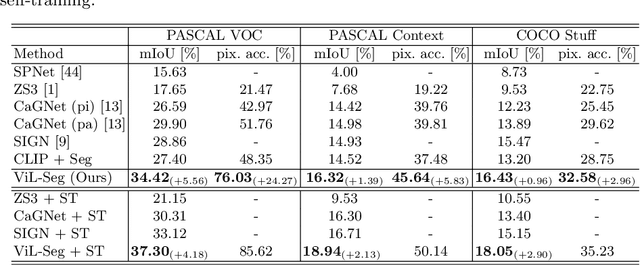
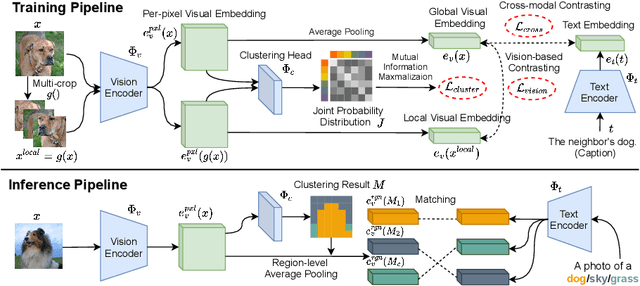
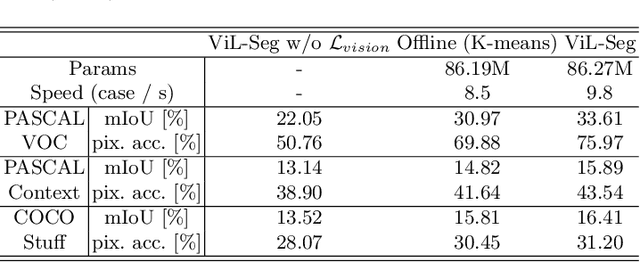
Abstract:To bridge the gap between supervised semantic segmentation and real-world applications that acquires one model to recognize arbitrary new concepts, recent zero-shot segmentation attracts a lot of attention by exploring the relationships between unseen and seen object categories, yet requiring large amounts of densely-annotated data with diverse base classes. In this paper, we propose a new open-world semantic segmentation pipeline that makes the first attempt to learn to segment semantic objects of various open-world categories without any efforts on dense annotations, by purely exploiting the image-caption data that naturally exist on the Internet. Our method, Vision-language-driven Semantic Segmentation (ViL-Seg), employs an image and a text encoder to generate visual and text embeddings for the image-caption data, with two core components that endow its segmentation ability: First, the image encoder is jointly trained with a vision-based contrasting and a cross-modal contrasting, which encourage the visual embeddings to preserve both fine-grained semantics and high-level category information that are crucial for the segmentation task. Furthermore, an online clustering head is devised over the image encoder, which allows to dynamically segment the visual embeddings into distinct semantic groups such that they can be classified by comparing with various text embeddings to complete our segmentation pipeline. Experiments show that without using any data with dense annotations, our method can directly segment objects of arbitrary categories, outperforming zero-shot segmentation methods that require data labeling on three benchmark datasets.
Single-domain Generalization in Medical Image Segmentation via Test-time Adaptation from Shape Dictionary
Jun 29, 2022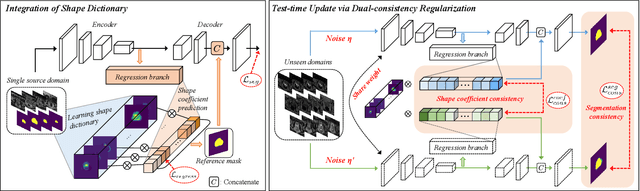
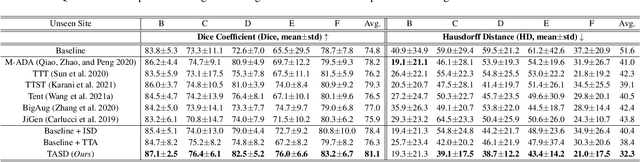

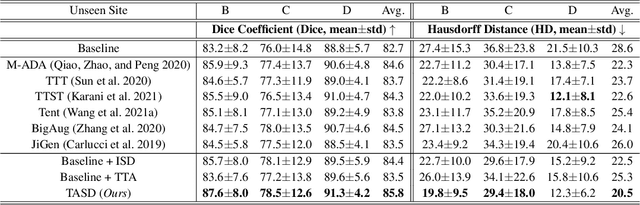
Abstract:Domain generalization typically requires data from multiple source domains for model learning. However, such strong assumption may not always hold in practice, especially in medical field where the data sharing is highly concerned and sometimes prohibitive due to privacy issue. This paper studies the important yet challenging single domain generalization problem, in which a model is learned under the worst-case scenario with only one source domain to directly generalize to different unseen target domains. We present a novel approach to address this problem in medical image segmentation, which extracts and integrates the semantic shape prior information of segmentation that are invariant across domains and can be well-captured even from single domain data to facilitate segmentation under distribution shifts. Besides, a test-time adaptation strategy with dual-consistency regularization is further devised to promote dynamic incorporation of these shape priors under each unseen domain to improve model generalizability. Extensive experiments on two medical image segmentation tasks demonstrate the consistent improvements of our method across various unseen domains, as well as its superiority over state-of-the-art approaches in addressing domain generalization under the worst-case scenario.
Dynamic Bank Learning for Semi-supervised Federated Image Diagnosis with Class Imbalance
Jun 27, 2022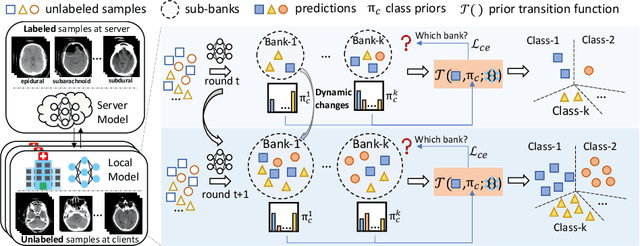
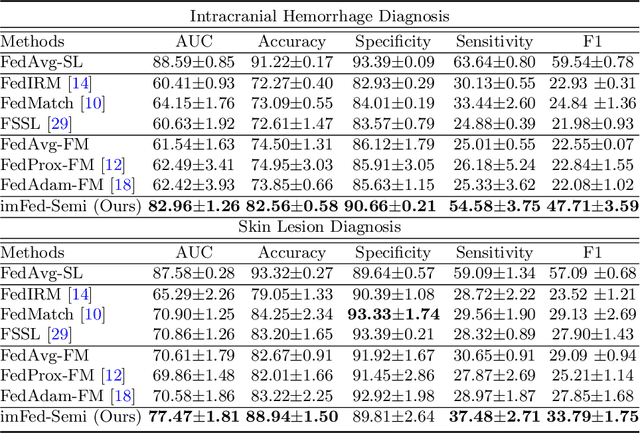
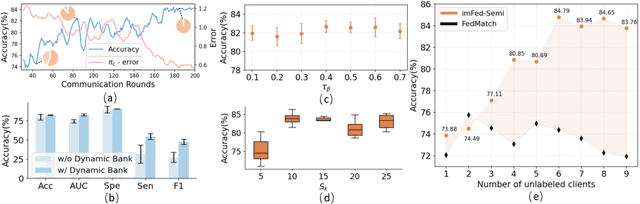
Abstract:Despite recent progress on semi-supervised federated learning (FL) for medical image diagnosis, the problem of imbalanced class distributions among unlabeled clients is still unsolved for real-world use. In this paper, we study a practical yet challenging problem of class imbalanced semi-supervised FL (imFed-Semi), which allows all clients to have only unlabeled data while the server just has a small amount of labeled data. This imFed-Semi problem is addressed by a novel dynamic bank learning scheme, which improves client training by exploiting class proportion information. This scheme consists of two parts, i.e., the dynamic bank construction to distill various class proportions for each local client, and the sub-bank classification to impose the local model to learn different class proportions. We evaluate our approach on two public real-world medical datasets, including the intracranial hemorrhage diagnosis with 25,000 CT slices and skin lesion diagnosis with 10,015 dermoscopy images. The effectiveness of our method has been validated with significant performance improvements (7.61% and 4.69%) compared with the second-best on the accuracy, as well as comprehensive analytical studies. Code is available at https://github.com/med-air/imFedSemi.
DLTTA: Dynamic Learning Rate for Test-time Adaptation on Cross-domain Medical Images
May 27, 2022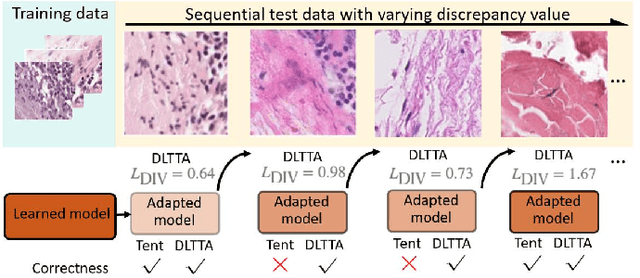
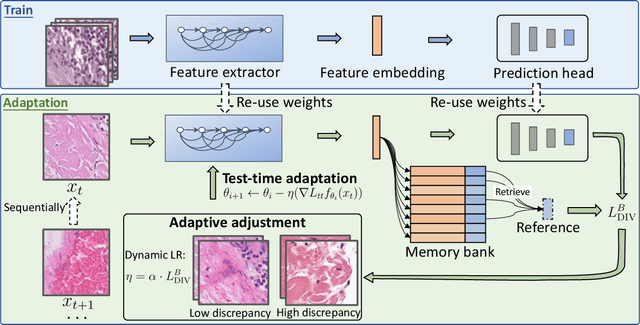
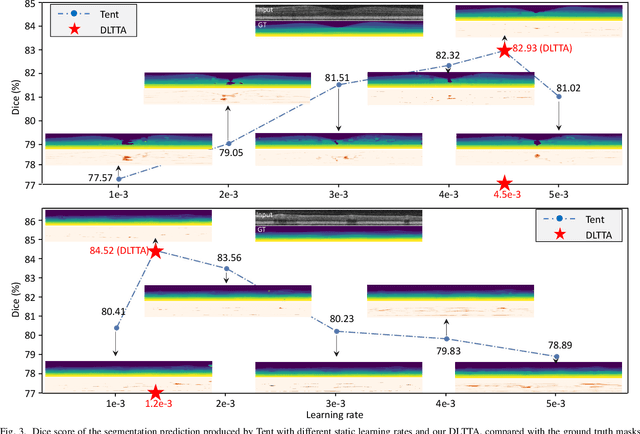
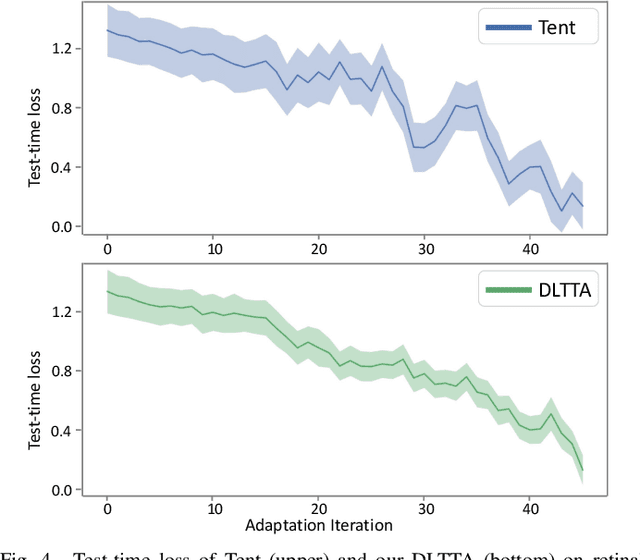
Abstract:Test-time adaptation (TTA) has increasingly been an important topic to efficiently tackle the cross-domain distribution shift at test time for medical images from different institutions. Previous TTA methods have a common limitation of using a fixed learning rate for all the test samples. Such a practice would be sub-optimal for TTA, because test data may arrive sequentially therefore the scale of distribution shift would change frequently. To address this problem, we propose a novel dynamic learning rate adjustment method for test-time adaptation, called DLTTA, which dynamically modulates the amount of weights update for each test image to account for the differences in their distribution shift. Specifically, our DLTTA is equipped with a memory bank based estimation scheme to effectively measure the discrepancy of a given test sample. Based on this estimated discrepancy, a dynamic learning rate adjustment strategy is then developed to achieve a suitable degree of adaptation for each test sample. The effectiveness and general applicability of our DLTTA is extensively demonstrated on three tasks including retinal optical coherence tomography (OCT) segmentation, histopathological image classification, and prostate 3D MRI segmentation. Our method achieves effective and fast test-time adaptation with consistent performance improvement over current state-of-the-art test-time adaptation methods. Code is available at: https://github.com/med-air/DLTTA.
Source-Free Domain Adaptive Fundus Image Segmentation with Denoised Pseudo-Labeling
Sep 19, 2021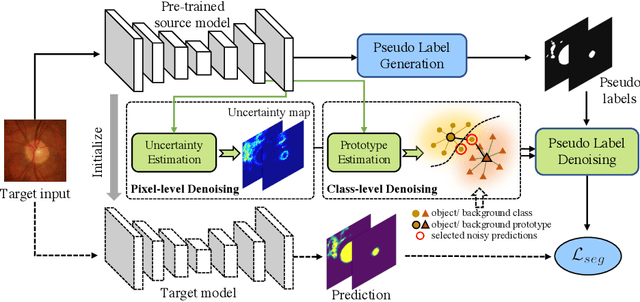
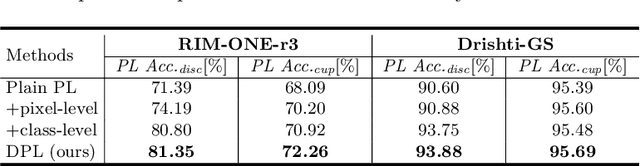
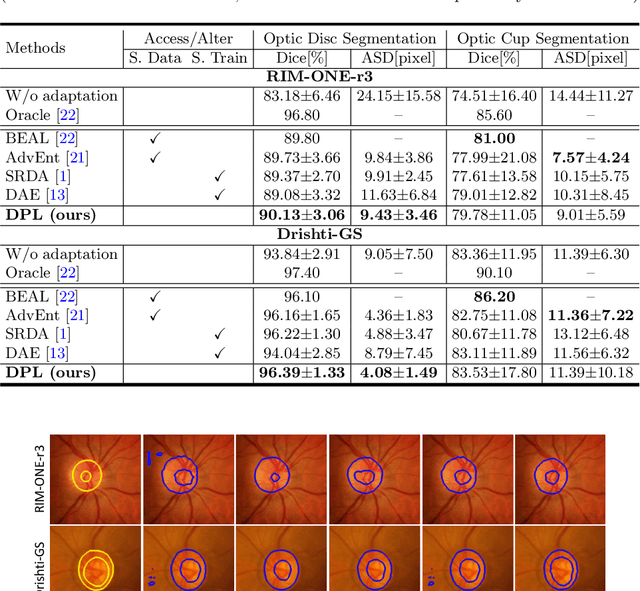

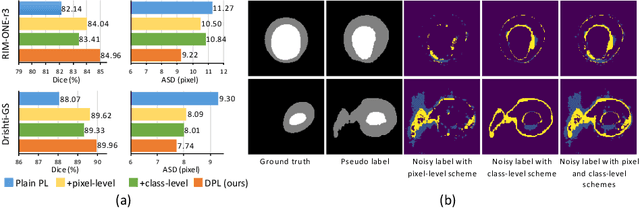
Abstract:Domain adaptation typically requires to access source domain data to utilize their distribution information for domain alignment with the target data. However, in many real-world scenarios, the source data may not be accessible during the model adaptation in the target domain due to privacy issue. This paper studies the practical yet challenging source-free unsupervised domain adaptation problem, in which only an existing source model and the unlabeled target data are available for model adaptation. We present a novel denoised pseudo-labeling method for this problem, which effectively makes use of the source model and unlabeled target data to promote model self-adaptation from pseudo labels. Importantly, considering that the pseudo labels generated from source model are inevitably noisy due to domain shift, we further introduce two complementary pixel-level and class-level denoising schemes with uncertainty estimation and prototype estimation to reduce noisy pseudo labels and select reliable ones to enhance the pseudo-labeling efficacy. Experimental results on cross-domain fundus image segmentation show that without using any source images or altering source training, our approach achieves comparable or even higher performance than state-of-the-art source-dependent unsupervised domain adaptation methods.
Federated Semi-supervised Medical Image Classification via Inter-client Relation Matching
Jun 16, 2021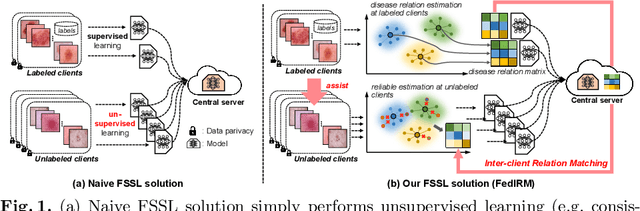
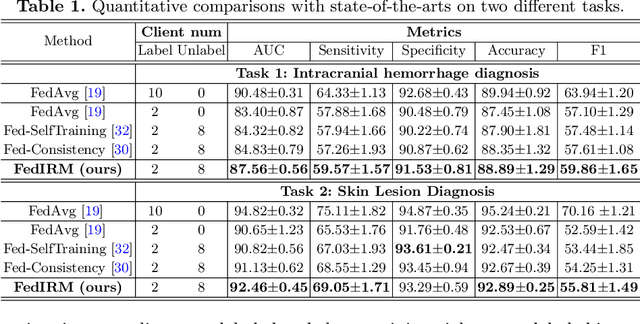
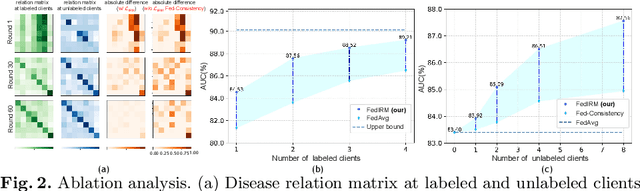
Abstract:Federated learning (FL) has emerged with increasing popularity to collaborate distributed medical institutions for training deep networks. However, despite existing FL algorithms only allow the supervised training setting, most hospitals in realistic usually cannot afford the intricate data labeling due to absence of budget or expertise. This paper studies a practical yet challenging FL problem, named \textit{Federated Semi-supervised Learning} (FSSL), which aims to learn a federated model by jointly utilizing the data from both labeled and unlabeled clients (i.e., hospitals). We present a novel approach for this problem, which improves over traditional consistency regularization mechanism with a new inter-client relation matching scheme. The proposed learning scheme explicitly connects the learning across labeled and unlabeled clients by aligning their extracted disease relationships, thereby mitigating the deficiency of task knowledge at unlabeled clients and promoting discriminative information from unlabeled samples. We validate our method on two large-scale medical image classification datasets. The effectiveness of our method has been demonstrated with the clear improvements over state-of-the-arts as well as the thorough ablation analysis on both tasks\footnote{Code will be made available at \url{https://github.com/liuquande/FedIRM}}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge