Qingyu Zhao
Dept. of Psychiatry and Behavioral Sciences, Stanford University, Stanford, CA, USA
Going Beyond Saliency Maps: Training Deep Models to Interpret Deep Models
Feb 16, 2021
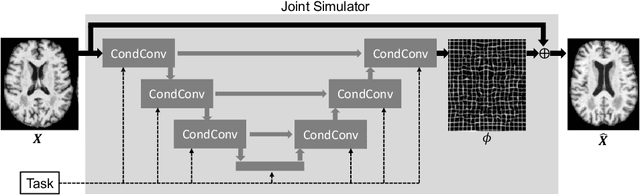
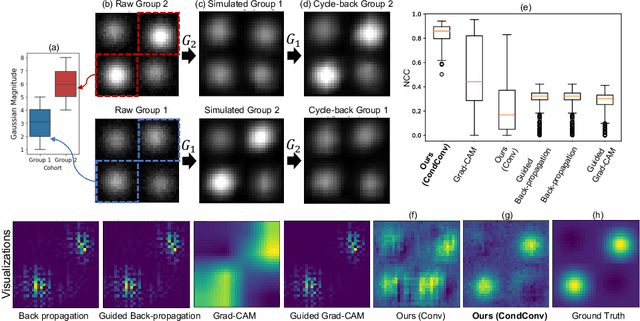
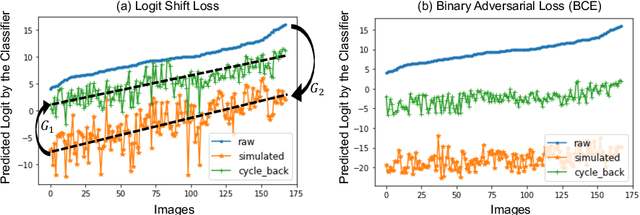
Abstract:Interpretability is a critical factor in applying complex deep learning models to advance the understanding of brain disorders in neuroimaging studies. To interpret the decision process of a trained classifier, existing techniques typically rely on saliency maps to quantify the voxel-wise or feature-level importance for classification through partial derivatives. Despite providing some level of localization, these maps are not human-understandable from the neuroscience perspective as they do not inform the specific meaning of the alteration linked to the brain disorder. Inspired by the image-to-image translation scheme, we propose to train simulator networks that can warp a given image to inject or remove patterns of the disease. These networks are trained such that the classifier produces consistently increased or decreased prediction logits for the simulated images. Moreover, we propose to couple all the simulators into a unified model based on conditional convolution. We applied our approach to interpreting classifiers trained on a synthetic dataset and two neuroimaging datasets to visualize the effect of the Alzheimer's disease and alcohol use disorder. Compared to the saliency maps generated by baseline approaches, our simulations and visualizations based on the Jacobian determinants of the warping field reveal meaningful and understandable patterns related to the diseases.
LSSL: Longitudinal Self-Supervised Learning
Jun 12, 2020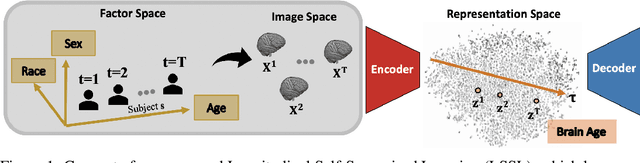
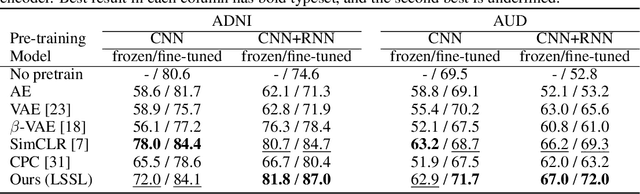
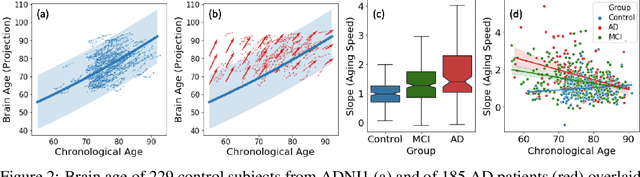
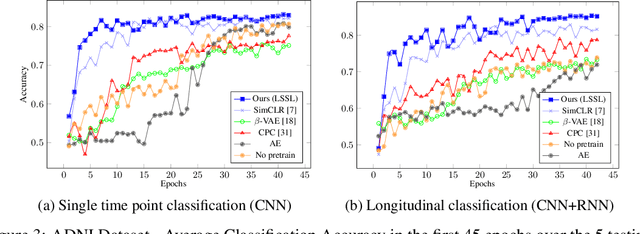
Abstract:Longitudinal neuroimaging or biomedical studies often acquire multiple observations from each individual over time, which entails repeated measures with highly interdependent variables. In this paper, we discuss the implication of repeated measures design on unsupervised learning by showing its tight conceptual connection to self-supervised learning and factor disentanglement. Leveraging the ability for `self-comparison' through repeated measures, we explicitly separate the definition of the factor space and the representation space enabling an exact disentanglement of time-related factors from the representations of the images. By formulating deterministic multivariate mapping functions between the two spaces, our model, named Longitudinal Self-Supervised Learning (LSSL), uses a standard autoencoding structure with a cosine loss to estimate the direction linked to the disentangled factor. We apply LSSL to two longitudinal neuroimaging studies to show its unique advantage in extracting the `brain-age' information from the data and in revealing informative characteristics associated with neurodegenerative and neuropsychological disorders. For a downstream task of supervised diagnosis classification, the representations learned by LSSL permit faster convergence and higher (or similar) prediction accuracy compared to several other representation learning techniques.
Recurrent Neural Networks with Longitudinal Pooling and Consistency Regularization
Mar 31, 2020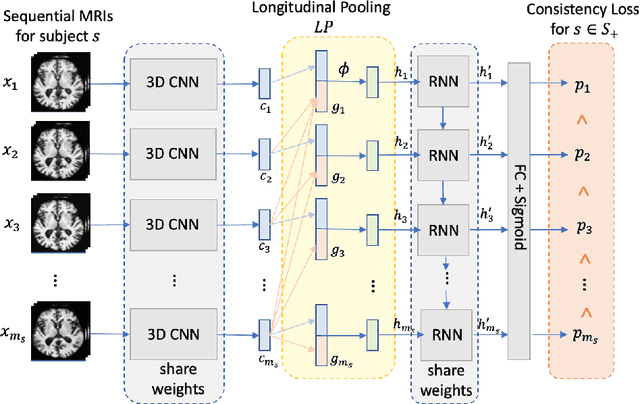
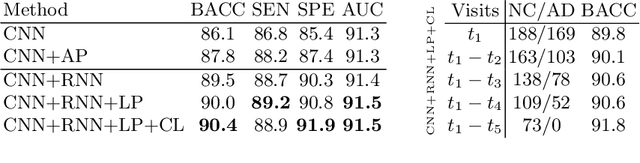
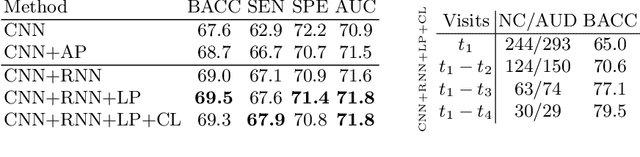

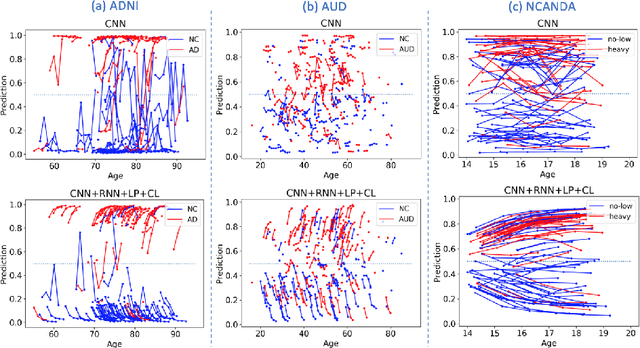

Abstract:Most neurological diseases are characterized by gradual deterioration of brain structure and function. To identify the impact of such diseases, studies have been acquiring large longitudinal MRI datasets and applied deep-learning to predict diagnosis label(s). These learning models apply Convolutional Neural Networks (CNN) to extract informative features from each time point of the longitudinal MRI and Recurrent Neural Networks (RNN) to classify each time point based on those features. However, they neglect the progressive nature of the disease, which may result in clinically implausible predictions across visits. In this paper, we propose a framework that injects the extracted features from CNNs at each time point to the RNN cells considering the dependencies across different time points in the longitudinal data. On the feature level, we propose a novel longitudinal pooling layer to couple features of a visit with those of proceeding ones. On the prediction level, we add a consistency regularization to the classification objective in line with the nature of the disease progression across visits. We evaluate the proposed method on the longitudinal structural MRIs from three neuroimaging datasets: Alzheimer's Disease Neuroimaging Initiative (ADNI, N=404), a dataset composed of 274 healthy controls and 329 patients with Alcohol Use Disorder (AUD), and 255 youths from the National Consortium on Alcohol and NeuroDevelopment in Adolescence (NCANDA). All three experiments show that our method is superior to the widely used methods. The code is available at https://github.com/ouyangjiahong/longitudinal-pooling.
Spatio-Temporal Graph Convolution for Functional MRI Analysis
Mar 24, 2020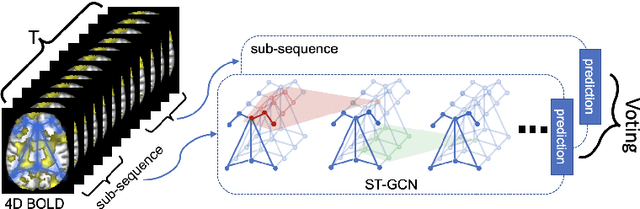
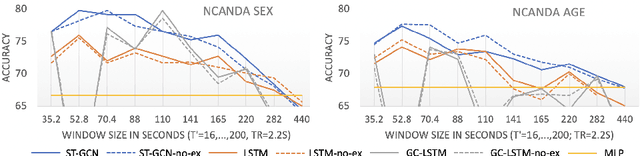
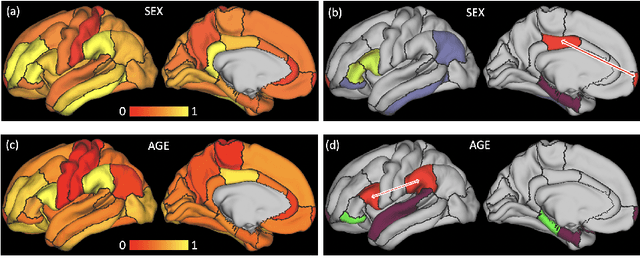
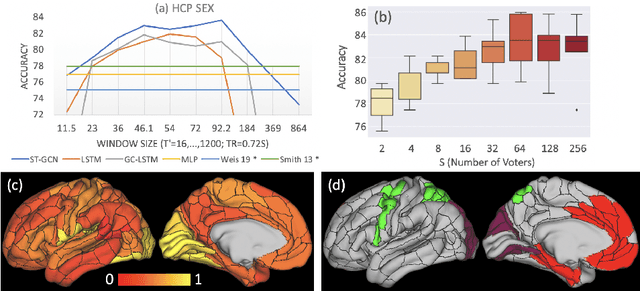
Abstract:The BOLD signal of resting-state fMRI (rs-fMRI) records the functional brain connectivity in a rich dynamic spatio-temporal setting. However, existing methods applied to rs-fMRI often fail to consider both spatial and temporal characteristics of the data. They either neglect the functional dependency between different brain regions in a network or discard the information in the temporal dynamics of brain activity. To overcome those shortcomings, we propose to formulate functional connectivity networks within the context of spatio-temporal graphs. We then train a spatio-temporal graph convolutional network (ST-GCN) on short sub-sequences of the BOLD time series to model the non-stationary nature of functional connectivity. We simultaneously learn the graph edge importance within ST-GCN to enable interpretation of functional connectivities contributing to the prediction model. In analyzing the rs-fMRI of the Human Connectome Project (HCP, N=1,091) and the National Consortium on Alcohol and Neurodevelopment in Adolescence (NCANDA, N=773), ST-GCN is significantly more accurate than common approaches in predicting gender and age based on BOLD signals. The matrix recording edge importance localizes brain regions and functional connections with significant aging and sex effects, which are verified by the neuroscience literature.
Bias-Resilient Neural Network
Oct 08, 2019
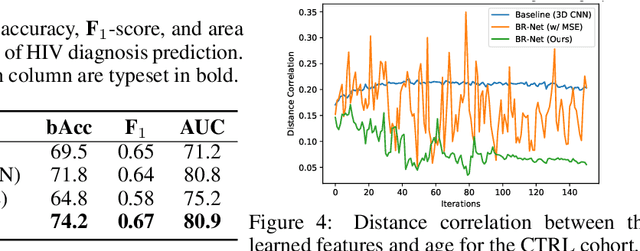
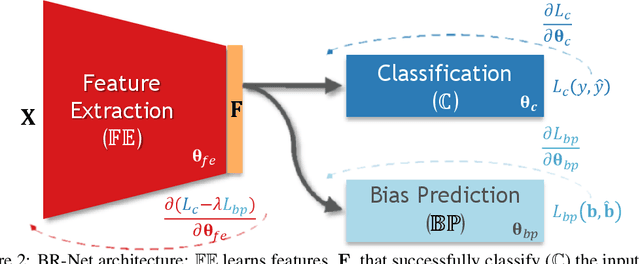
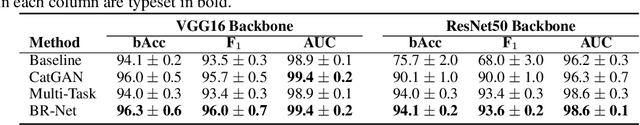
Abstract:Presence of bias and confounding effects is inarguably one of the most critical challenges in machine learning applications that has alluded to pivotal debates in the recent years. Such challenges range from spurious associations of confounding variables in medical studies to the bias of race in gender or face recognition systems. One solution is to enhance datasets and organize them such that they do not reflect biases, which is a cumbersome and intensive task. The alternative is to make use of available data and build models considering these biases. Traditional statistical methods apply straightforward techniques such as residualization or stratification to precomputed features to account for confounding variables. However, these techniques are generally not suitable for end-to-end deep learning methods. In this paper, we propose a method based on the adversarial training strategy to learn discriminative features unbiased and invariant to the confounder(s). This is enabled by incorporating a new adversarial loss function that encourages a vanished correlation between the bias and learned features. We apply our method to synthetic data, medical images, and a gender classification (Gender Shades Pilot Parliaments Benchmark) dataset. Our results show that the learned features by our method not only result in superior prediction performance but also are uncorrelated with the bias or confounder variables. The code is available at http://github.com/QingyuZhao/BR-Net/.
Confounder-Aware Visualization of ConvNets
Jul 30, 2019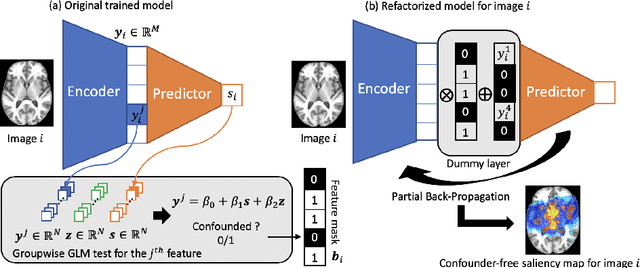
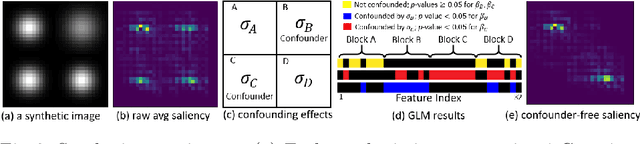
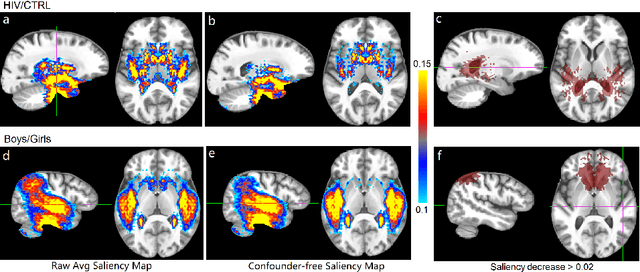
Abstract:With recent advances in deep learning, neuroimaging studies increasingly rely on convolutional networks (ConvNets) to predict diagnosis based on MR images. To gain a better understanding of how a disease impacts the brain, the studies visualize the salience maps of the ConvNet highlighting voxels within the brain majorly contributing to the prediction. However, these salience maps are generally confounded, i.e., some salient regions are more predictive of confounding variables (such as age) than the diagnosis. To avoid such misinterpretation, we propose in this paper an approach that aims to visualize confounder-free saliency maps that only highlight voxels predictive of the diagnosis. The approach incorporates univariate statistical tests to identify confounding effects within the intermediate features learned by ConvNet. The influence from the subset of confounded features is then removed by a novel partial back-propagation procedure. We use this two-step approach to visualize confounder-free saliency maps extracted from synthetic and two real datasets. These experiments reveal the potential of our visualization in producing unbiased model-interpretation.
Variational AutoEncoder For Regression: Application to Brain Aging Analysis
Apr 11, 2019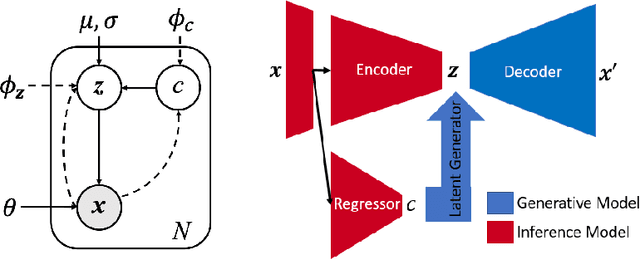
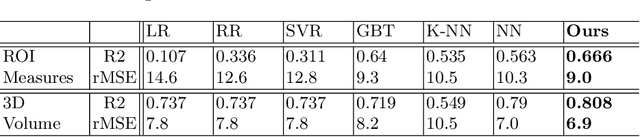
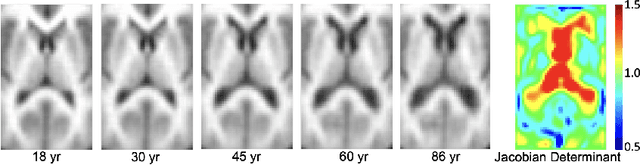
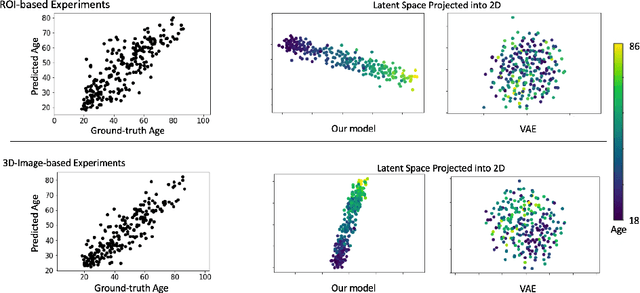
Abstract:While unsupervised variational autoencoders (VAE) have become a powerful tool in neuroimage analysis, their application to supervised learning is under-explored. We aim to close this gap by proposing a unified probabilistic model for learning the latent space of imaging data and performing supervised regression. Based on recent advances in learning disentangled representations, the novel generative process explicitly models the conditional distribution of latent representations with respect to the regression target variable. Performing a variational inference procedure on this model leads to joint regularization between the VAE and a neural-network regressor. In predicting the age of 245 subjects from their structural Magnetic Resonance (MR) images, our model is more accurate than state-of-the-art methods when applied to either region-of-interest (ROI) measurements or raw 3D volume images. More importantly, unlike simple feed-forward neural-networks, disentanglement of age in latent representations allows for intuitive interpretation of the structural developmental patterns of the human brain.
Truncated Gaussian-Mixture Variational AutoEncoder
Mar 08, 2019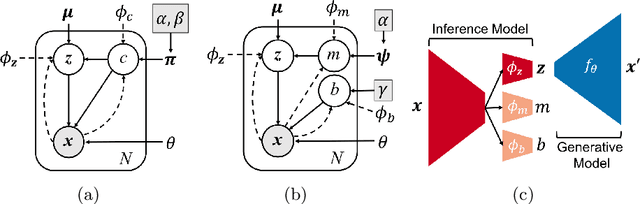
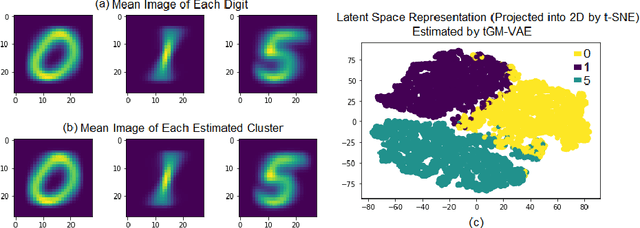
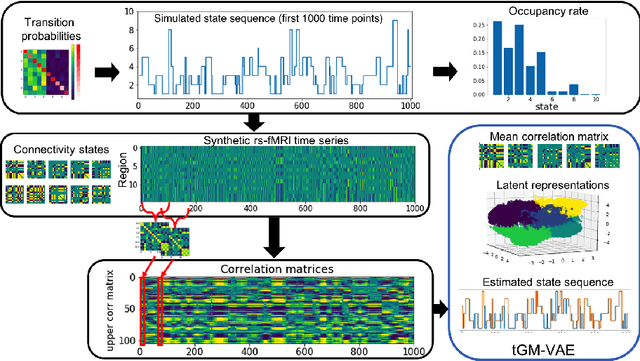
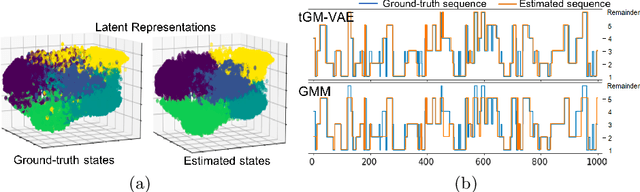
Abstract:Variation Autoencoder (VAE) has become a powerful tool in modeling the non-linear generative process of data from a low-dimensional latent space. Recently, several studies have proposed to use VAE for unsupervised clustering by using mixture models to capture the multi-modal structure of latent representations. This strategy, however, is ineffective when there are outlier data samples whose latent representations are meaningless, yet contaminating the estimation of key major clusters in the latent space. This exact problem arises in the context of resting-state fMRI (rs-fMRI) analysis, where clustering major functional connectivity patterns is often hindered by heavy noise of rs-fMRI and many minor clusters (rare connectivity patterns) of no interest to analysis. In this paper we propose a novel generative process, in which we use a Gaussian-mixture to model a few major clusters in the data, and use a non-informative uniform distribution to capture the remaining data. We embed this truncated Gaussian-Mixture model in a Variational AutoEncoder framework to obtain a general joint clustering and outlier detection approach, called tGM-VAE. We demonstrated the applicability of tGM-VAE on the MNIST dataset and further validated it in the context of rs-fMRI connectivity analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge