Paul Aljabar
Confidence Matters: Uncertainty Quantification and Precision Assessment of Deep Learning-based CMR Biomarker Estimates Using Scan-rescan Data
Mar 25, 2026Abstract:The performance of deep learning (DL) methods for the analysis of cine cardiovascular magnetic resonance (CMR) is typically assessed in terms of accuracy, overlooking precision. In this work, uncertainty estimation techniques, namely deep ensemble, test-time augmentation, and Monte Carlo dropout, are applied to a state-of-the-art DL pipeline for cardiac functional biomarker estimation, and new distribution-based metrics are proposed for the assessment of biomarker precision. The model achieved high accuracy (average Dice 87%) and point estimate precision on two external validation scan-rescan CMR datasets. However, distribution-based metrics showed that the overlap between scan/rescan confidence intervals was >50% in less than 45% of the cases. Statistical similarity tests between scan and rescan biomarkers also resulted in significant differences for over 65% of the cases. We conclude that, while point estimate metrics might suggest good performance, distributional analyses reveal lower precision, highlighting the need to use more representative metrics to assess scan-rescan agreement.
Improving the Scan-rescan Precision of AI-based CMR Biomarker Estimation
Aug 21, 2024Abstract:Quantification of cardiac biomarkers from cine cardiovascular magnetic resonance (CMR) data using deep learning (DL) methods offers many advantages, such as increased accuracy and faster analysis. However, only a few studies have focused on the scan-rescan precision of the biomarker estimates, which is important for reproducibility and longitudinal analysis. Here, we propose a cardiac biomarker estimation pipeline that not only focuses on achieving high segmentation accuracy but also on improving the scan-rescan precision of the computed biomarkers, namely left and right ventricular ejection fraction, and left ventricular myocardial mass. We evaluate two approaches to improve the apical-basal resolution of the segmentations used for estimating the biomarkers: one based on image interpolation and one based on segmentation interpolation. Using a database comprising scan-rescan cine CMR data acquired from 92 subjects, we compare the performance of these two methods against ground truth (GT) segmentations and DL segmentations obtained before interpolation (baseline). The results demonstrate that both the image-based and segmentation-based interpolation methods were able to narrow Bland-Altman scan-rescan confidence intervals for all biomarkers compared to the GT and baseline performances. Our findings highlight the importance of focusing not only on segmentation accuracy but also on the consistency of biomarkers across repeated scans, which is crucial for longitudinal analysis of cardiac function.
SVRDA: A Web-based Dataset Annotation Tool for Slice-to-Volume Registration
Nov 27, 2023



Abstract:Background and Objective: The lack of benchmark datasets has impeded the development of slice-to-volume registration algorithms. Such datasets are difficult to annotate, primarily due to the dimensional difference within data and the dearth of task-specific software. We aim to develop a user-friendly tool to streamline dataset annotation for slice-to-volume registration. Methods: The proposed tool, named SVRDA, is an installation-free web application for platform-agnostic collaborative dataset annotation. It enables efficient transformation manipulation via keyboard shortcuts and smooth case transitions with auto-saving. SVRDA supports configuration-based data loading and adheres to the separation of concerns, offering great flexibility and extensibility for future research. Various supplementary features have been implemented to facilitate slice-to-volume registration. Results: We validated the effectiveness of SVRDA by indirectly evaluating the post-registration segmentation quality on UK Biobank data, observing a dramatic overall improvement (24.02% in the Dice Similarity Coefficient and 48.93% in the 95th percentile Hausdorff distance, respectively) supported by highly statistically significant evidence ($p<0.001$).We further showcased the clinical usage of SVRDA by integrating it into test-retest T1 quantification on in-house magnetic resonance images, leading to more consistent results after registration. Conclusions: SVRDA can facilitate collaborative annotation of benchmark datasets while being potentially applicable to other pipelines incorporating slice-to-volume registration. Full source code and documentation are available at https://github.com/Roldbach/SVRDA
Multi-Atlas Segmentation using Partially Annotated Data: Methods and Annotation Strategies
Apr 29, 2016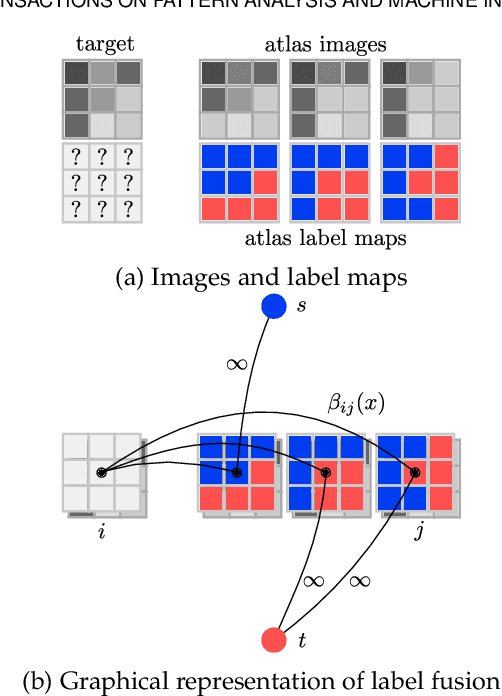
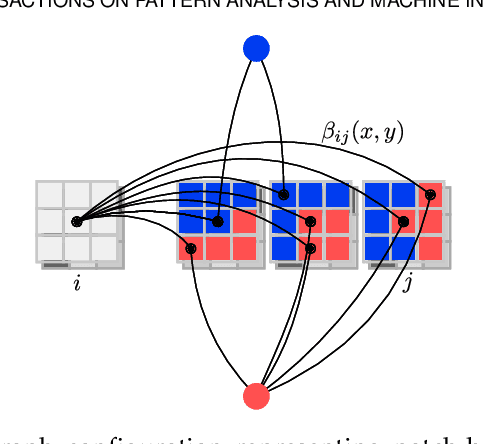
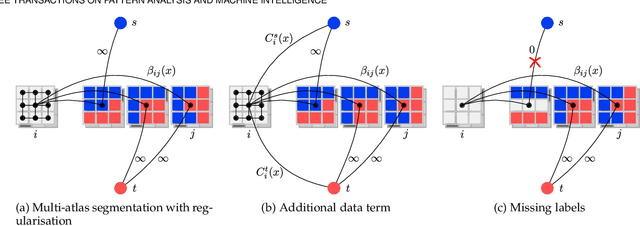
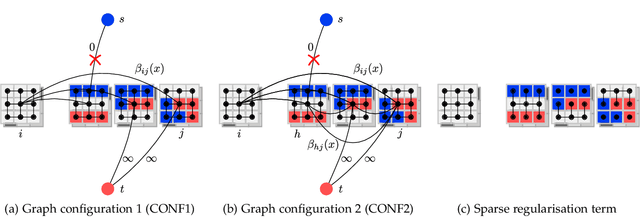
Abstract:Multi-atlas segmentation is a widely used tool in medical image analysis, providing robust and accurate results by learning from annotated atlas datasets. However, the availability of fully annotated atlas images for training is limited due to the time required for the labelling task. Segmentation methods requiring only a proportion of each atlas image to be labelled could therefore reduce the workload on expert raters tasked with annotating atlas images. To address this issue, we first re-examine the labelling problem common in many existing approaches and formulate its solution in terms of a Markov Random Field energy minimisation problem on a graph connecting atlases and the target image. This provides a unifying framework for multi-atlas segmentation. We then show how modifications in the graph configuration of the proposed framework enable the use of partially annotated atlas images and investigate different partial annotation strategies. The proposed method was evaluated on two Magnetic Resonance Imaging (MRI) datasets for hippocampal and cardiac segmentation. Experiments were performed aimed at (1) recreating existing segmentation techniques with the proposed framework and (2) demonstrating the potential of employing sparsely annotated atlas data for multi-atlas segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge