Or Zuk
Hebrew University of Jerusalem
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
Mar 10, 2026Abstract:Recovering a tree that represents the evolutionary history of a group of species is a key task in phylogenetics. Performing this task using sequence data from multiple genetic markers poses two key challenges. The first is the discordance between the evolutionary history of individual genes and that of the species. The second challenge is computational, as contemporary studies involve thousands of species. Here we present SDSR, a scalable divide-and-conquer approach for species tree reconstruction based on spectral graph theory. The algorithm recursively partitions the species into subsets until their sizes are below a given threshold. The trees of these subsets are reconstructed by a user-chosen species tree algorithm. Finally, these subtrees are merged to form the full tree. On the theoretical front, we derive recovery guarantees for SDSR, under the multispecies coalescent (MSC) model. We also perform a runtime complexity analysis. We show that SDSR, when combined with a species tree reconstruction algorithm as a subroutine, yields substantial runtime savings as compared to applying the same algorithm on the full data. Empirically, we evaluate SDSR on synthetic benchmark datasets with incomplete lineage sorting and horizontal gene transfer. In accordance with our theoretical analysis, the simulations show that combining SDSR with common species tree methods, such as CA-ML or ASTRAL, yields up to 10-fold faster runtimes. In addition, SDSR achieves a comparable tree reconstruction accuracy to that obtained by applying these methods on the full data.
Clustering Noisy Signals with Structured Sparsity Using Time-Frequency Representation
Oct 18, 2015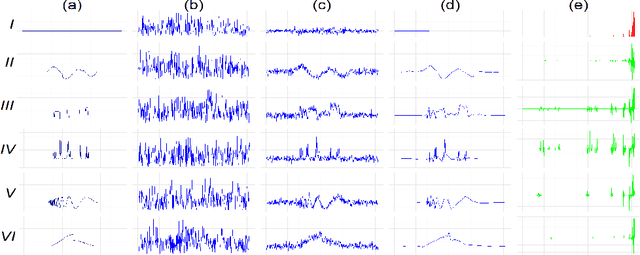
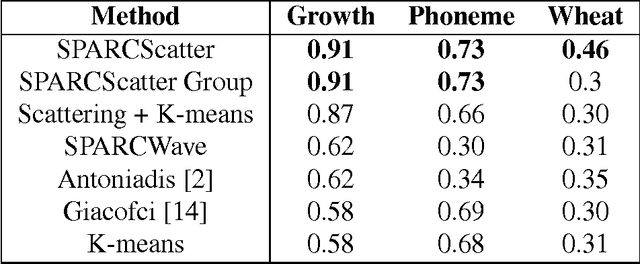
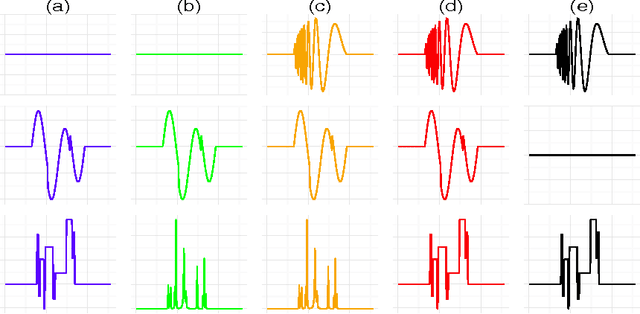
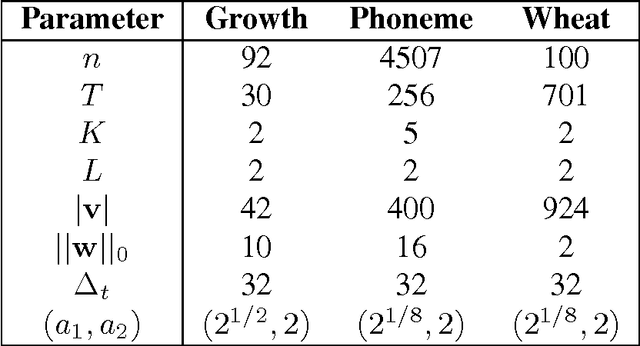
Abstract:We propose a simple and efficient time-series clustering framework particularly suited for low Signal-to-Noise Ratio (SNR), by simultaneous smoothing and dimensionality reduction aimed at preserving clustering information. We extend the sparse K-means algorithm by incorporating structured sparsity, and use it to exploit the multi-scale property of wavelets and group structure in multivariate signals. Finally, we extract features invariant to translation and scaling with the scattering transform, which corresponds to a convolutional network with filters given by a wavelet operator, and use the network's structure in sparse clustering. By promoting sparsity, this transform can yield a low-dimensional representation of signals that gives improved clustering results on several real datasets.
Low-Rank Matrix Recovery from Row-and-Column Affine Measurements
May 23, 2015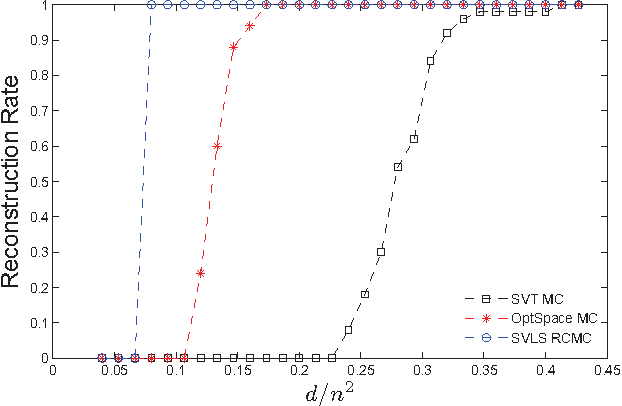
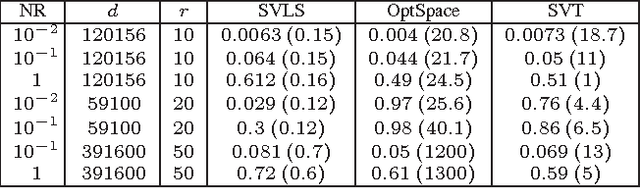
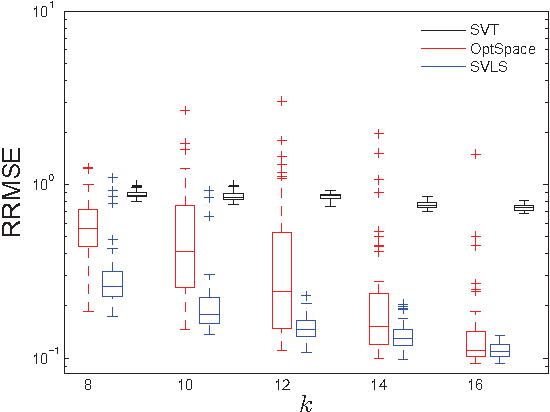
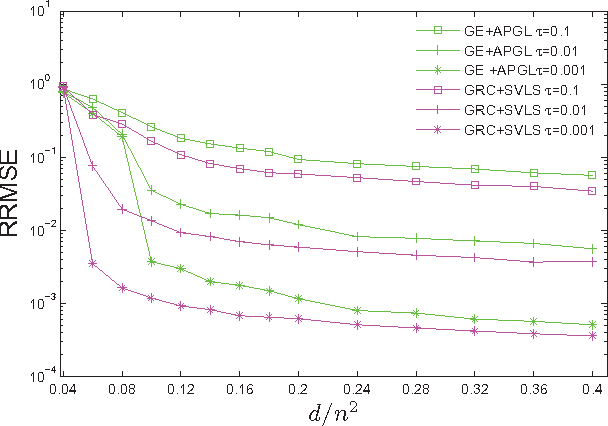
Abstract:We propose and study a row-and-column affine measurement scheme for low-rank matrix recovery. Each measurement is a linear combination of elements in one row or one column of a matrix $X$. This setting arises naturally in applications from different domains. However, current algorithms developed for standard matrix recovery problems do not perform well in our case, hence the need for developing new algorithms and theory for our problem. We propose a simple algorithm for the problem based on Singular Value Decomposition ($SVD$) and least-squares ($LS$), which we term \alg. We prove that (a simplified version of) our algorithm can recover $X$ exactly with the minimum possible number of measurements in the noiseless case. In the general noisy case, we prove performance guarantees on the reconstruction accuracy under the Frobenius norm. In simulations, our row-and-column design and \alg algorithm show improved speed, and comparable and in some cases better accuracy compared to standard measurements designs and algorithms. Our theoretical and experimental results suggest that the proposed row-and-column affine measurements scheme, together with our recovery algorithm, may provide a powerful framework for affine matrix reconstruction.
On the Number of Samples Needed to Learn the Correct Structure of a Bayesian Network
Jun 27, 2012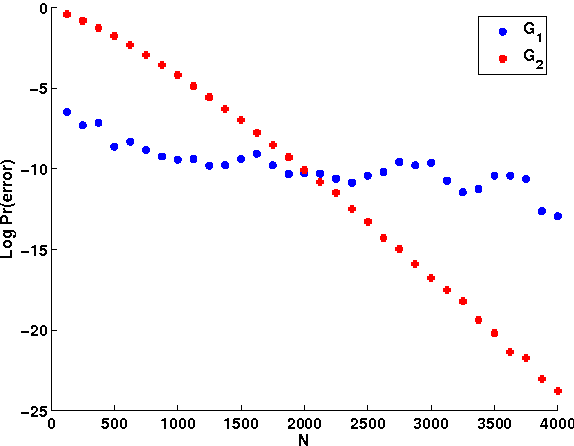
Abstract:Bayesian Networks (BNs) are useful tools giving a natural and compact representation of joint probability distributions. In many applications one needs to learn a Bayesian Network (BN) from data. In this context, it is important to understand the number of samples needed in order to guarantee a successful learning. Previous work have studied BNs sample complexity, yet it mainly focused on the requirement that the learned distribution will be close to the original distribution which generated the data. In this work, we study a different aspect of the learning, namely the number of samples needed in order to learn the correct structure of the network. We give both asymptotic results, valid in the large sample limit, and experimental results, demonstrating the learning behavior for feasible sample sizes. We show that structure learning is a more difficult task, compared to approximating the correct distribution, in the sense that it requires a much larger number of samples, regardless of the computational power available for the learner.
Ranking Under Uncertainty
Jun 20, 2012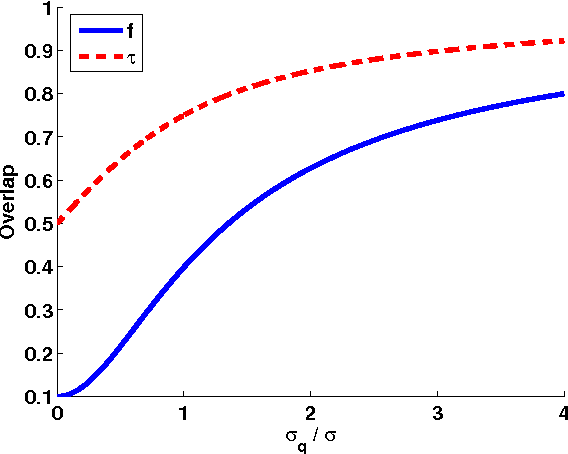
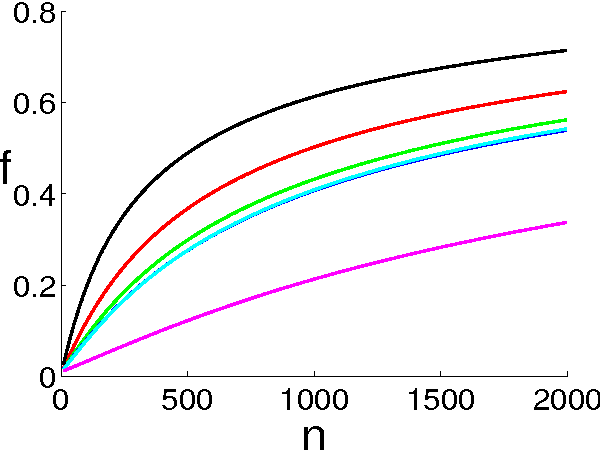
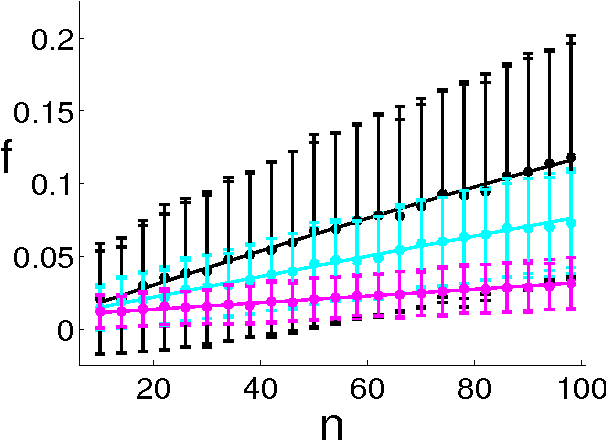
Abstract:Ranking objects is a simple and natural procedure for organizing data. It is often performed by assigning a quality score to each object according to its relevance to the problem at hand. Ranking is widely used for object selection, when resources are limited and it is necessary to select a subset of most relevant objects for further processing. In real world situations, the object's scores are often calculated from noisy measurements, casting doubt on the ranking reliability. We introduce an analytical method for assessing the influence of noise levels on the ranking reliability. We use two similarity measures for reliability evaluation, Top-K-List overlap and Kendall's tau measure, and show that the former is much more sensitive to noise than the latter. We apply our method to gene selection in a series of microarray experiments of several cancer types. The results indicate that the reliability of the lists obtained from these experiments is very poor, and that experiment sizes which are necessary for attaining reasonably stable Top-K-Lists are much larger than those currently available. Simulations support our analytical results.
Rare-Allele Detection Using Compressed Se(que)nsing
Sep 02, 2009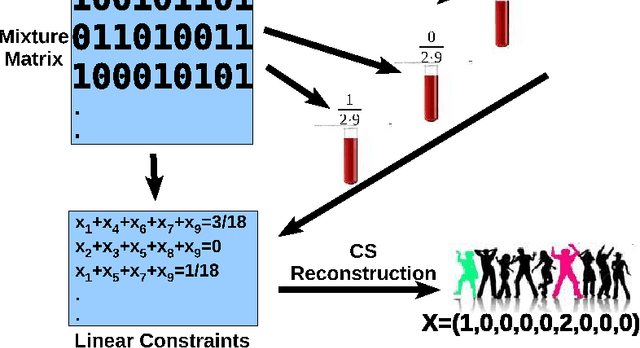
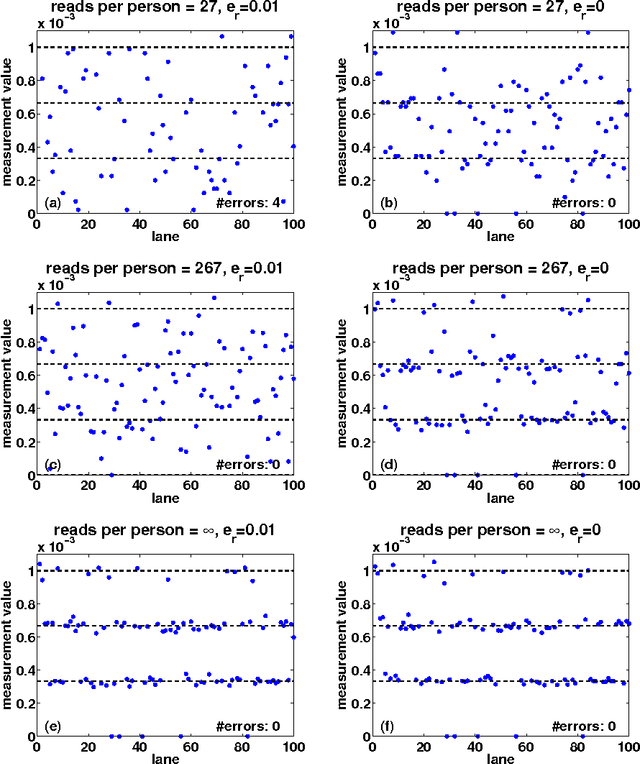
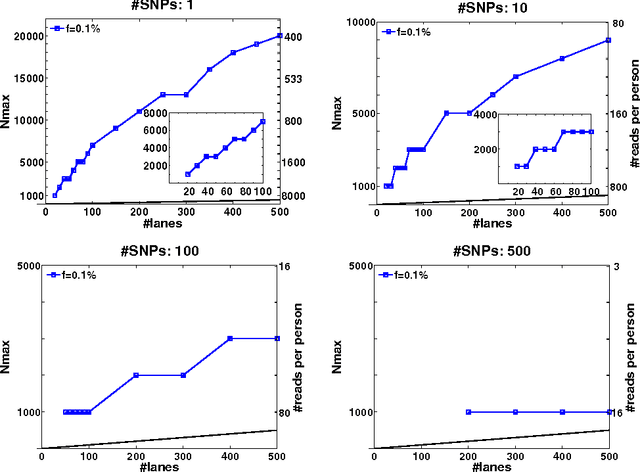
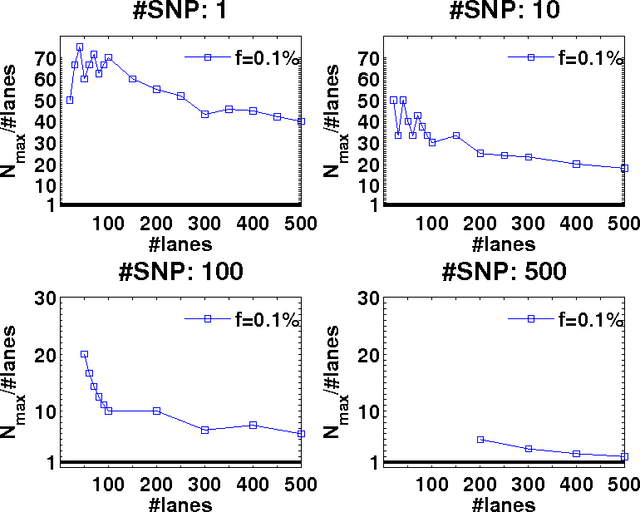
Abstract:Detection of rare variants by resequencing is important for the identification of individuals carrying disease variants. Rapid sequencing by new technologies enables low-cost resequencing of target regions, although it is still prohibitive to test more than a few individuals. In order to improve cost trade-offs, it has recently been suggested to apply pooling designs which enable the detection of carriers of rare alleles in groups of individuals. However, this was shown to hold only for a relatively low number of individuals in a pool, and requires the design of pooling schemes for particular cases. We propose a novel pooling design, based on a compressed sensing approach, which is both general, simple and efficient. We model the experimental procedure and show via computer simulations that it enables the recovery of rare allele carriers out of larger groups than were possible before, especially in situations where high coverage is obtained for each individual. Our approach can also be combined with barcoding techniques to enhance performance and provide a feasible solution based on current resequencing costs. For example, when targeting a small enough genomic region (~100 base-pairs) and using only ~10 sequencing lanes and ~10 distinct barcodes, one can recover the identity of 4 rare allele carriers out of a population of over 4000 individuals.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge