Ariel Jaffe
Hebrew University of Jerusalem
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
Mar 10, 2026Abstract:Recovering a tree that represents the evolutionary history of a group of species is a key task in phylogenetics. Performing this task using sequence data from multiple genetic markers poses two key challenges. The first is the discordance between the evolutionary history of individual genes and that of the species. The second challenge is computational, as contemporary studies involve thousands of species. Here we present SDSR, a scalable divide-and-conquer approach for species tree reconstruction based on spectral graph theory. The algorithm recursively partitions the species into subsets until their sizes are below a given threshold. The trees of these subsets are reconstructed by a user-chosen species tree algorithm. Finally, these subtrees are merged to form the full tree. On the theoretical front, we derive recovery guarantees for SDSR, under the multispecies coalescent (MSC) model. We also perform a runtime complexity analysis. We show that SDSR, when combined with a species tree reconstruction algorithm as a subroutine, yields substantial runtime savings as compared to applying the same algorithm on the full data. Empirically, we evaluate SDSR on synthetic benchmark datasets with incomplete lineage sorting and horizontal gene transfer. In accordance with our theoretical analysis, the simulations show that combining SDSR with common species tree methods, such as CA-ML or ASTRAL, yields up to 10-fold faster runtimes. In addition, SDSR achieves a comparable tree reconstruction accuracy to that obtained by applying these methods on the full data.
Graph-based Semi-Supervised Learning via Maximum Discrimination
Feb 08, 2026Abstract:Semi-supervised learning (SSL) addresses the critical challenge of training accurate models when labeled data is scarce but unlabeled data is abundant. Graph-based SSL (GSSL) has emerged as a popular framework that captures data structure through graph representations. Classic graph SSL methods, such as Label Propagation and Label Spreading, aim to compute low-dimensional representations where points with the same labels are close in representation space. Although often effective, these methods can be suboptimal on data with complex label distributions. In our work, we develop AUC-spec, a graph approach that computes a low-dimensional representation that maximizes class separation. We compute this representation by optimizing the Area Under the ROC Curve (AUC) as estimated via the labeled points. We provide a detailed analysis of our approach under a product-of-manifold model, and show that the required number of labeled points for AUC-spec is polynomial in the model parameters. Empirically, we show that AUC-spec balances class separation with graph smoothness. It demonstrates competitive results on synthetic and real-world datasets while maintaining computational efficiency comparable to the field's classic and state-of-the-art methods.
Spectral Self-supervised Feature Selection
Jul 12, 2024Abstract:Choosing a meaningful subset of features from high-dimensional observations in unsupervised settings can greatly enhance the accuracy of downstream analysis, such as clustering or dimensionality reduction, and provide valuable insights into the sources of heterogeneity in a given dataset. In this paper, we propose a self-supervised graph-based approach for unsupervised feature selection. Our method's core involves computing robust pseudo-labels by applying simple processing steps to the graph Laplacian's eigenvectors. The subset of eigenvectors used for computing pseudo-labels is chosen based on a model stability criterion. We then measure the importance of each feature by training a surrogate model to predict the pseudo-labels from the observations. Our approach is shown to be robust to challenging scenarios, such as the presence of outliers and complex substructures. We demonstrate the effectiveness of our method through experiments on real-world datasets, showing its robustness across multiple domains, particularly its effectiveness on biological datasets.
Multi-modal Differentiable Unsupervised Feature Selection
Mar 16, 2023



Abstract:Multi-modal high throughput biological data presents a great scientific opportunity and a significant computational challenge. In multi-modal measurements, every sample is observed simultaneously by two or more sets of sensors. In such settings, many observed variables in both modalities are often nuisance and do not carry information about the phenomenon of interest. Here, we propose a multi-modal unsupervised feature selection framework: identifying informative variables based on coupled high-dimensional measurements. Our method is designed to identify features associated with two types of latent low-dimensional structures: (i) shared structures that govern the observations in both modalities and (ii) differential structures that appear in only one modality. To that end, we propose two Laplacian-based scoring operators. We incorporate the scores with differentiable gates that mask nuisance features and enhance the accuracy of the structure captured by the graph Laplacian. The performance of the new scheme is illustrated using synthetic and real datasets, including an extended biological application to single-cell multi-omics.
DiSC: Differential Spectral Clustering of Features
Nov 10, 2022



Abstract:Selecting subsets of features that differentiate between two conditions is a key task in a broad range of scientific domains. In many applications, the features of interest form clusters with similar effects on the data at hand. To recover such clusters we develop DiSC, a data-driven approach for detecting groups of features that differentiate between conditions. For each condition, we construct a graph whose nodes correspond to the features and whose weights are functions of the similarity between them for that condition. We then apply a spectral approach to compute subsets of nodes whose connectivity differs significantly between the condition-specific feature graphs. On the theoretical front, we analyze our approach with a toy example based on the stochastic block model. We evaluate DiSC on a variety of datasets, including MNIST, hyperspectral imaging, simulated scRNA-seq and task fMRI, and demonstrate that DiSC uncovers features that better differentiate between conditions compared to competing methods.
Weakly Supervised Indoor Localization via Manifold Matching
Feb 07, 2022



Abstract:Inferring the location of a mobile device in an indoor setting is an open problem of utmost significance. A leading approach that does not require the deployment of expensive infrastructure is fingerprinting, where a classifier is trained to predict the location of a device based on its captured signal. The main caveat of this approach is that acquiring a sufficiently large and accurate training set may be prohibitively expensive. Here, we propose a weakly supervised method that only requires the location of a small number of devices. The localization is done by matching a low-dimensional spectral representation of the signals to a given sketch of the indoor environment. We test our approach on simulated and real data and show that it yields an accuracy of a few meters, which is on par with fully supervised approaches. The simplicity of our method and its accuracy with minimal supervision makes it ideal for implementation in indoor localization systems.
Spectral Top-Down Recovery of Latent Tree Models
Feb 26, 2021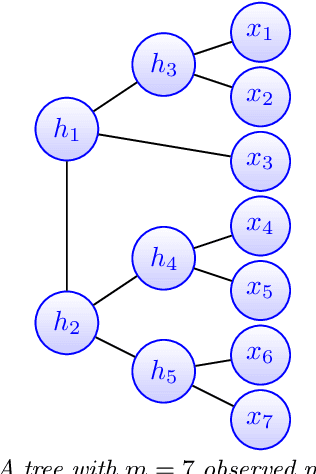
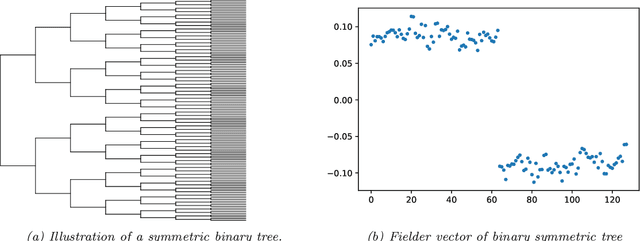
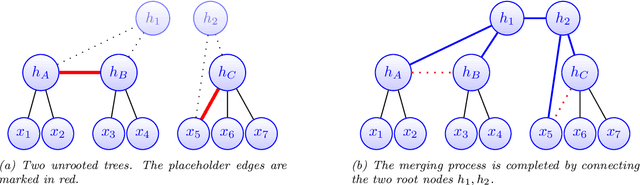
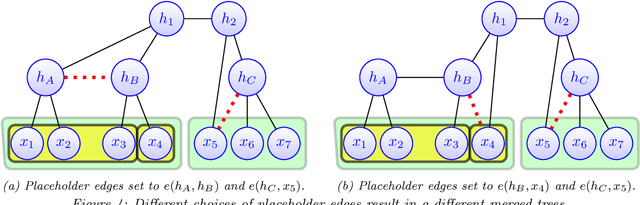
Abstract:Modeling the distribution of high dimensional data by a latent tree graphical model is a common approach in multiple scientific domains. A common task is to infer the underlying tree structure given only observations of the terminal nodes. Many algorithms for tree recovery are computationally intensive, which limits their applicability to trees of moderate size. For large trees, a common approach, termed divide-and-conquer, is to recover the tree structure in two steps. First, recover the structure separately for multiple randomly selected subsets of the terminal nodes. Second, merge the resulting subtrees to form a full tree. Here, we develop Spectral Top-Down Recovery (STDR), a divide-and-conquer approach for inference of large latent tree models. Unlike previous methods, STDR's partitioning step is non-random. Instead, it is based on the Fiedler vector of a suitable Laplacian matrix related to the observed nodes. We prove that under certain conditions this partitioning is consistent with the tree structure. This, in turn leads to a significantly simpler merging procedure of the small subtrees. We prove that STDR is statistically consistent, and bound the number of samples required to accurately recover the tree with high probability. Using simulated data from several common tree models in phylogenetics, we demonstrate that STDR has a significant advantage in terms of runtime, with improved or similar accuracy.
Spectral neighbor joining for reconstruction of latent tree models
Feb 28, 2020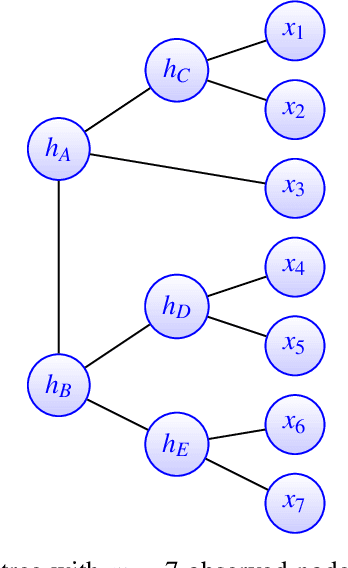
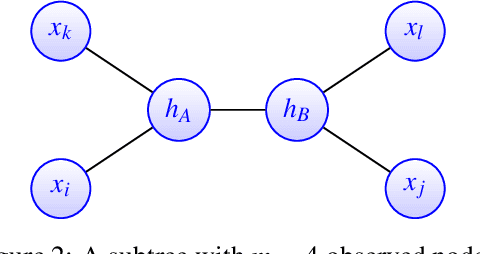
Abstract:A key assumption in multiple scientific applications is that the distribution of observed data can be modeled by a latent tree graphical model. An important example is phylogenetics, where the tree models the evolutionary lineages of various organisms. Given a set of independent realizations of the random variables at the leaves of the tree, a common task is to infer the underlying tree topology. In this work we develop Spectral Neighbor Joining (SNJ), a novel method to recover latent tree graphical models. In contrast to distance based methods, SNJ is based on a spectral measure of similarity between all pairs of observed variables. We prove that SNJ is consistent, and derive a sufficient condition for correct tree recovery from an estimated similarity matrix. Combining this condition with a concentration of measure result on the similarity matrix, we bound the number of samples required to recover the tree with high probability. We illustrate via extensive simulations that SNJ requires fewer samples to accurately recover trees in regimes where the tree contains a large number of leaves or long edges. We provide theoretical support for this observation by analyzing the model of a perfect binary tree.
The Spectral Underpinning of word2vec
Feb 27, 2020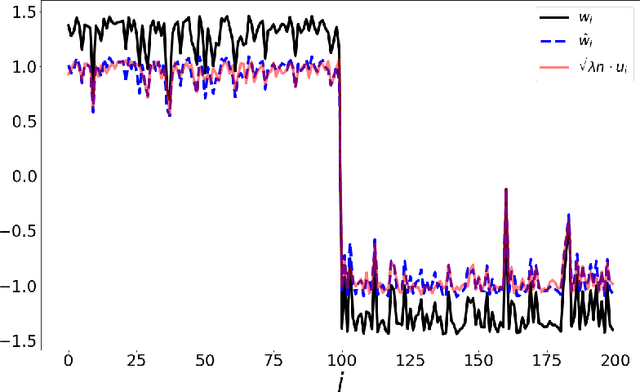
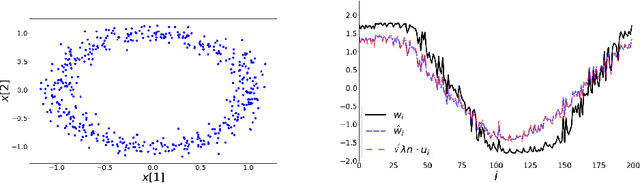
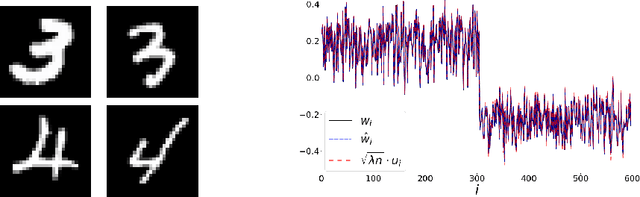
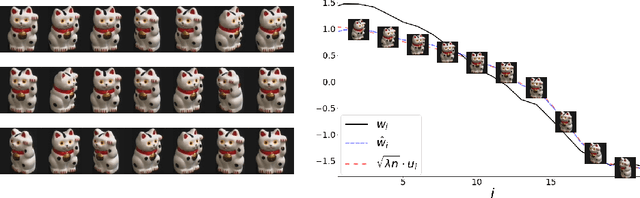
Abstract:word2vec due to Mikolov \textit{et al.} (2013) is a word embedding method that is widely used in natural language processing. Despite its great success and frequent use, theoretical justification is still lacking. The main contribution of our paper is to propose a rigorous analysis of the highly nonlinear functional of word2vec. Our results suggest that word2vec may be primarily driven by an underlying spectral method. This insight may open the door to obtaining provable guarantees for word2vec. We support these findings by numerical simulations. One fascinating open question is whether the nonlinear properties of word2vec that are not captured by the spectral method are beneficial and, if so, by what mechanism.
Learning Binary Latent Variable Models: A Tensor Eigenpair Approach
Feb 27, 2018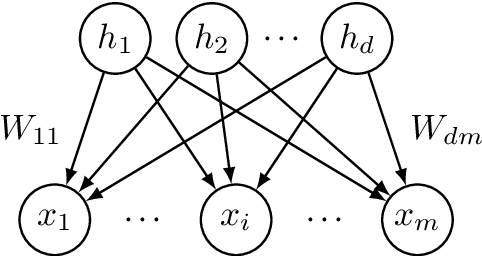
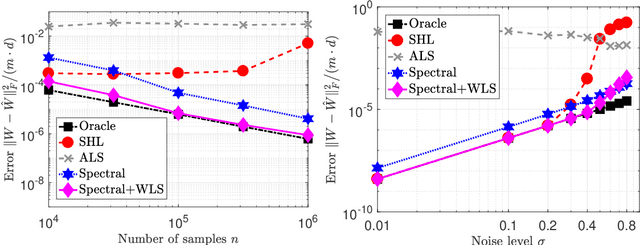
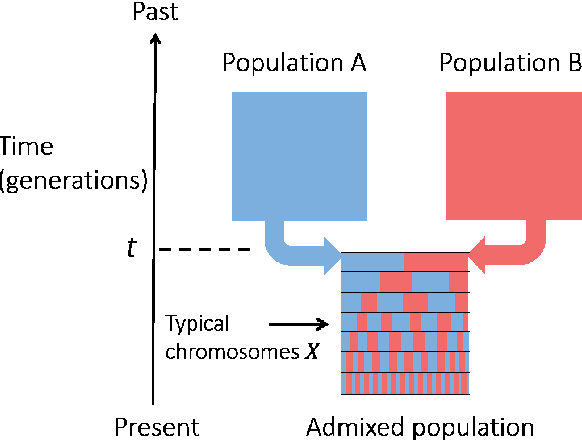
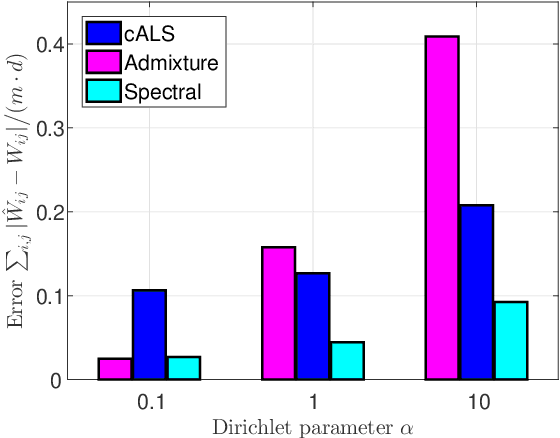
Abstract:Latent variable models with hidden binary units appear in various applications. Learning such models, in particular in the presence of noise, is a challenging computational problem. In this paper we propose a novel spectral approach to this problem, based on the eigenvectors of both the second order moment matrix and third order moment tensor of the observed data. We prove that under mild non-degeneracy conditions, our method consistently estimates the model parameters at the optimal parametric rate. Our tensor-based method generalizes previous orthogonal tensor decomposition approaches, where the hidden units were assumed to be either statistically independent or mutually exclusive. We illustrate the consistency of our method on simulated data and demonstrate its usefulness in learning a common model for population mixtures in genetics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge