Nikos Paragios
Therapanacea
Deforming Autoencoders: Unsupervised Disentangling of Shape and Appearance
Jun 18, 2018
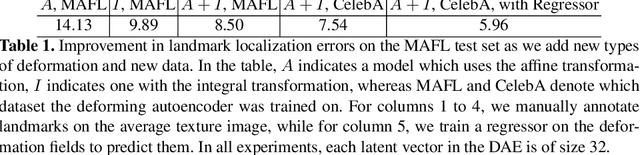

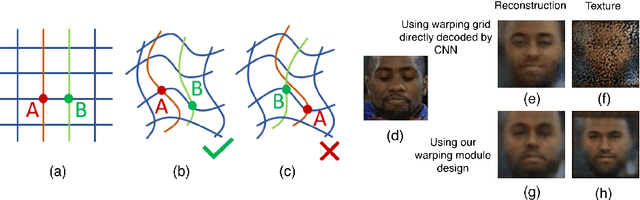
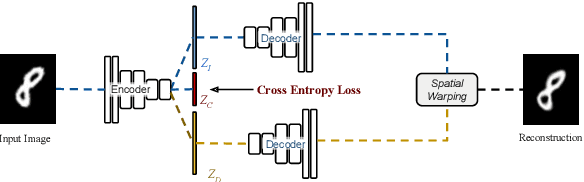
Abstract:In this work we introduce Deforming Autoencoders, a generative model for images that disentangles shape from appearance in an unsupervised manner. As in the deformable template paradigm, shape is represented as a deformation between a canonical coordinate system (`template') and an observed image, while appearance is modeled in `canonical', template, coordinates, thus discarding variability due to deformations. We introduce novel techniques that allow this approach to be deployed in the setting of autoencoders and show that this method can be used for unsupervised group-wise image alignment. We show experiments with expression morphing in humans, hands, and digits, face manipulation, such as shape and appearance interpolation, as well as unsupervised landmark localization. A more powerful form of unsupervised disentangling becomes possible in template coordinates, allowing us to successfully decompose face images into shading and albedo, and further manipulate face images.
Continuous Relaxation of MAP Inference: A Nonconvex Perspective
Feb 25, 2018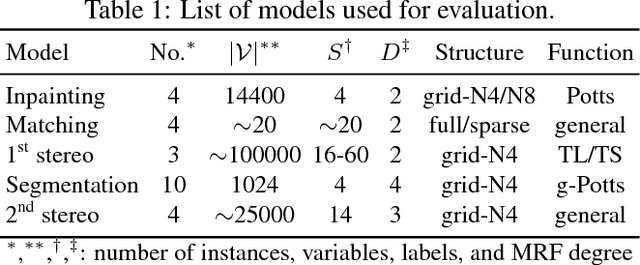
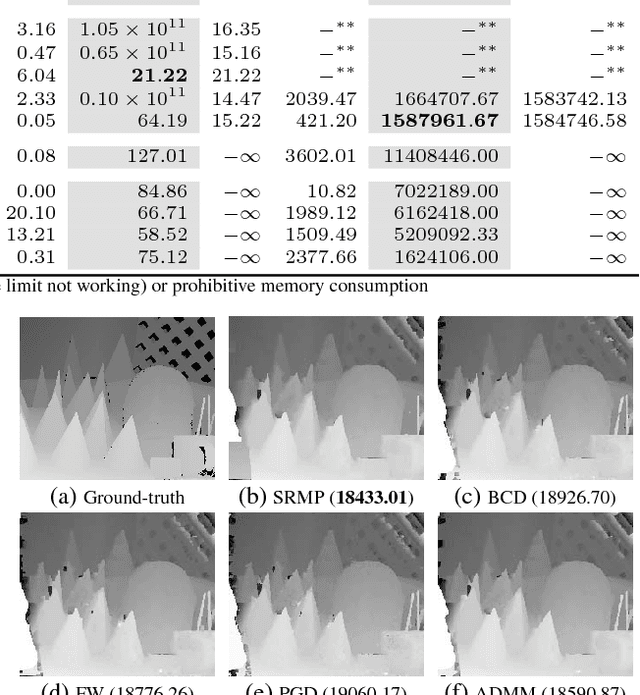
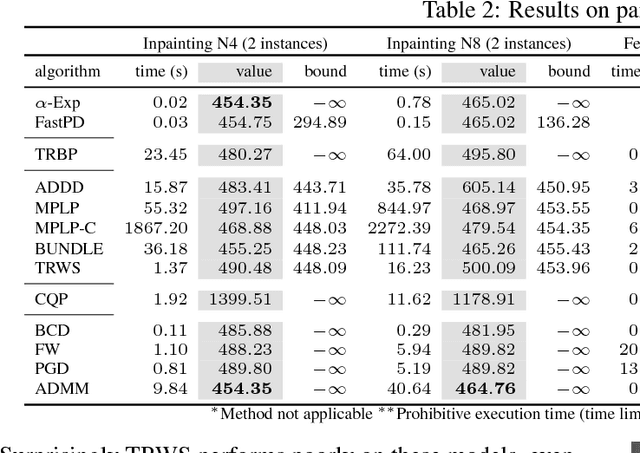
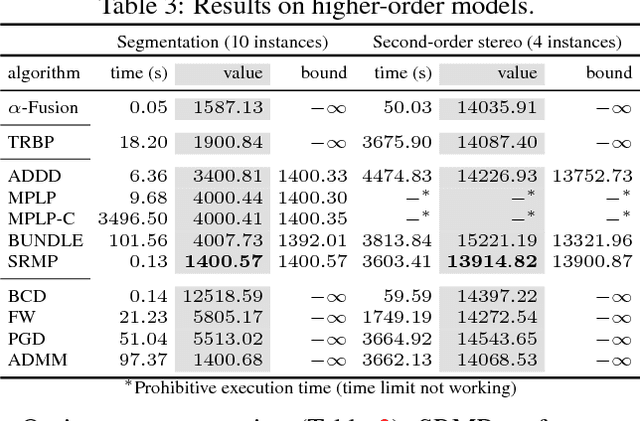
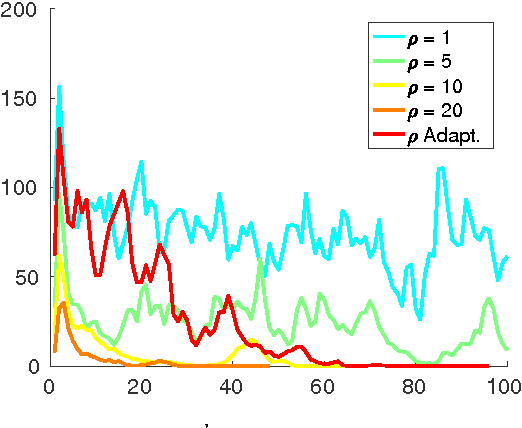
Abstract:In this paper, we study a nonconvex continuous relaxation of MAP inference in discrete Markov random fields (MRFs). We show that for arbitrary MRFs, this relaxation is tight, and a discrete stationary point of it can be easily reached by a simple block coordinate descent algorithm. In addition, we study the resolution of this relaxation using popular gradient methods, and further propose a more effective solution using a multilinear decomposition framework based on the alternating direction method of multipliers (ADMM). Experiments on many real-world problems demonstrate that the proposed ADMM significantly outperforms other nonconvex relaxation based methods, and compares favorably with state of the art MRF optimization algorithms in different settings.
Alternating Direction Graph Matching
Feb 23, 2018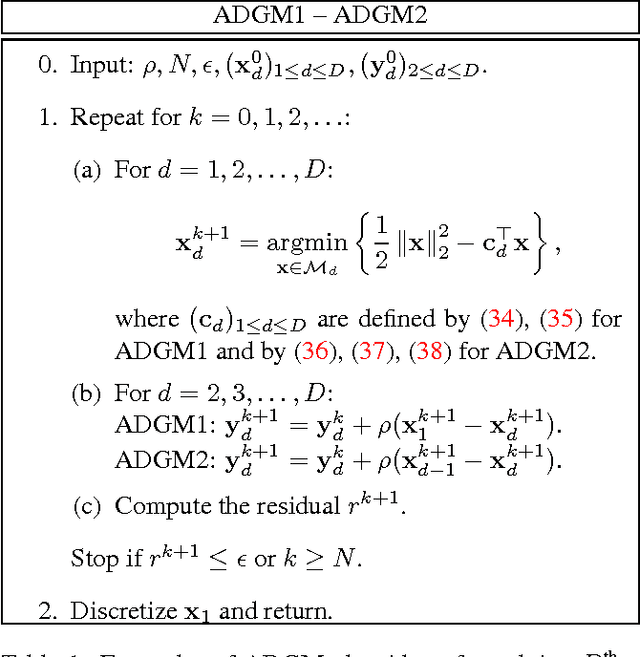
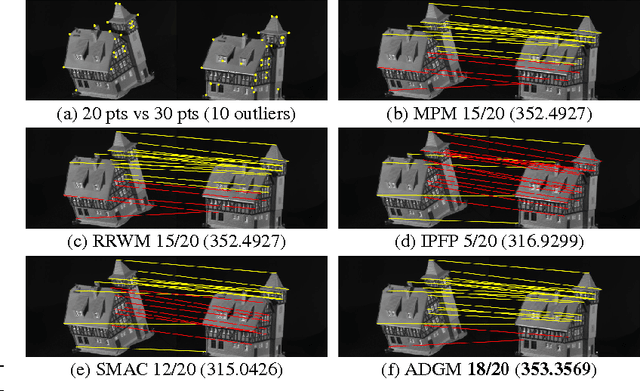
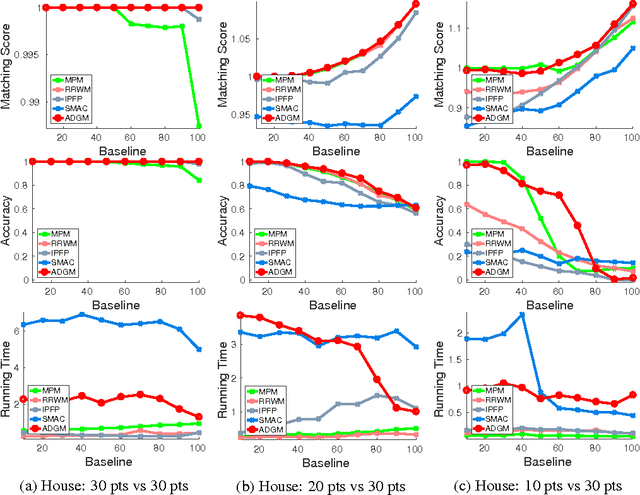
Abstract:In this paper, we introduce a graph matching method that can account for constraints of arbitrary order, with arbitrary potential functions. Unlike previous decomposition approaches that rely on the graph structures, we introduce a decomposition of the matching constraints. Graph matching is then reformulated as a non-convex non-separable optimization problem that can be split into smaller and much-easier-to-solve subproblems, by means of the alternating direction method of multipliers. The proposed framework is modular, scalable, and can be instantiated into different variants. Two instantiations are studied exploring pairwise and higher-order constraints. Experimental results on widely adopted benchmarks involving synthetic and real examples demonstrate that the proposed solutions outperform existing pairwise graph matching methods, and competitive with the state of the art in higher-order settings.
Newton-type Methods for Inference in Higher-Order Markov Random Fields
Sep 05, 2017
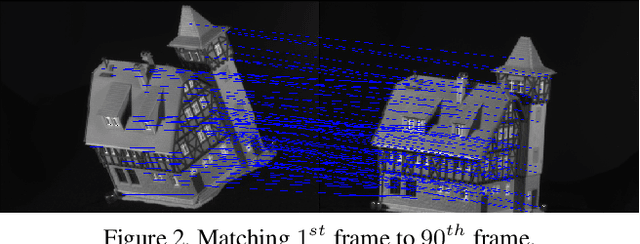
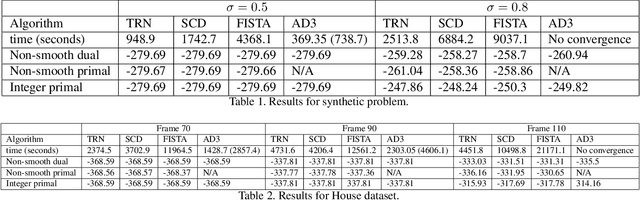


Abstract:Linear programming relaxations are central to {\sc map} inference in discrete Markov Random Fields. The ability to properly solve the Lagrangian dual is a critical component of such methods. In this paper, we study the benefit of using Newton-type methods to solve the Lagrangian dual of a smooth version of the problem. We investigate their ability to achieve superior convergence behavior and to better handle the ill-conditioned nature of the formulation, as compared to first order methods. We show that it is indeed possible to efficiently apply a trust region Newton method for a broad range of {\sc map} inference problems. In this paper we propose a provably convergent and efficient framework that includes (i) excellent compromise between computational complexity and precision concerning the Hessian matrix construction, (ii) a damping strategy that aids efficient optimization, (iii) a truncation strategy coupled with a generic pre-conditioner for Conjugate Gradients, (iv) efficient sum-product computation for sparse clique potentials. Results for higher-order Markov Random Fields demonstrate the potential of this approach.
* 10 pages, 3 figures, 3 tables, CVPR 2017
Deformable Registration through Learning of Context-Specific Metric Aggregation
Jul 19, 2017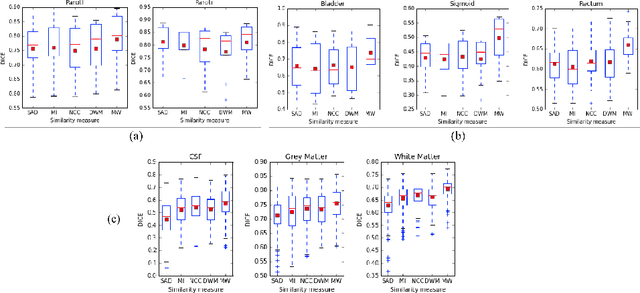
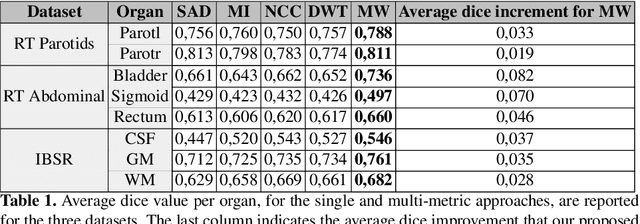
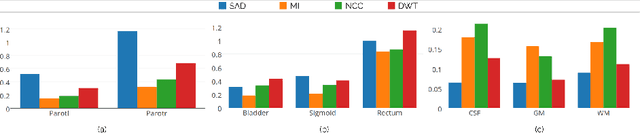
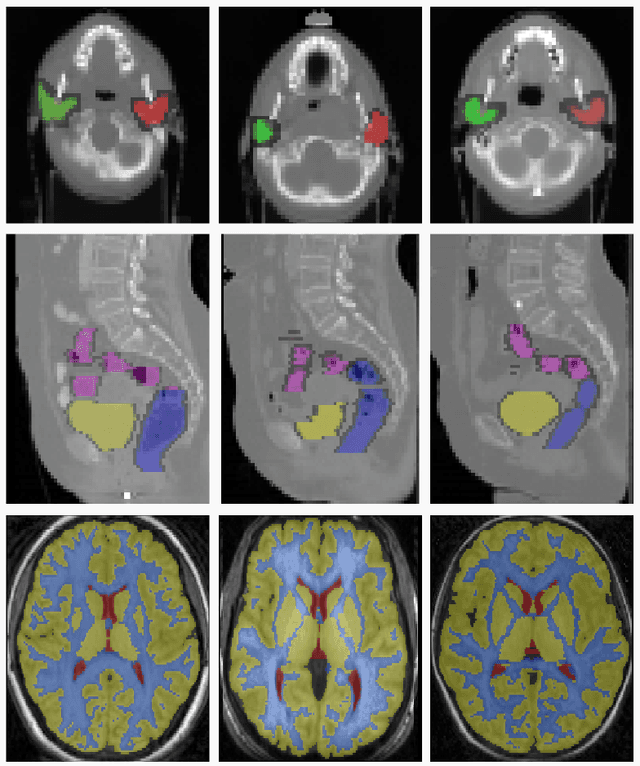
Abstract:We propose a novel weakly supervised discriminative algorithm for learning context specific registration metrics as a linear combination of conventional similarity measures. Conventional metrics have been extensively used over the past two decades and therefore both their strengths and limitations are known. The challenge is to find the optimal relative weighting (or parameters) of different metrics forming the similarity measure of the registration algorithm. Hand-tuning these parameters would result in sub optimal solutions and quickly become infeasible as the number of metrics increases. Furthermore, such hand-crafted combination can only happen at global scale (entire volume) and therefore will not be able to account for the different tissue properties. We propose a learning algorithm for estimating these parameters locally, conditioned to the data semantic classes. The objective function of our formulation is a special case of non-convex function, difference of convex function, which we optimize using the concave convex procedure. As a proof of concept, we show the impact of our approach on three challenging datasets for different anatomical structures and modalities.
EnzyNet: enzyme classification using 3D convolutional neural networks on spatial representation
Jul 19, 2017



Abstract:During the past decade, with the significant progress of computational power as well as ever-rising data availability, deep learning techniques became increasingly popular due to their excellent performance on computer vision problems. The size of the Protein Data Bank has increased more than 15 fold since 1999, which enabled the expansion of models that aim at predicting enzymatic function via their amino acid composition. Amino acid sequence however is less conserved in nature than protein structure and therefore considered a less reliable predictor of protein function. This paper presents EnzyNet, a novel 3D-convolutional neural networks classifier that predicts the Enzyme Commission number of enzymes based only on their voxel-based spatial structure. The spatial distribution of biochemical properties was also examined as complementary information. The 2-layer architecture was investigated on a large dataset of 63,558 enzymes from the Protein Data Bank and achieved an accuracy of 78.4% by exploiting only the binary representation of the protein shape. Code and datasets are available at https://github.com/shervinea/enzynet.
Slice-to-volume medical image registration: a survey
Apr 27, 2017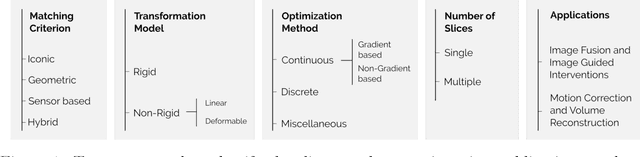
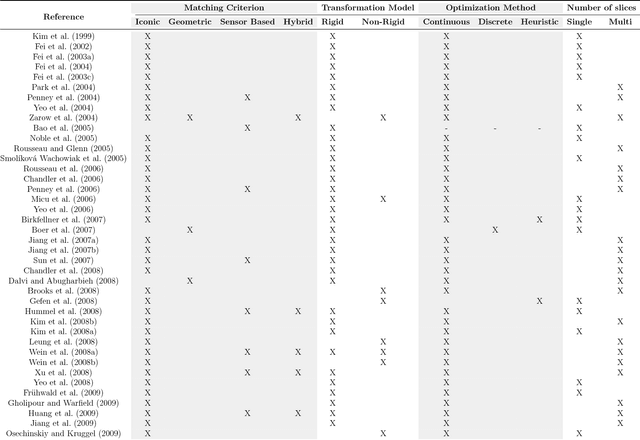
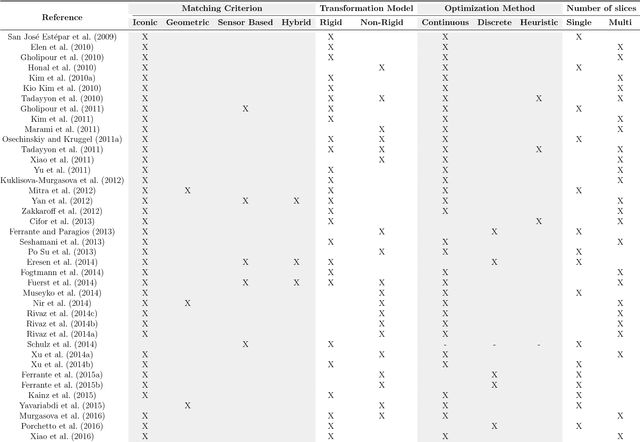
Abstract:During the last decades, the research community of medical imaging has witnessed continuous advances in image registration methods, which pushed the limits of the state-of-the-art and enabled the development of novel medical procedures. A particular type of image registration problem, known as slice-to-volume registration, played a fundamental role in areas like image guided surgeries and volumetric image reconstruction. However, to date, and despite the extensive literature available on this topic, no survey has been written to discuss this challenging problem. This paper introduces the first comprehensive survey of the literature about slice-to-volume registration, presenting a categorical study of the algorithms according to an ad-hoc taxonomy and analyzing advantages and disadvantages of every category. We draw some general conclusions from this analysis and present our perspectives on the future of the field.
Rigid Slice-To-Volume Medical Image Registration through Markov Random Fields
Aug 19, 2016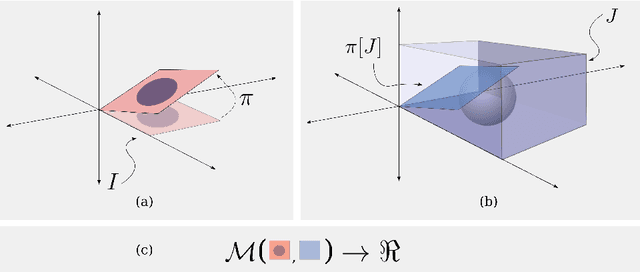
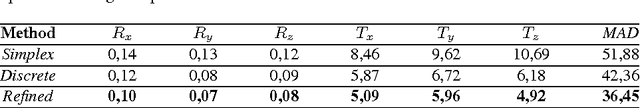
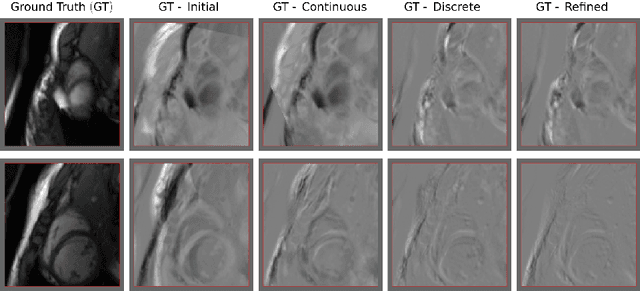
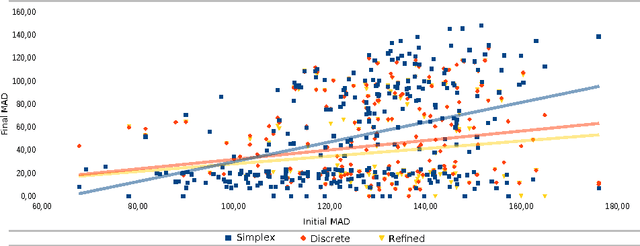
Abstract:Rigid slice-to-volume registration is a challenging task, which finds application in medical imaging problems like image fusion for image guided surgeries and motion correction for volume reconstruction. It is usually formulated as an optimization problem and solved using standard continuous methods. In this paper, we discuss how this task be formulated as a discrete labeling problem on a graph. Inspired by previous works on discrete estimation of linear transformations using Markov Random Fields (MRFs), we model it using a pairwise MRF, where the nodes are associated to the rigid parameters, and the edges encode the relation between the variables. We compare the performance of the proposed method to a continuous formulation optimized using simplex, and we discuss how it can be used to further improve the accuracy of our approach. Promising results are obtained using a monomodal dataset composed of magnetic resonance images (MRI) of a beating heart.
Prior-based Coregistration and Cosegmentation
Jul 22, 2016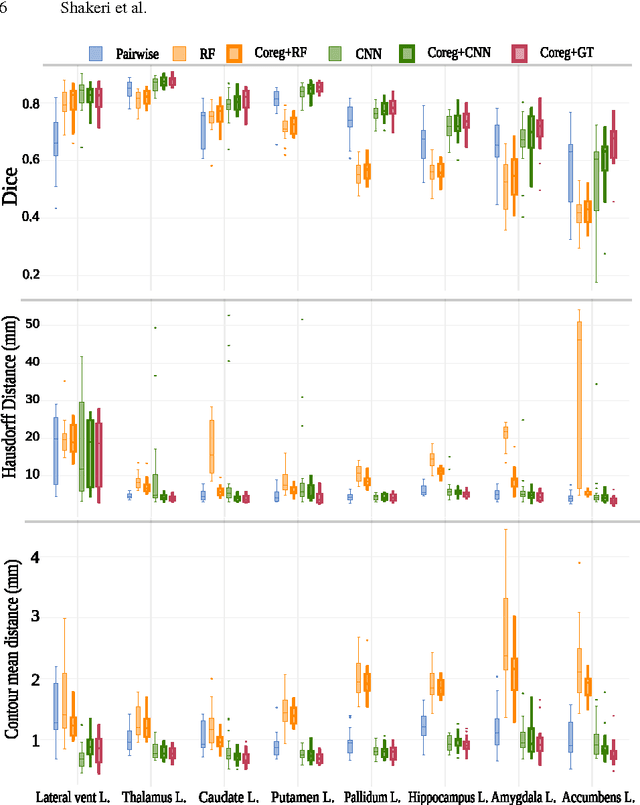
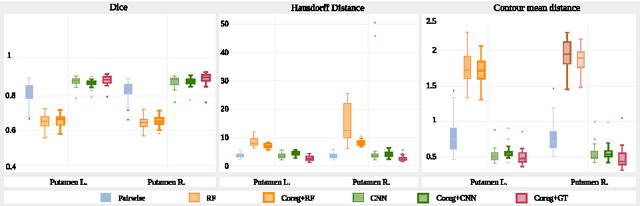
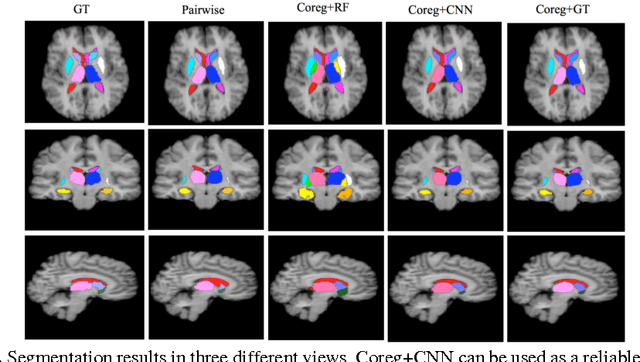
Abstract:We propose a modular and scalable framework for dense coregistration and cosegmentation with two key characteristics: first, we substitute ground truth data with the semantic map output of a classifier; second, we combine this output with population deformable registration to improve both alignment and segmentation. Our approach deforms all volumes towards consensus, taking into account image similarities and label consistency. Our pipeline can incorporate any classifier and similarity metric. Results on two datasets, containing annotations of challenging brain structures, demonstrate the potential of our method.
* The first two authors contributed equally
Sub-cortical brain structure segmentation using F-CNN's
Feb 05, 2016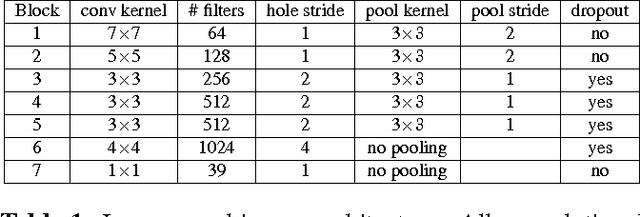
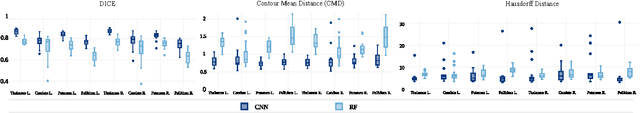
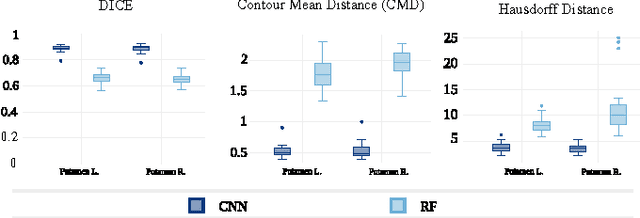
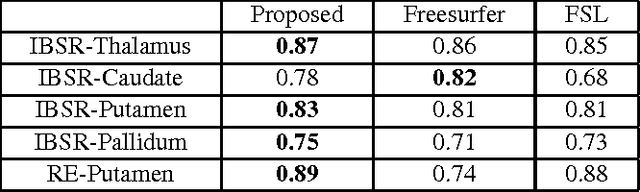
Abstract:In this paper we propose a deep learning approach for segmenting sub-cortical structures of the human brain in Magnetic Resonance (MR) image data. We draw inspiration from a state-of-the-art Fully-Convolutional Neural Network (F-CNN) architecture for semantic segmentation of objects in natural images, and adapt it to our task. Unlike previous CNN-based methods that operate on image patches, our model is applied on a full blown 2D image, without any alignment or registration steps at testing time. We further improve segmentation results by interpreting the CNN output as potentials of a Markov Random Field (MRF), whose topology corresponds to a volumetric grid. Alpha-expansion is used to perform approximate inference imposing spatial volumetric homogeneity to the CNN priors. We compare the performance of the proposed pipeline with a similar system using Random Forest-based priors, as well as state-of-art segmentation algorithms, and show promising results on two different brain MRI datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge