Murat Bilgel
from the iSTAGING consortium, for the ADNI
Unsupervised learning of acquisition variability in structural connectomes via hybrid latent space modeling
May 13, 2026Abstract:Acquisition differences across sites, scanners, and protocols in dMRI introduce variability that complicates structural connectome analysis. This motivates deep learning models that can represent high-dimensional connectomes in a low-dimensional space while explicitly separating acquisition-related effects from biological variation. Conventional dimensionality reduction methods model all variance as continuous, so acquisition effects often get absorbed into a continuous latent space. Recent hybrid latent-space models combine discrete and continuous components to address this, but typically require manual capacity tuning to ensure the discrete component captures the intended variability. We introduce an unsupervised framework that removes this manual tuning by architecturally annealing encoder outputs before decoding, allowing the model to adaptively balance discrete and continuous latent variables during training. To evaluate it, we curated a dataset of N=7,416 structural connectomes derived from dMRI, spanning ages 2 to 102 and 13 studies with 25 unique acquisition-parameter combinations. Of these, 5,900 are cognitively unimpaired, 877 have mild cognitive impairment (MCI), and 639 have Alzheimer's disease (AD). We compare against a standard VAE, PCA with k-means clustering, and hybrid models that anneal only through the loss function. Our architectural annealing produces stronger site learning (ARI=0.53, p<0.05) than these baselines. Results show that a hybrid continuous-discrete latent space, with architectural rather than loss-based annealing, provides a useful unsupervised mechanism for capturing acquisition variability in dMRI: by jointly modeling smooth and categorical structure, the Joint-VAE recovers clusters aligned with scanner and protocol differences.
Harmonizing MR Images Across 100+ Scanners: Multi-site Validation with Traveling Subjects and Real-world Protocols
Apr 21, 2026Abstract:Reliable harmonization of heterogeneous magnetic resonance~(MR) image datasets, especially those acquired in pragmatic clinical trials, is critical to advance multi-center neuroimaging studies and translational machine learning in healthcare. We present an enhanced and rigorously validated version of the HACA3 harmonization algorithm, which we refer to as HACA3$^+$, incorporating key methodological enhancements: (1)~an improved artifact encoder to better isolate and mitigate image artifacts, (2)~background and foreground-sensitive attention mechanisms to increase harmonization specificity, and (3)~extensive training using data spanning 100+ scanners from 64 independent sites, providing a broader diversity of scanners than other harmonization methods. Our study focuses on four commonly acquired MR image contrasts (T1-weighted, T2-weighted, proton density, \& fluid-attenuated inversion recovery), reflecting realistic clinical protocols. We perform inter-site harmonization experiments using traveling subjects to assess the generalization and robustness of the harmonization model. We compare the results of the publicly available version of HACA3 and our implementation, HACA3$^+$. Downstream relevance is further established through whole brain segmentation and image imputation. Finally, we justify each enhancement through an ablation experiment. Pre-trained weights and code for HACA3$^+$ are made publicly available at https://github.com/shays15/haca3-plus.
HACA3: A Unified Approach for Multi-site MR Image Harmonization
Dec 12, 2022



Abstract:The lack of standardization is a prominent issue in magnetic resonance (MR) imaging. This often causes undesired contrast variations due to differences in hardware and acquisition parameters. In recent years, MR harmonization using image synthesis with disentanglement has been proposed to compensate for the undesired contrast variations. Despite the success of existing methods, we argue that three major improvements can be made. First, most existing methods are built upon the assumption that multi-contrast MR images of the same subject share the same anatomy. This assumption is questionable since different MR contrasts are specialized to highlight different anatomical features. Second, these methods often require a fixed set of MR contrasts for training (e.g., both Tw-weighted and T2-weighted images must be available), which limits their applicability. Third, existing methods generally are sensitive to imaging artifacts. In this paper, we present a novel approach, Harmonization with Attention-based Contrast, Anatomy, and Artifact Awareness (HACA3), to address these three issues. We first propose an anatomy fusion module that enables HACA3 to respect the anatomical differences between MR contrasts. HACA3 is also robust to imaging artifacts and can be trained and applied to any set of MR contrasts. Experiments show that HACA3 achieves state-of-the-art performance under multiple image quality metrics. We also demonstrate the applicability of HACA3 on downstream tasks with diverse MR datasets acquired from 21 sites with different field strengths, scanner platforms, and acquisition protocols.
Disentangling A Single MR Modality
May 10, 2022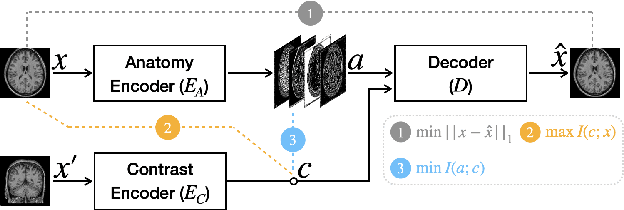
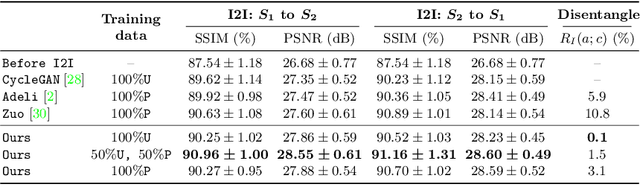
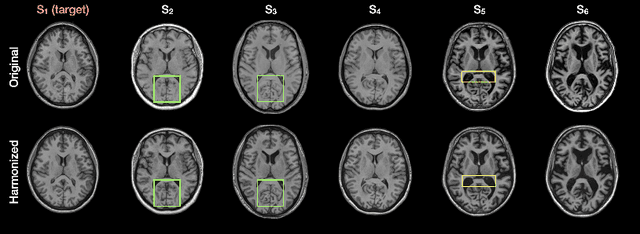
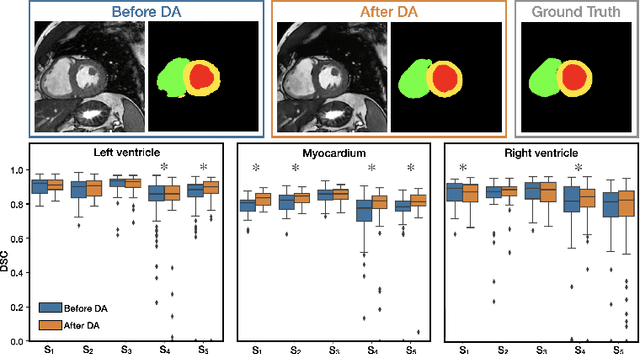
Abstract:Disentangling anatomical and contrast information from medical images has gained attention recently, demonstrating benefits for various image analysis tasks. Current methods learn disentangled representations using either paired multi-modal images with the same underlying anatomy or auxiliary labels (e.g., manual delineations) to provide inductive bias for disentanglement. However, these requirements could significantly increase the time and cost in data collection and limit the applicability of these methods when such data are not available. Moreover, these methods generally do not guarantee disentanglement. In this paper, we present a novel framework that learns theoretically and practically superior disentanglement from single modality magnetic resonance images. Moreover, we propose a new information-based metric to quantitatively evaluate disentanglement. Comparisons over existing disentangling methods demonstrate that the proposed method achieves superior performance in both disentanglement and cross-domain image-to-image translation tasks.
Disentangling Alzheimer's disease neurodegeneration from typical brain aging using machine learning
Sep 08, 2021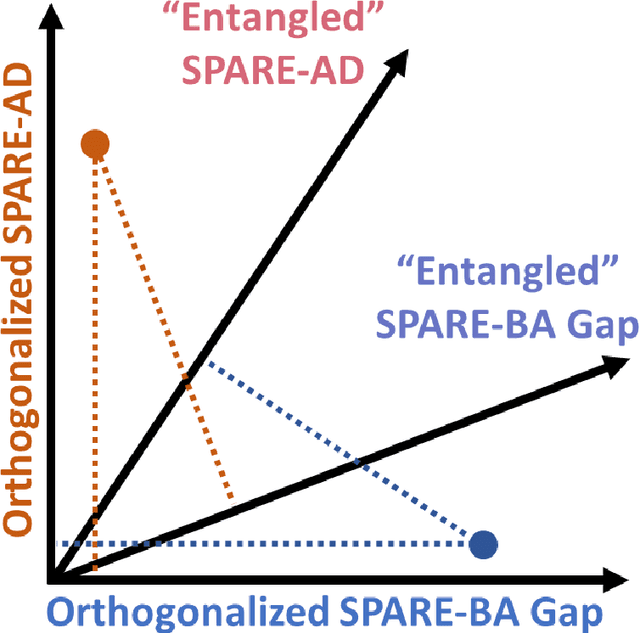
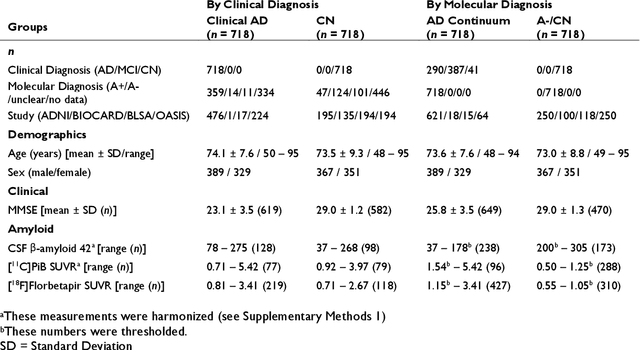
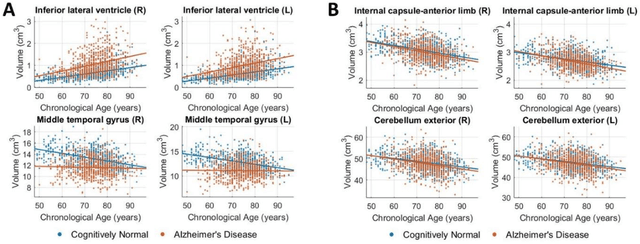
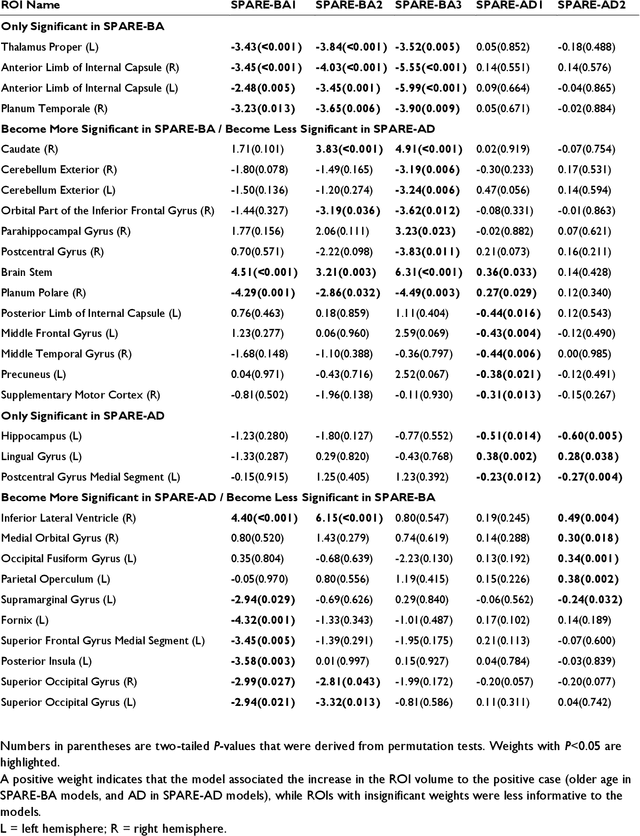
Abstract:Neuroimaging biomarkers that distinguish between typical brain aging and Alzheimer's disease (AD) are valuable for determining how much each contributes to cognitive decline. Machine learning models can derive multi-variate brain change patterns related to the two processes, including the SPARE-AD (Spatial Patterns of Atrophy for Recognition of Alzheimer's Disease) and SPARE-BA (of Brain Aging) investigated herein. However, substantial overlap between brain regions affected in the two processes confounds measuring them independently. We present a methodology toward disentangling the two. T1-weighted MRI images of 4,054 participants (48-95 years) with AD, mild cognitive impairment (MCI), or cognitively normal (CN) diagnoses from the iSTAGING (Imaging-based coordinate SysTem for AGIng and NeurodeGenerative diseases) consortium were analyzed. First, a subset of AD patients and CN adults were selected based purely on clinical diagnoses to train SPARE-BA1 (regression of age using CN individuals) and SPARE-AD1 (classification of CN versus AD). Second, analogous groups were selected based on clinical and molecular markers to train SPARE-BA2 and SPARE-AD2: amyloid-positive (A+) AD continuum group (consisting of A+AD, A+MCI, and A+ and tau-positive CN individuals) and amyloid-negative (A-) CN group. Finally, the combined group of the AD continuum and A-/CN individuals was used to train SPARE-BA3, with the intention to estimate brain age regardless of AD-related brain changes. Disentangled SPARE models derived brain patterns that were more specific to the two types of the brain changes. Correlation between the SPARE-BA and SPARE-AD was significantly reduced. Correlation of disentangled SPARE-AD was non-inferior to the molecular measurements and to the number of APOE4 alleles, but was less to AD-related psychometric test scores, suggesting contribution of advanced brain aging to these scores.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge