Muhammad E. H. Chowdhury
Advanced Artificial Intelligence Algorithms in Cochlear Implants: Review of Healthcare Strategies, Challenges, and Perspectives
Mar 17, 2024



Abstract:Automatic speech recognition (ASR) plays a pivotal role in our daily lives, offering utility not only for interacting with machines but also for facilitating communication for individuals with either partial or profound hearing impairments. The process involves receiving the speech signal in analogue form, followed by various signal processing algorithms to make it compatible with devices of limited capacity, such as cochlear implants (CIs). Unfortunately, these implants, equipped with a finite number of electrodes, often result in speech distortion during synthesis. Despite efforts by researchers to enhance received speech quality using various state-of-the-art signal processing techniques, challenges persist, especially in scenarios involving multiple sources of speech, environmental noise, and other circumstances. The advent of new artificial intelligence (AI) methods has ushered in cutting-edge strategies to address the limitations and difficulties associated with traditional signal processing techniques dedicated to CIs. This review aims to comprehensively review advancements in CI-based ASR and speech enhancement, among other related aspects. The primary objective is to provide a thorough overview of metrics and datasets, exploring the capabilities of AI algorithms in this biomedical field, summarizing and commenting on the best results obtained. Additionally, the review will delve into potential applications and suggest future directions to bridge existing research gaps in this domain.
Deep Learning in Computed Tomography Pulmonary Angiography Imaging: A Dual-Pronged Approach for Pulmonary Embolism Detection
Nov 09, 2023



Abstract:Pulmonary Embolism (PE) is a critical medical condition characterized by obstructions in the pulmonary arteries. Despite being a major health concern, it often goes underdiagnosed leading to detrimental clinical outcomes. The increasing reliance on Computed Tomography Pulmonary Angiography for diagnosis presents challenges and a pressing need for enhanced diagnostic solutions. The primary objective of this study is to leverage deep learning techniques to enhance the Computer Assisted Diagnosis of PE. This study presents a comprehensive dual-pronged approach combining classification and detection for PE diagnosis. We introduce an Attention-Guided Convolutional Neural Network (AG-CNN) for classification, addressing both global and local lesion region. For detection, state-of-the-art models are employed to pinpoint potential PE regions. Different ensembling techniques further improve detection accuracy by combining predictions from different models. Finally, a heuristic strategy integrates classifier outputs with detection results, ensuring robust and accurate PE identification. Our attention-guided classification approach, tested on the Ferdowsi University of Mashhad's Pulmonary Embolism (FUMPE) dataset, outperformed the baseline model DenseNet-121 by achieving an 8.1% increase in the Area Under the Receiver Operating Characteristic. By employing ensemble techniques with detection models, the mean average precision (mAP) was considerably enhanced by a 4.7% increase. The classifier-guided framework further refined the mAP and F1 scores over the ensemble models. Our research offers a comprehensive approach to PE diagnostics using deep learning, addressing the prevalent issues of underdiagnosis and misdiagnosis. We aim to improve PE patient care by integrating AI solutions into clinical workflows, highlighting the potential of human-AI collaboration in medical diagnostics.
Deep Learning based Automatic Quantification of Urethral Plate Quality using the Plate Objective Scoring Tool (POST)
Sep 28, 2022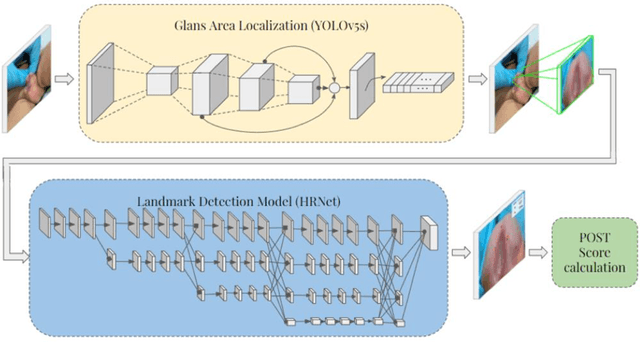
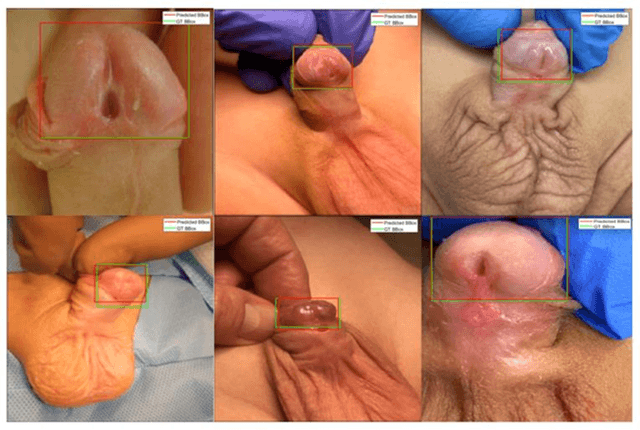
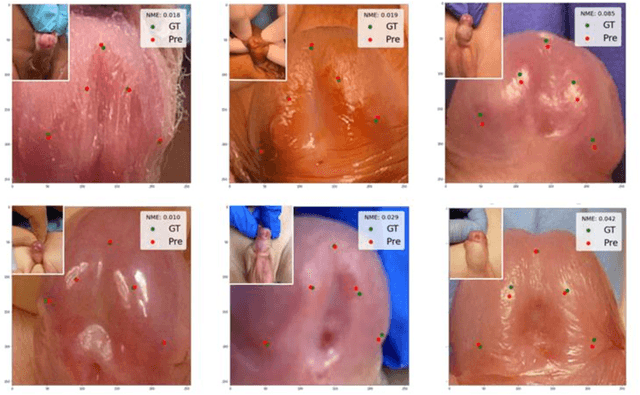
Abstract:Objectives: To explore the capacity of deep learning algorithm to further streamline and optimize urethral plate (UP) quality appraisal on 2D images using the plate objective scoring tool (POST), aiming to increase the objectivity and reproducibility of UP appraisal in hypospadias repair. Methods: The five key POST landmarks were marked by specialists in a 691-image dataset of prepubertal boys undergoing primary hypospadias repair. This dataset was then used to develop and validate a deep learning-based landmark detection model. The proposed framework begins with glans localization and detection, where the input image is cropped using the predicted bounding box. Next, a deep convolutional neural network (CNN) architecture is used to predict the coordinates of the five POST landmarks. These predicted landmarks are then used to assess UP quality in distal hypospadias. Results: The proposed model accurately localized the glans area, with a mean average precision (mAP) of 99.5% and an overall sensitivity of 99.1%. A normalized mean error (NME) of 0.07152 was achieved in predicting the coordinates of the landmarks, with a mean squared error (MSE) of 0.001 and a 20.2% failure rate at a threshold of 0.1 NME. Conclusions: This deep learning application shows robustness and high precision in using POST to appraise UP quality. Further assessment using international multi-centre image-based databases is ongoing. External validation could benefit deep learning algorithms and lead to better assessments, decision-making and predictions for surgical outcomes.
BIO-CXRNET: A Robust Multimodal Stacking Machine Learning Technique for Mortality Risk Prediction of COVID-19 Patients using Chest X-Ray Images and Clinical Data
Jun 15, 2022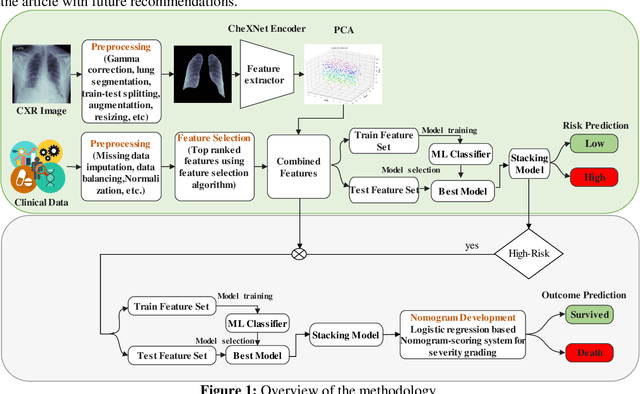
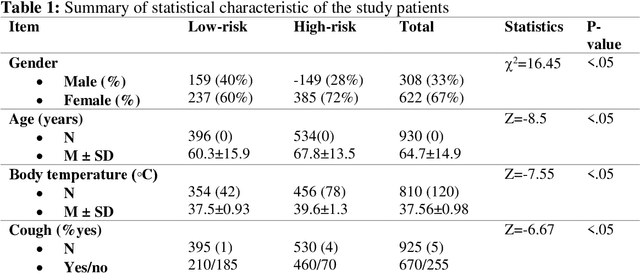
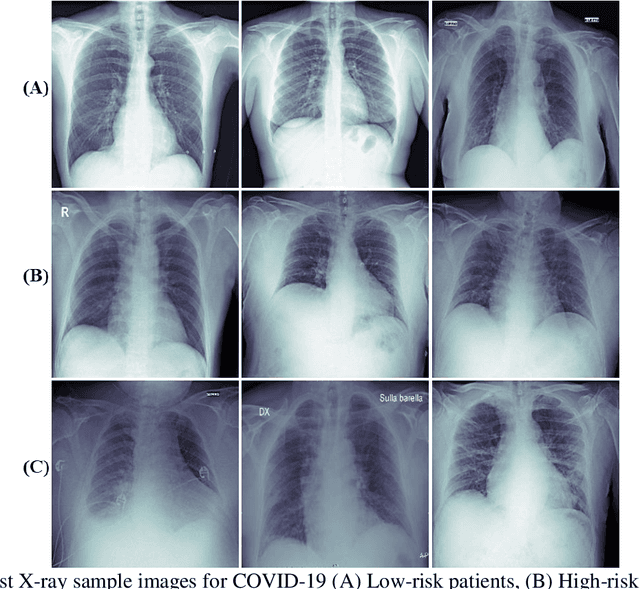
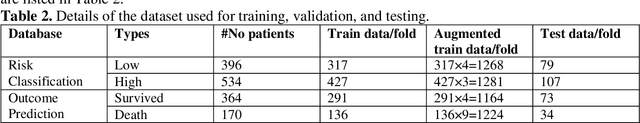
Abstract:Fast and accurate detection of the disease can significantly help in reducing the strain on the healthcare facility of any country to reduce the mortality during any pandemic. The goal of this work is to create a multimodal system using a novel machine learning framework that uses both Chest X-ray (CXR) images and clinical data to predict severity in COVID-19 patients. In addition, the study presents a nomogram-based scoring technique for predicting the likelihood of death in high-risk patients. This study uses 25 biomarkers and CXR images in predicting the risk in 930 COVID-19 patients admitted during the first wave of COVID-19 (March-June 2020) in Italy. The proposed multimodal stacking technique produced the precision, sensitivity, and F1-score, of 89.03%, 90.44%, and 89.03%, respectively to identify low or high-risk patients. This multimodal approach improved the accuracy by 6% in comparison to the CXR image or clinical data alone. Finally, nomogram scoring system using multivariate logistic regression -- was used to stratify the mortality risk among the high-risk patients identified in the first stage. Lactate Dehydrogenase (LDH), O2 percentage, White Blood Cells (WBC) Count, Age, and C-reactive protein (CRP) were identified as useful predictor using random forest feature selection model. Five predictors parameters and a CXR image based nomogram score was developed for quantifying the probability of death and categorizing them into two risk groups: survived (<50%), and death (>=50%), respectively. The multi-modal technique was able to predict the death probability of high-risk patients with an F1 score of 92.88 %. The area under the curves for the development and validation cohorts are 0.981 and 0.939, respectively.
Motion Artifacts Correction from Single-Channel EEG and fNIRS Signals using Novel Wavelet Packet Decomposition in Combination with Canonical Correlation Analysis
Apr 09, 2022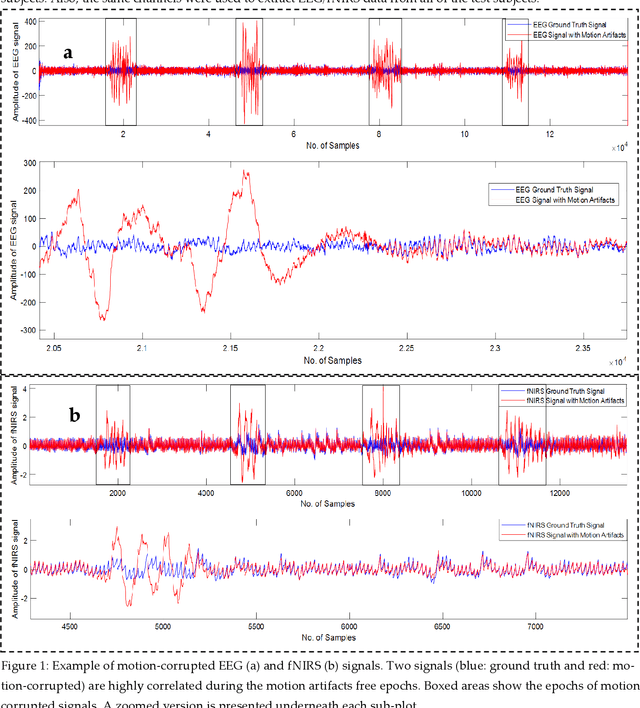
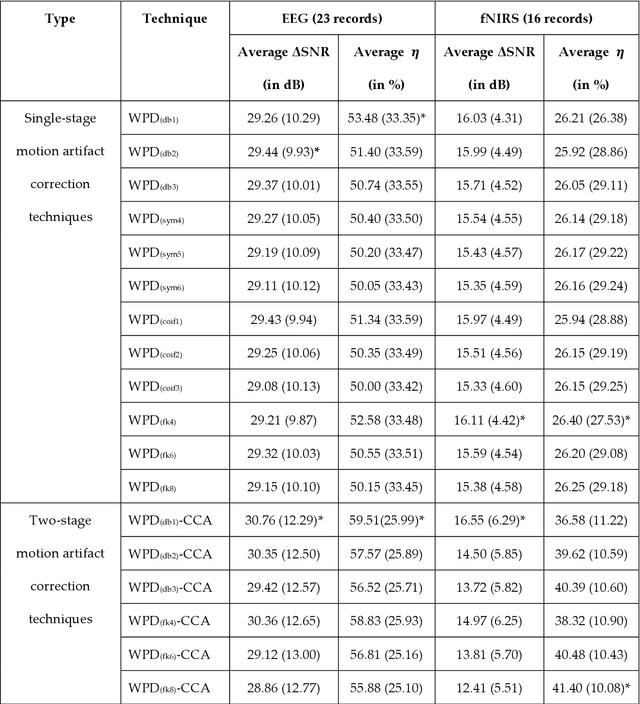
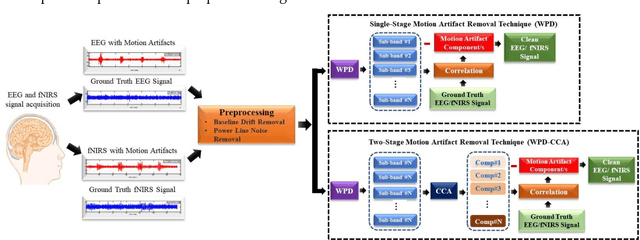
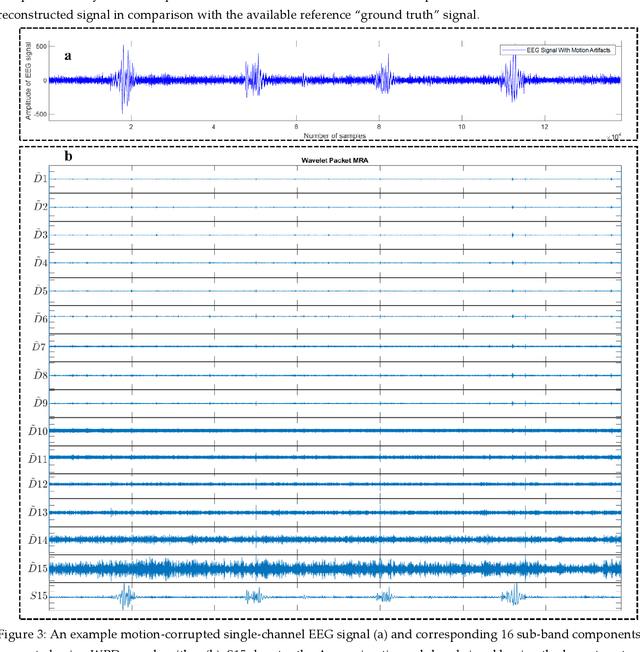
Abstract:The electroencephalogram (EEG) and functional near-infrared spectroscopy (fNIRS) signals, highly non-stationary in nature, greatly suffers from motion artifacts while recorded using wearable sensors. This paper proposes two robust methods: i) Wavelet packet decomposition (WPD), and ii) WPD in combination with canonical correlation analysis (WPD-CCA), for motion artifact correction from single-channel EEG and fNIRS signals. The efficacy of these proposed techniques is tested using a benchmark dataset and the performance of the proposed methods is measured using two well-established performance matrices: i) Difference in the signal to noise ratio ({\Delta}SNR) and ii) Percentage reduction in motion artifacts ({\eta}). The proposed WPD-based single-stage motion artifacts correction technique produces the highest average {\Delta}SNR (29.44 dB) when db2 wavelet packet is incorporated whereas the greatest average {\eta} (53.48%) is obtained using db1 wavelet packet for all the available 23 EEG recordings. Our proposed two-stage motion artifacts correction technique i.e. the WPD-CCA method utilizing db1 wavelet packet has shown the best denoising performance producing an average {\Delta}SNR and {\eta} values of 30.76 dB and 59.51%, respectively for all the EEG recordings. On the other hand, the two-stage motion artifacts removal technique i.e. WPD-CCA has produced the best average {\Delta}SNR (16.55 dB, utilizing db1 wavelet packet) and largest average {\eta} (41.40%, using fk8 wavelet packet). The highest average {\Delta}SNR and {\eta} using single-stage artifacts removal techniques (WPD) are found as 16.11 dB and 26.40%, respectively for all the fNIRS signals using fk4 wavelet packet. In both EEG and fNIRS modalities, the percentage reduction in motion artifacts increases by 11.28% and 56.82%, respectively when two-stage WPD-CCA techniques are employed.
A machine learning-based severity prediction tool for diabetic sensorimotor polyneuropathy using Michigan neuropathy screening instrumentations
Mar 28, 2022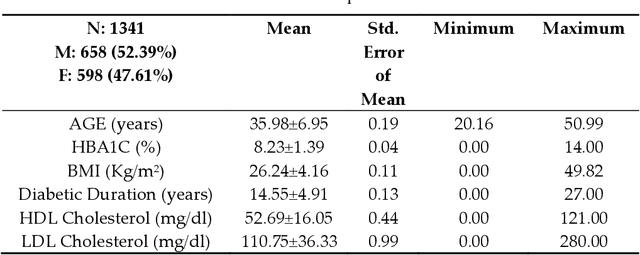
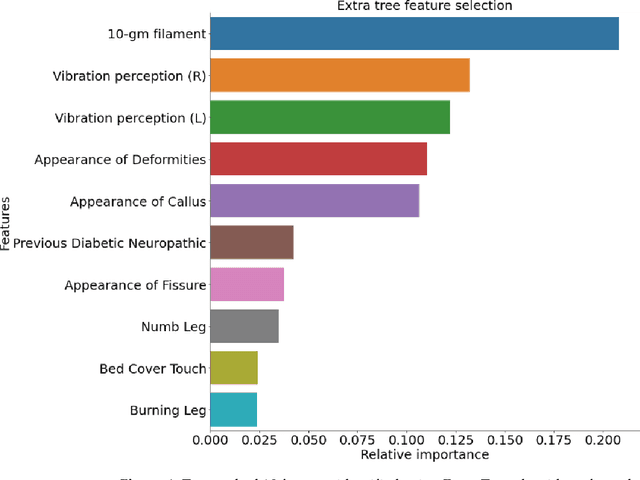
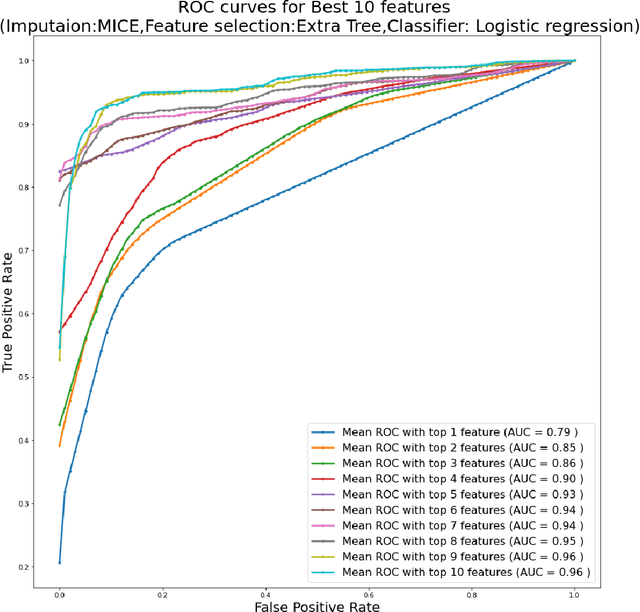
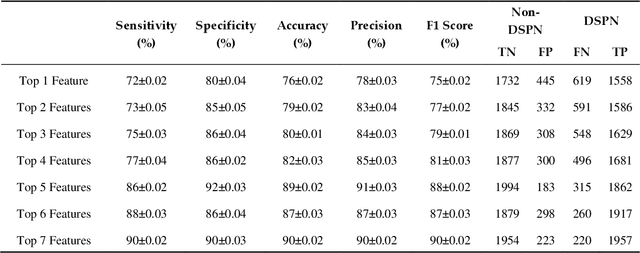
Abstract:Background: Diabetic Sensorimotor polyneuropathy (DSPN) is a major long-term complication in diabetic patients associated with painful neuropathy, foot ulceration and amputation. The Michigan neuropathy screening instrument (MNSI) is one of the most common screening techniques for DSPN, however, it does not provide any direct severity grading system. Method: For designing and modelling the DSPN severity grading systems for MNSI, 19 years of data from Epidemiology of Diabetes Interventions and Complications (EDIC) clinical trials were used. MNSI variables and patient outcomes were investigated using machine learning tools to identify the features having higher association in DSPN identification. A multivariable logistic regression-based nomogram was generated and validated for DSPN severity grading. Results: The top-7 ranked features from MNSI: 10-gm filament, Vibration perception (R), Vibration perception (L), previous diabetic neuropathy, the appearance of deformities, appearance of callus and appearance of fissure were identified as key features for identifying DSPN using the extra tree model. The area under the curve (AUC) of the nomogram for the internal and external datasets were 0.9421 and 0.946, respectively. From the developed nomogram, the probability of having DSPN was predicted and a DSPN severity scoring system for MNSI was developed from the probability score. The model performance was validated on an independent dataset. Patients were stratified into four severity levels: absent, mild, moderate, and severe using a cut-off value of 10.5, 12.7 and 15 for a DSPN probability less than 50%, 75% to 90%, and above 90%, respectively. Conclusions: This study provides a simple, easy-to-use and reliable algorithm for defining the prognosis and management of patients with DSPN.
OSegNet: Operational Segmentation Network for COVID-19 Detection using Chest X-ray Images
Feb 21, 2022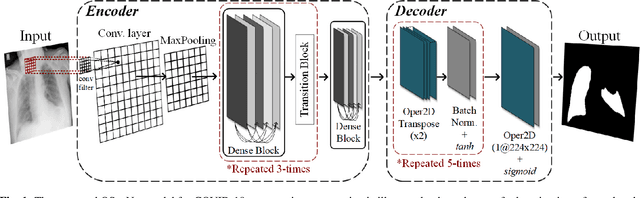
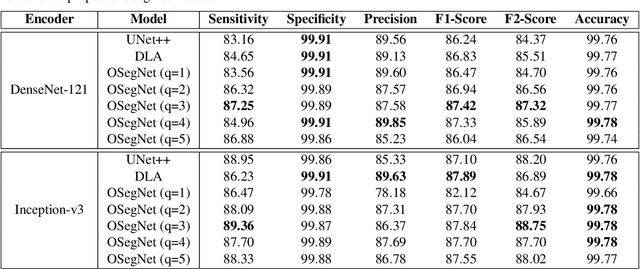
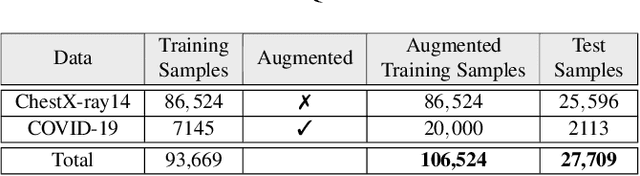
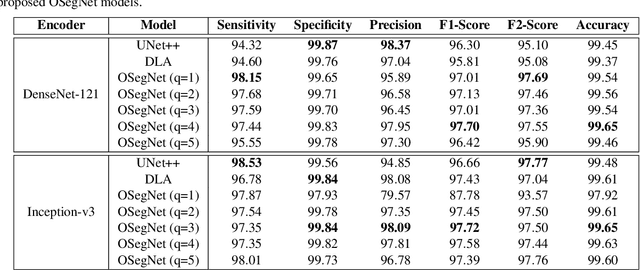
Abstract:Coronavirus disease 2019 (COVID-19) has been diagnosed automatically using Machine Learning algorithms over chest X-ray (CXR) images. However, most of the earlier studies used Deep Learning models over scarce datasets bearing the risk of overfitting. Additionally, previous studies have revealed the fact that deep networks are not reliable for classification since their decisions may originate from irrelevant areas on the CXRs. Therefore, in this study, we propose Operational Segmentation Network (OSegNet) that performs detection by segmenting COVID-19 pneumonia for a reliable diagnosis. To address the data scarcity encountered in training and especially in evaluation, this study extends the largest COVID-19 CXR dataset: QaTa-COV19 with 121,378 CXRs including 9258 COVID-19 samples with their corresponding ground-truth segmentation masks that are publicly shared with the research community. Consequently, OSegNet has achieved a detection performance with the highest accuracy of 99.65% among the state-of-the-art deep models with 98.09% precision.
RamanNet: A generalized neural network architecture for Raman Spectrum Analysis
Jan 20, 2022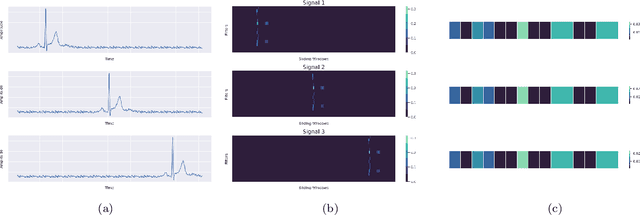
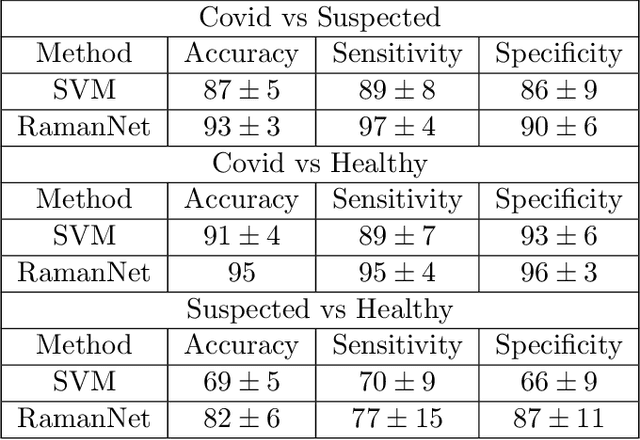
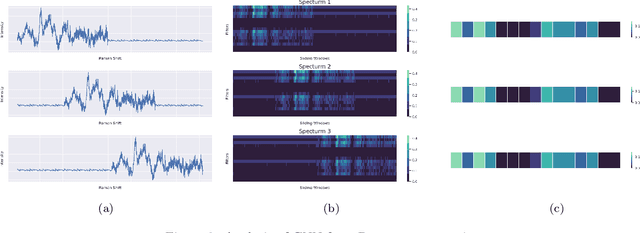
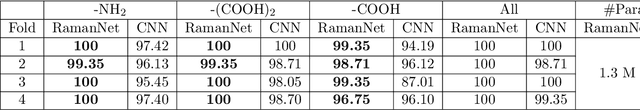
Abstract:Raman spectroscopy provides a vibrational profile of the molecules and thus can be used to uniquely identify different kind of materials. This sort of fingerprinting molecules has thus led to widespread application of Raman spectrum in various fields like medical dignostics, forensics, mineralogy, bacteriology and virology etc. Despite the recent rise in Raman spectra data volume, there has not been any significant effort in developing generalized machine learning methods for Raman spectra analysis. We examine, experiment and evaluate existing methods and conjecture that neither current sequential models nor traditional machine learning models are satisfactorily sufficient to analyze Raman spectra. Both has their perks and pitfalls, therefore we attempt to mix the best of both worlds and propose a novel network architecture RamanNet. RamanNet is immune to invariance property in CNN and at the same time better than traditional machine learning models for the inclusion of sparse connectivity. Our experiments on 4 public datasets demonstrate superior performance over the much complex state-of-the-art methods and thus RamanNet has the potential to become the defacto standard in Raman spectra data analysis
A Shallow U-Net Architecture for Reliably Predicting Blood Pressure from Photoplethysmogram and Electrocardiogram Signals
Nov 12, 2021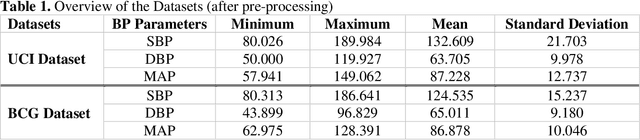
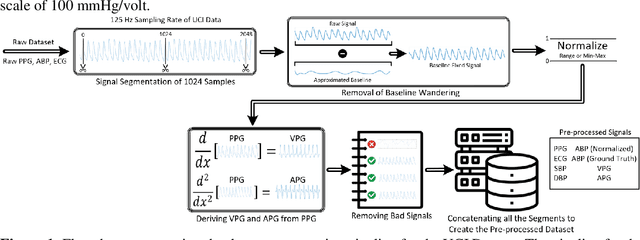
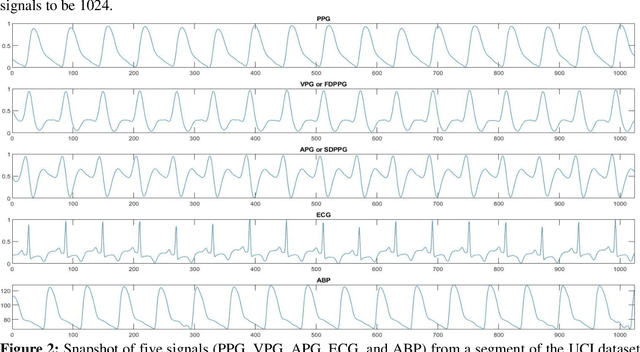
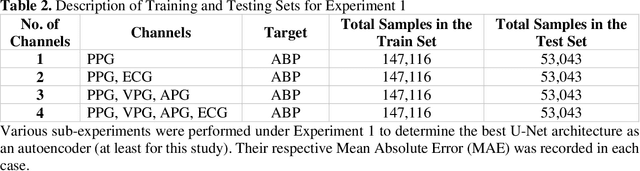
Abstract:Cardiovascular diseases are the most common causes of death around the world. To detect and treat heart-related diseases, continuous Blood Pressure (BP) monitoring along with many other parameters are required. Several invasive and non-invasive methods have been developed for this purpose. Most existing methods used in the hospitals for continuous monitoring of BP are invasive. On the contrary, cuff-based BP monitoring methods, which can predict Systolic Blood Pressure (SBP) and Diastolic Blood Pressure (DBP), cannot be used for continuous monitoring. Several studies attempted to predict BP from non-invasively collectible signals such as Photoplethysmogram (PPG) and Electrocardiogram (ECG), which can be used for continuous monitoring. In this study, we explored the applicability of autoencoders in predicting BP from PPG and ECG signals. The investigation was carried out on 12,000 instances of 942 patients of the MIMIC-II dataset and it was found that a very shallow, one-dimensional autoencoder can extract the relevant features to predict the SBP and DBP with the state-of-the-art performance on a very large dataset. Independent test set from a portion of the MIMIC-II dataset provides an MAE of 2.333 and 0.713 for SBP and DBP, respectively. On an external dataset of forty subjects, the model trained on the MIMIC-II dataset, provides an MAE of 2.728 and 1.166 for SBP and DBP, respectively. For both the cases, the results met British Hypertension Society (BHS) Grade A and surpassed the studies from the current literature.
A Machine Learning Model for Early Detection of Diabetic Foot using Thermogram Images
Jun 27, 2021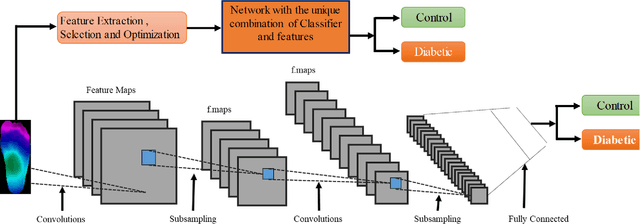
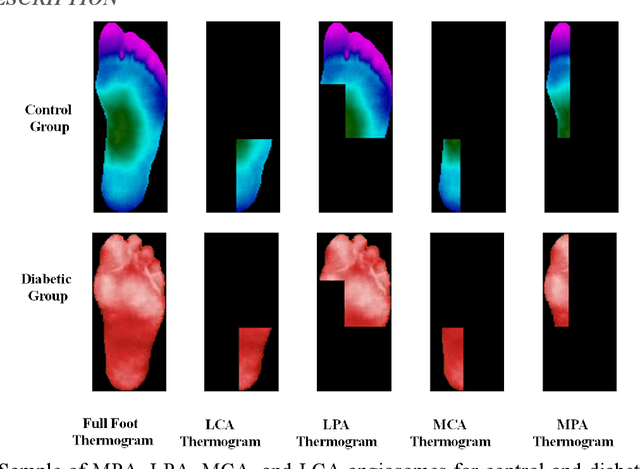
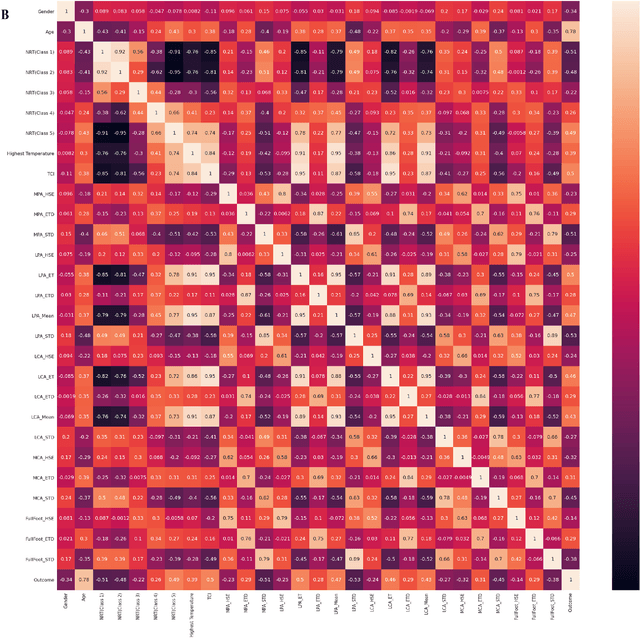
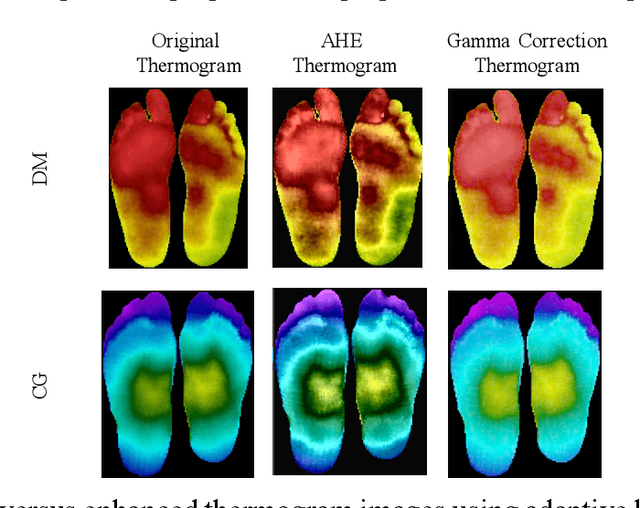
Abstract:Diabetes foot ulceration (DFU) and amputation are a cause of significant morbidity. The prevention of DFU may be achieved by the identification of patients at risk of DFU and the institution of preventative measures through education and offloading. Several studies have reported that thermogram images may help to detect an increase in plantar temperature prior to DFU. However, the distribution of plantar temperature may be heterogeneous, making it difficult to quantify and utilize to predict outcomes. We have compared a machine learning-based scoring technique with feature selection and optimization techniques and learning classifiers to several state-of-the-art Convolutional Neural Networks (CNNs) on foot thermogram images and propose a robust solution to identify the diabetic foot. A comparatively shallow CNN model, MobilenetV2 achieved an F1 score of ~95% for a two-feet thermogram image-based classification and the AdaBoost Classifier used 10 features and achieved an F1 score of 97 %. A comparison of the inference time for the best-performing networks confirmed that the proposed algorithm can be deployed as a smartphone application to allow the user to monitor the progression of the DFU in a home setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge