Moinak Bhattacharya
Gaze2Report: Radiology Report Generation via Visual-Gaze Prompt Tuning of LLMs
Apr 07, 2026Abstract:Existing deep learning methods for radiology report generation enhance diagnostic efficiency but often overlook physician-informed medical priors. This leads to a suboptimal alignment between the structured explanations and disease manifestations. Eye gaze data provides critical insights into a radiologist's visual attention, enhancing the relevance and interpretability of extracted features while aligning with human decision-making processes. However, despite its promising potential, the integration of eye gaze information into AI-driven medical imaging workflows is impeded by challenges such as the complexity of multimodal data fusion and the high cost of gaze acquisition, particularly its absence during inference, limiting its practical applicability in real-world clinical settings. To address these issues, we introduce Gaze2Report, a framework which leverages a scanpath prediction module and Graph Neural Network (GNN) to generate joint visual-gaze tokens. Combined with instruction and report tokens, these form a multimodal prompt used to fine-tune LoRA layers of large language models (LLMs) for autoregressive report generation. Gaze2Report enhances report quality through eye-gaze-guided visual learning and incorporates on-the-fly scanpath prediction, enabling the model to operate without gaze input during inference.
NeuroRAD-FM: A Foundation Model for Neuro-Oncology with Distributionally Robust Training
Sep 18, 2025



Abstract:Neuro-oncology poses unique challenges for machine learning due to heterogeneous data and tumor complexity, limiting the ability of foundation models (FMs) to generalize across cohorts. Existing FMs also perform poorly in predicting uncommon molecular markers, which are essential for treatment response and risk stratification. To address these gaps, we developed a neuro-oncology specific FM with a distributionally robust loss function, enabling accurate estimation of tumor phenotypes while maintaining cross-institution generalization. We pretrained self-supervised backbones (BYOL, DINO, MAE, MoCo) on multi-institutional brain tumor MRI and applied distributionally robust optimization (DRO) to mitigate site and class imbalance. Downstream tasks included molecular classification of common markers (MGMT, IDH1, 1p/19q, EGFR), uncommon alterations (ATRX, TP53, CDKN2A/2B, TERT), continuous markers (Ki-67, TP53), and overall survival prediction in IDH1 wild-type glioblastoma at UCSF, UPenn, and CUIMC. Our method improved molecular prediction and reduced site-specific embedding differences. At CUIMC, mean balanced accuracy rose from 0.744 to 0.785 and AUC from 0.656 to 0.676, with the largest gains for underrepresented endpoints (CDKN2A/2B accuracy 0.86 to 0.92, AUC 0.73 to 0.92; ATRX AUC 0.69 to 0.82; Ki-67 accuracy 0.60 to 0.69). For survival, c-index improved at all sites: CUIMC 0.592 to 0.597, UPenn 0.647 to 0.672, UCSF 0.600 to 0.627. Grad-CAM highlighted tumor and peri-tumoral regions, confirming interpretability. Overall, coupling FMs with DRO yields more site-invariant representations, improves prediction of common and uncommon markers, and enhances survival discrimination, underscoring the need for prospective validation and integration of longitudinal and interventional signals to advance precision neuro-oncology.
ImmunoDiff: A Diffusion Model for Immunotherapy Response Prediction in Lung Cancer
May 29, 2025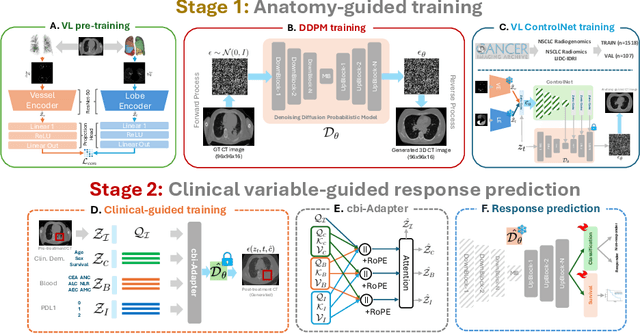

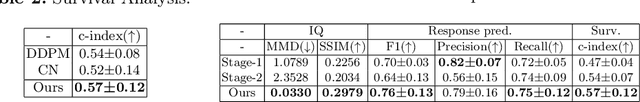

Abstract:Accurately predicting immunotherapy response in Non-Small Cell Lung Cancer (NSCLC) remains a critical unmet need. Existing radiomics and deep learning-based predictive models rely primarily on pre-treatment imaging to predict categorical response outcomes, limiting their ability to capture the complex morphological and textural transformations induced by immunotherapy. This study introduces ImmunoDiff, an anatomy-aware diffusion model designed to synthesize post-treatment CT scans from baseline imaging while incorporating clinically relevant constraints. The proposed framework integrates anatomical priors, specifically lobar and vascular structures, to enhance fidelity in CT synthesis. Additionally, we introduce a novel cbi-Adapter, a conditioning module that ensures pairwise-consistent multimodal integration of imaging and clinical data embeddings, to refine the generative process. Additionally, a clinical variable conditioning mechanism is introduced, leveraging demographic data, blood-based biomarkers, and PD-L1 expression to refine the generative process. Evaluations on an in-house NSCLC cohort treated with immune checkpoint inhibitors demonstrate a 21.24% improvement in balanced accuracy for response prediction and a 0.03 increase in c-index for survival prediction. Code will be released soon.
BrainMRDiff: A Diffusion Model for Anatomically Consistent Brain MRI Synthesis
Apr 06, 2025



Abstract:Accurate brain tumor diagnosis relies on the assessment of multiple Magnetic Resonance Imaging (MRI) sequences. However, in clinical practice, the acquisition of certain sequences may be affected by factors like motion artifacts or contrast agent contraindications, leading to suboptimal outcome, such as poor image quality. This can then affect image interpretation by radiologists. Synthesizing high quality MRI sequences has thus become a critical research focus. Though recent advancements in controllable generative AI have facilitated the synthesis of diagnostic quality MRI, ensuring anatomical accuracy remains a significant challenge. Preserving critical structural relationships between different anatomical regions is essential, as even minor structural or topological inconsistencies can compromise diagnostic validity. In this work, we propose BrainMRDiff, a novel topology-preserving, anatomy-guided diffusion model for synthesizing brain MRI, leveraging brain and tumor anatomies as conditioning inputs. To achieve this, we introduce two key modules: Tumor+Structure Aggregation (TSA) and Topology-Guided Anatomy Preservation (TGAP). TSA integrates diverse anatomical structures with tumor information, forming a comprehensive conditioning mechanism for the diffusion process. TGAP enforces topological consistency during reverse denoising diffusion process; both these modules ensure that the generated image respects anatomical integrity. Experimental results demonstrate that BrainMRDiff surpasses existing baselines, achieving performance improvements of 23.33% on the BraTS-AG dataset and 33.33% on the BraTS-Met dataset. Code will be made publicly available soon.
RadioTransformer: A Cascaded Global-Focal Transformer for Visual Attention-guided Disease Classification
Feb 23, 2022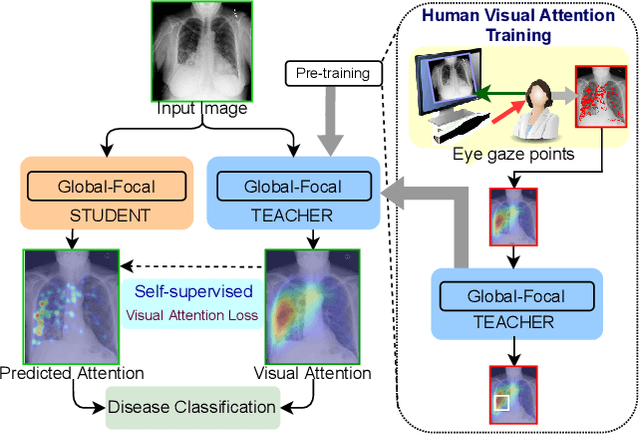
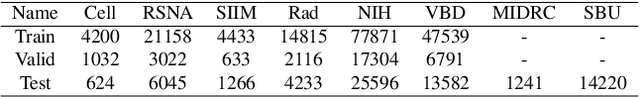
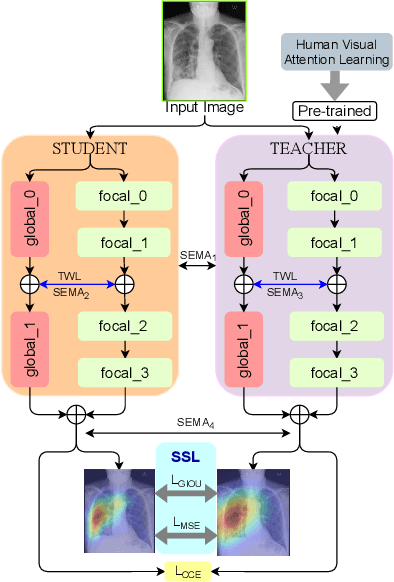
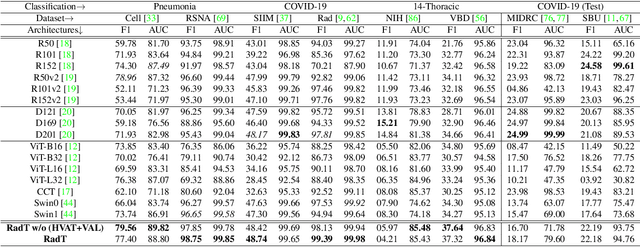
Abstract:In this work, we present RadioTransformer, a novel visual attention-driven transformer framework, that leverages radiologists' gaze patterns and models their visuo-cognitive behavior for disease diagnosis on chest radiographs. Domain experts, such as radiologists, rely on visual information for medical image interpretation. On the other hand, deep neural networks have demonstrated significant promise in similar tasks even where visual interpretation is challenging. Eye-gaze tracking has been used to capture the viewing behavior of domain experts, lending insights into the complexity of visual search. However, deep learning frameworks, even those that rely on attention mechanisms, do not leverage this rich domain information. RadioTransformer fills this critical gap by learning from radiologists' visual search patterns, encoded as 'human visual attention regions' in a cascaded global-focal transformer framework. The overall 'global' image characteristics and the more detailed 'local' features are captured by the proposed global and focal modules, respectively. We experimentally validate the efficacy of our student-teacher approach for 8 datasets involving different disease classification tasks where eye-gaze data is not available during the inference phase.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge