Michael C. Hughes
Dept. of Computer Science, Tufts University
SINCERE: Supervised Information Noise-Contrastive Estimation REvisited
Sep 25, 2023



Abstract:The information noise-contrastive estimation (InfoNCE) loss function provides the basis of many self-supervised deep learning methods due to its strong empirical results and theoretic motivation. Previous work suggests a supervised contrastive (SupCon) loss to extend InfoNCE to learn from available class labels. This SupCon loss has been widely-used due to reports of good empirical performance. However, in this work we suggest that the specific SupCon loss formulated by prior work has questionable theoretic justification, because it can encourage images from the same class to repel one another in the learned embedding space. This problematic behavior gets worse as the number of inputs sharing one class label increases. We propose the Supervised InfoNCE REvisited (SINCERE) loss as a remedy. SINCERE is a theoretically justified solution for a supervised extension of InfoNCE that never causes images from the same class to repel one another. We further show that minimizing our new loss is equivalent to maximizing a bound on the KL divergence between class conditional embedding distributions. We compare SINCERE and SupCon losses in terms of learning trajectories during pretraining and in ultimate linear classifier performance after finetuning. Our proposed SINCERE loss better separates embeddings from different classes during pretraining while delivering competitive accuracy.
Accuracy versus time frontiers of semi-supervised and self-supervised learning on medical images
Jul 18, 2023



Abstract:For many applications of classifiers to medical images, a trustworthy label for each image can be difficult or expensive to obtain. In contrast, images without labels are more readily available. Two major research directions both promise that additional unlabeled data can improve classifier performance: self-supervised learning pretrains useful representations on unlabeled data only, then fine-tunes a classifier on these representations via the labeled set; semi-supervised learning directly trains a classifier on labeled and unlabeled data simultaneously. Recent methods from both directions have claimed significant gains on non-medical tasks, but do not systematically assess medical images and mostly compare only to methods in the same direction. This study contributes a carefully-designed benchmark to help answer a practitioner's key question: given a small labeled dataset and a limited budget of hours to spend on training, what gains from additional unlabeled images are possible and which methods best achieve them? Unlike previous benchmarks, ours uses realistic-sized validation sets to select hyperparameters, assesses runtime-performance tradeoffs, and bridges two research fields. By comparing 6 semi-supervised methods and 5 self-supervised methods to strong labeled-only baselines on 3 medical datasets with 30-1000 labels per class, we offer insights to resource-constrained, results-focused practitioners: MixMatch, SimCLR, and BYOL represent strong choices that were not surpassed by more recent methods. After much effort selecting hyperparameters on one dataset, we publish settings that enable strong methods to perform well on new medical tasks within a few hours, with further search over dozens of hours delivering modest additional gains.
Detecting Heart Disease from Multi-View Ultrasound Images via Supervised Attention Multiple Instance Learning
May 25, 2023Abstract:Aortic stenosis (AS) is a degenerative valve condition that causes substantial morbidity and mortality. This condition is under-diagnosed and under-treated. In clinical practice, AS is diagnosed with expert review of transthoracic echocardiography, which produces dozens of ultrasound images of the heart. Only some of these views show the aortic valve. To automate screening for AS, deep networks must learn to mimic a human expert's ability to identify views of the aortic valve then aggregate across these relevant images to produce a study-level diagnosis. We find previous approaches to AS detection yield insufficient accuracy due to relying on inflexible averages across images. We further find that off-the-shelf attention-based multiple instance learning (MIL) performs poorly. We contribute a new end-to-end MIL approach with two key methodological innovations. First, a supervised attention technique guides the learned attention mechanism to favor relevant views. Second, a novel self-supervised pretraining strategy applies contrastive learning on the representation of the whole study instead of individual images as commonly done in prior literature. Experiments on an open-access dataset and an external validation set show that our approach yields higher accuracy while reducing model size.
Non-Parametric and Regularized Dynamical Wasserstein Barycenters for Time-Series Analysis
Oct 07, 2022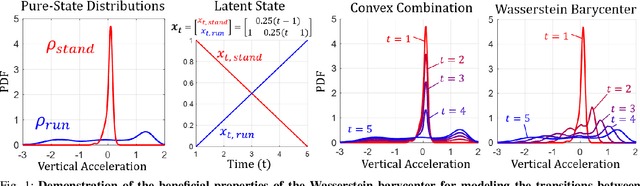
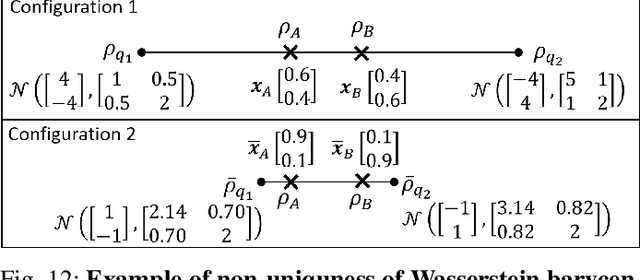
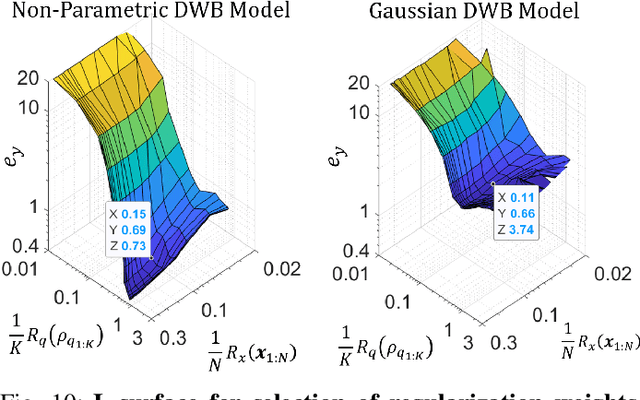
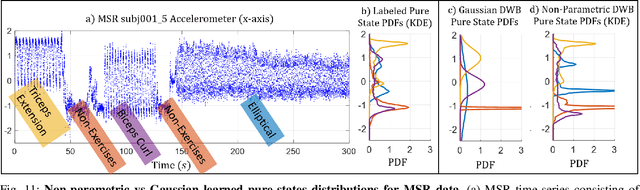

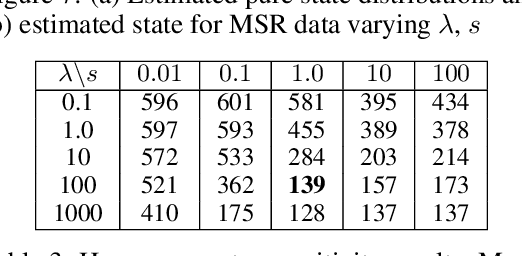
Abstract:We consider probabilistic time-series models for systems that gradually transition among a finite number of states. We are particularly motivated by applications such as human activity analysis where the observed time-series contains segments representing distinct activities such as running or walking as well as segments characterized by continuous transition among these states. Accordingly, the dynamical Wasserstein barycenter (DWB) model introduced in Cheng et al. in 2021 [1] associates with each state, which we call a pure state, its own probability distribution, and models these continuous transitions with the dynamics of the barycentric weights that combine the pure state distributions via the Wasserstein barycenter. Here, focusing on the univariate case where Wasserstein distances and barycenters can be computed in closed form, we extend [1] by discussing two challenges associated with learning a DWB model and two improvements. First, we highlight the issue of uniqueness in identifying the model parameters. Secondly, we discuss the challenge of estimating a dynamically evolving distribution given a limited number of samples. The uncertainty associated with this estimation may cause a model's learned dynamics to not reflect the gradual transitions characteristic of the system. The first improvement introduces a regularization framework that addresses this uncertainty by imposing temporal smoothness on the dynamics of the barycentric weights while leveraging the understanding of the non-uniqueness of the problem. This is done without defining an entire stochastic model for the dynamics of the system as in [1]. Our second improvement lifts the Gaussian assumption on the pure states distributions in [1] by proposing a quantile-based non-parametric representation. We pose model estimation in a variational framework and propose a finite approximation to the infinite dimensional problem.
Fix-A-Step: Effective Semi-supervised Learning from Uncurated Unlabeled Sets
Aug 25, 2022
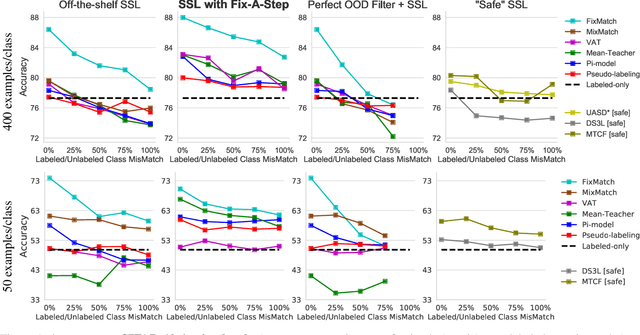
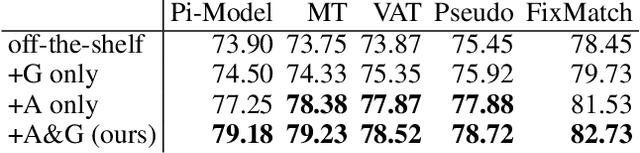
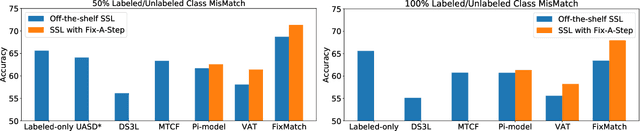
Abstract:Semi-supervised learning (SSL) promises gains in accuracy compared to training classifiers on small labeled datasets by also training on many unlabeled images. In realistic applications like medical imaging, unlabeled sets will be collected for expediency and thus uncurated: possibly different from the labeled set in represented classes or class frequencies. Unfortunately, modern deep SSL often makes accuracy worse when given uncurated unlabeled sets. Recent remedies suggest filtering approaches that detect out-of-distribution unlabeled examples and then discard or downweight them. Instead, we view all unlabeled examples as potentially helpful. We introduce a procedure called Fix-A-Step that can improve heldout accuracy of common deep SSL methods despite lack of curation. The key innovations are augmentations of the labeled set inspired by all unlabeled data and a modification of gradient descent updates to prevent following the multi-task SSL loss from hurting labeled-set accuracy. Though our method is simpler than alternatives, we show consistent accuracy gains on CIFAR-10 and CIFAR-100 benchmarks across all tested levels of artificial contamination for the unlabeled sets. We further suggest a real medical benchmark for SSL: recognizing the view type of ultrasound images of the heart. Our method can learn from 353,500 truly uncurated unlabeled images to deliver gains that generalize across hospitals.
NovelCraft: A Dataset for Novelty Detection and Discovery in Open Worlds
Jun 23, 2022
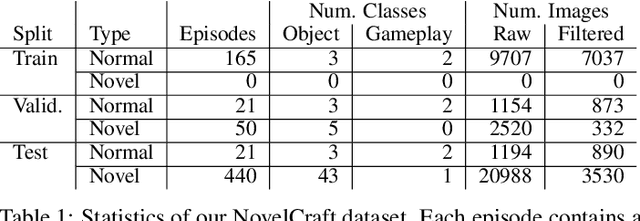
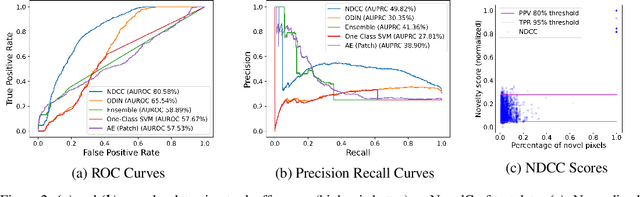
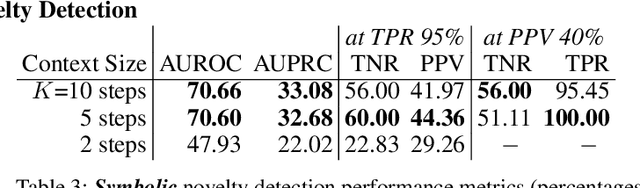
Abstract:In order for artificial agents to perform useful tasks in changing environments, they must be able to both detect and adapt to novelty. However, visual novelty detection research often only evaluates on repurposed datasets such as CIFAR-10 originally intended for object classification. This practice restricts novelties to well-framed images of distinct object types. We suggest that new benchmarks are needed to represent the challenges of navigating an open world. Our new NovelCraft dataset contains multi-modal episodic data of the images and symbolic world-states seen by an agent completing a pogo-stick assembly task within a video game world. In some episodes, we insert novel objects that can impact gameplay. Novelty can vary in size, position, and occlusion within complex scenes. We benchmark state-of-the-art novelty detection and generalized category discovery models with a focus on comprehensive evaluation. Results suggest an opportunity for future research: models aware of task-specific costs of different types of mistakes could more effectively detect and adapt to novelty in open worlds.
Easy Variational Inference for Categorical Models via an Independent Binary Approximation
May 31, 2022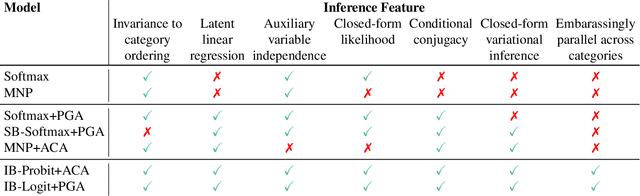
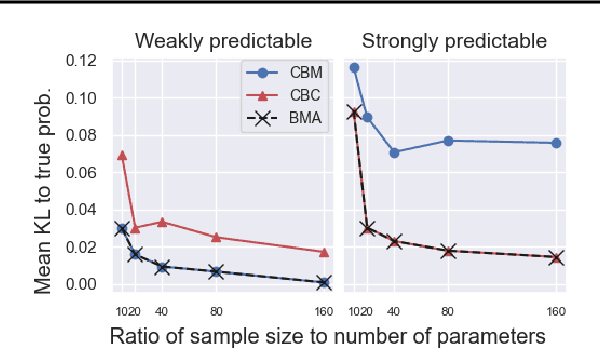
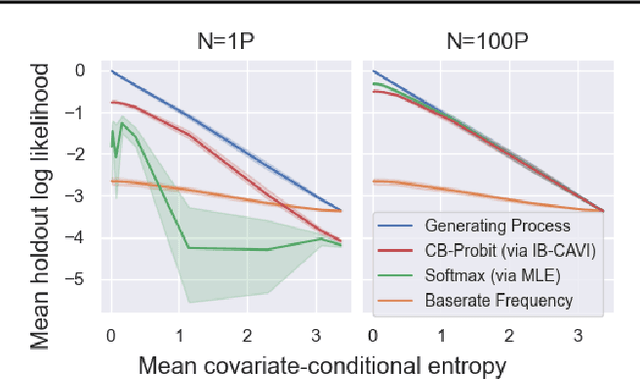
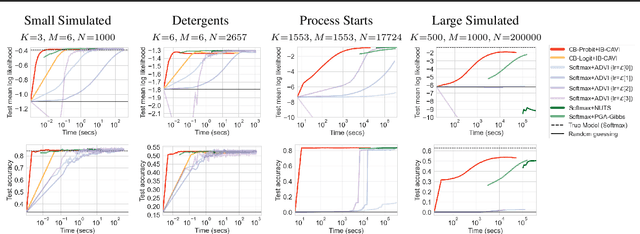
Abstract:We pursue tractable Bayesian analysis of generalized linear models (GLMs) for categorical data. Thus far, GLMs are difficult to scale to more than a few dozen categories due to non-conjugacy or strong posterior dependencies when using conjugate auxiliary variable methods. We define a new class of GLMs for categorical data called categorical-from-binary (CB) models. Each CB model has a likelihood that is bounded by the product of binary likelihoods, suggesting a natural posterior approximation. This approximation makes inference straightforward and fast; using well-known auxiliary variables for probit or logistic regression, the product of binary models admits conjugate closed-form variational inference that is embarrassingly parallel across categories and invariant to category ordering. Moreover, an independent binary model simultaneously approximates multiple CB models. Bayesian model averaging over these can improve the quality of the approximation for any given dataset. We show that our approach scales to thousands of categories, outperforming posterior estimation competitors like Automatic Differentiation Variational Inference (ADVI) and No U-Turn Sampling (NUTS) in the time required to achieve fixed prediction quality.
Dynamical Wasserstein Barycenters for Time-series Modeling
Oct 29, 2021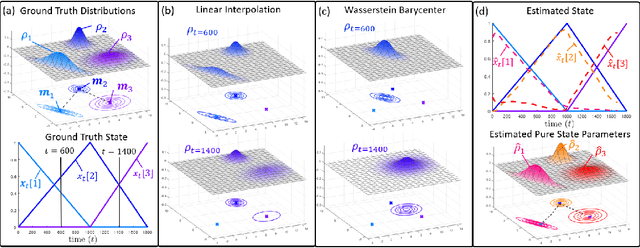
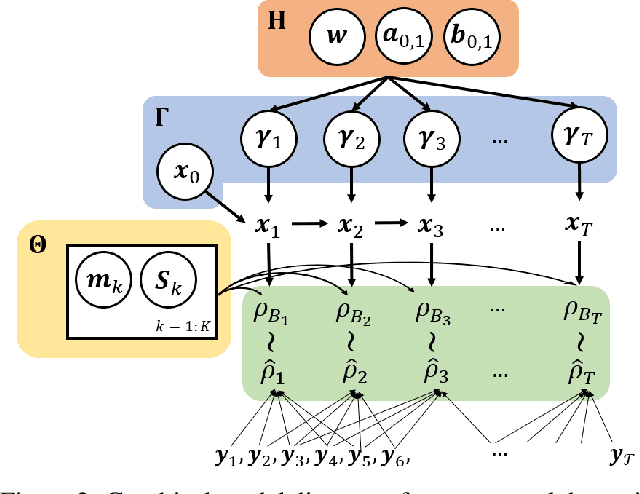
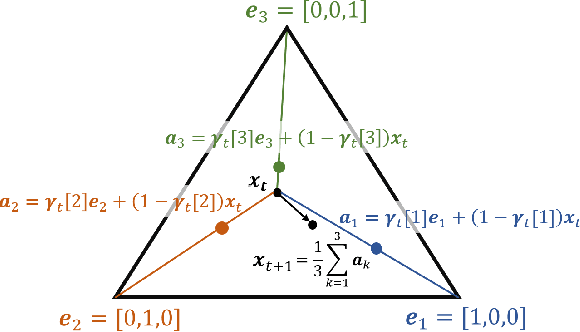
Abstract:Many time series can be modeled as a sequence of segments representing high-level discrete states, such as running and walking in a human activity application. Flexible models should describe the system state and observations in stationary "pure-state" periods as well as transition periods between adjacent segments, such as a gradual slowdown between running and walking. However, most prior work assumes instantaneous transitions between pure discrete states. We propose a dynamical Wasserstein barycentric (DWB) model that estimates the system state over time as well as the data-generating distributions of pure states in an unsupervised manner. Our model assumes each pure state generates data from a multivariate normal distribution, and characterizes transitions between states via displacement-interpolation specified by the Wasserstein barycenter. The system state is represented by a barycentric weight vector which evolves over time via a random walk on the simplex. Parameter learning leverages the natural Riemannian geometry of Gaussian distributions under the Wasserstein distance, which leads to improved convergence speeds. Experiments on several human activity datasets show that our proposed DWB model accurately learns the generating distribution of pure states while improving state estimation for transition periods compared to the commonly used linear interpolation mixture models.
A New Semi-supervised Learning Benchmark for Classifying View and Diagnosing Aortic Stenosis from Echocardiograms
Jul 30, 2021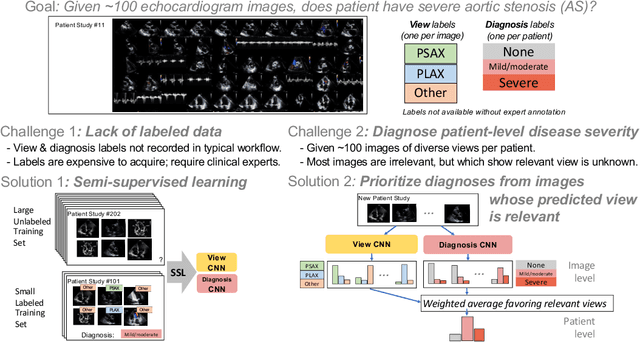
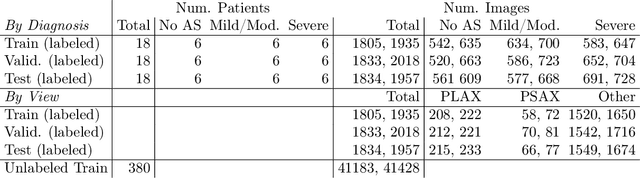
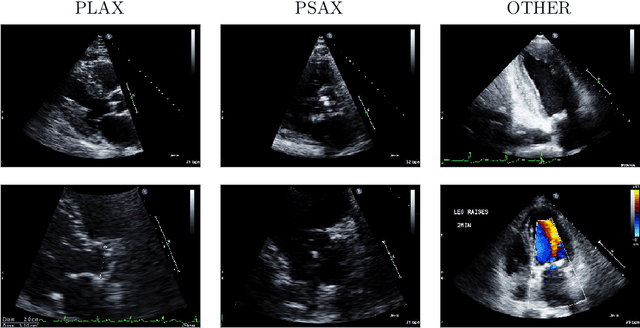
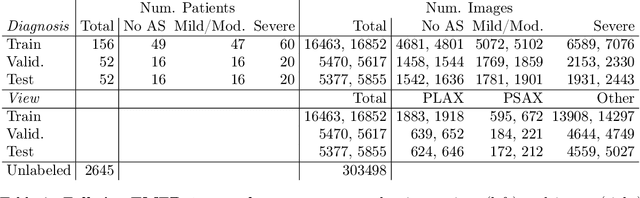
Abstract:Semi-supervised image classification has shown substantial progress in learning from limited labeled data, but recent advances remain largely untested for clinical applications. Motivated by the urgent need to improve timely diagnosis of life-threatening heart conditions, especially aortic stenosis, we develop a benchmark dataset to assess semi-supervised approaches to two tasks relevant to cardiac ultrasound (echocardiogram) interpretation: view classification and disease severity classification. We find that a state-of-the-art method called MixMatch achieves promising gains in heldout accuracy on both tasks, learning from a large volume of truly unlabeled images as well as a labeled set collected at great expense to achieve better performance than is possible with the labeled set alone. We further pursue patient-level diagnosis prediction, which requires aggregating across hundreds of images of diverse view types, most of which are irrelevant, to make a coherent prediction. The best patient-level performance is achieved by new methods that prioritize diagnosis predictions from images that are predicted to be clinically-relevant views and transfer knowledge from the view task to the diagnosis task. We hope our released Tufts Medical Echocardiogram Dataset and evaluation framework inspire further improvements in multi-task semi-supervised learning for clinical applications.
Evaluating the Use of Reconstruction Error for Novelty Localization
Jul 28, 2021
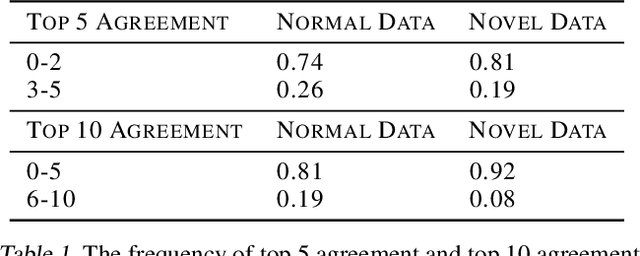
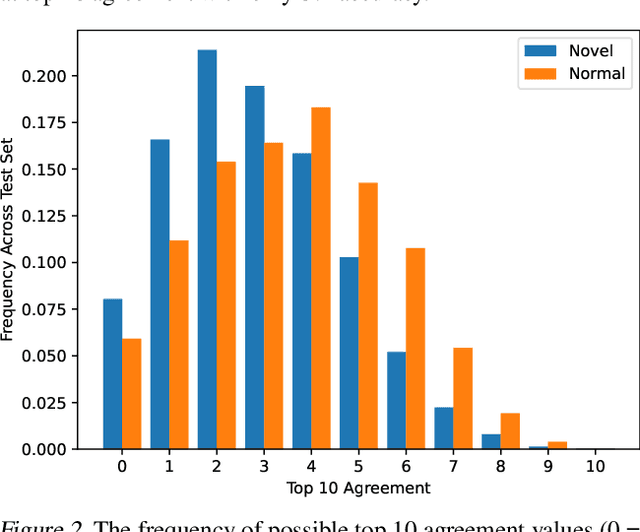
Abstract:The pixelwise reconstruction error of deep autoencoders is often utilized for image novelty detection and localization under the assumption that pixels with high error indicate which parts of the input image are unfamiliar and therefore likely to be novel. This assumed correlation between pixels with high reconstruction error and novel regions of input images has not been verified and may limit the accuracy of these methods. In this paper we utilize saliency maps to evaluate whether this correlation exists. Saliency maps reveal directly how much a change in each input pixel would affect reconstruction loss, while each pixel's reconstruction error may be attributed to many input pixels when layers are fully connected. We compare saliency maps to reconstruction error maps via qualitative visualizations as well as quantitative correspondence between the top K elements of the maps for both novel and normal images. Our results indicate that reconstruction error maps do not closely correlate with the importance of pixels in the input images, making them insufficient for novelty localization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge