Matthew Thompson
WavesFM: Hierarchical Representation Learning for Longitudinal Wearable Sensor Waveforms
May 09, 2026Abstract:Wearable sensors enable the continuous acquisition of high-resolution physiological waveforms, such as photoplethysmography and accelerometry, under free-living conditions. However, inferring health-related phenotypes from these signals presents significant challenges due to high sampling frequencies, multimodal dependencies, and extreme sequence lengths (e.g., weeks of recordings), compounded by a scarcity of ground-truth labels. To address these challenges, existing self-supervised learning (SSL) methodologies typically follow two paradigms: (1) learning rich morphological representations from short waveform segments while collapsing longitudinal dynamics through simple aggregation, or (2) modeling behavioral patterns from coarse, hand-crafted features (e.g. heart rate, step counts) spanning longer horizons but foregoing subtle, predictive signatures in raw waveforms. To bridge this gap, we propose WavesFM, a foundation model utilizing a two-stage SSL framework for longitudinal physiological data. Specifically, we decompose the learning problem into two stages: first, a segment-level encoder is pretrained to extract local embeddings from short waveforms; subsequently, a temporal encoder is trained to model the sequence of these embeddings across a multi-day horizon. This hierarchical approach overcomes the computational complexity of high-resolution, long-sequence data, allowing the overall model to capture both local signal semantics and the complex circadian and inter-day variations governing physiological dynamics. Pretrained on over 6.8M hours (N=324k individuals) of recordings for the first stage and 5.3M hours (N=10k) for the second stage, WavesFM demonstrates superior performance across 58 diverse tasks spanning demographics, lifestyle, health conditions, and medications.
SymptomAI: Towards a Conversational AI Agent for Everyday Symptom Assessment
May 05, 2026Abstract:Language models excel at diagnostic assessments on currated medical case-studies and vignettes, performing on par with, or better than, clinical professionals. However, existing studies focus on complex scenarios with rich context making it difficult to draw conclusions about how these systems perform for patients reporting symptoms in everyday life. We deployed SymptomAI, a set of conversational AI agents for end-to-end patient interviewing and differential diagnosis (DDx), via the Fitbit app in a study that randomized participants (N=13,917) to interact with five AI agents. This corpus captures diverse communication and a realistic distribution of illnesses from a real world population. A subset of 1,228 participants reported a clinician-provided diagnosis, and 517 of these were further evaluated by a panel of clinicians during over 250 hours of annotation. SymptomAI DDx were significantly more accurate (OR = 2.47, p < 0.001) than those from independent clinicians given the same dialogue in a blinded randomized comparison. Moreover, agentic strategies which conduct a dedicated symptom interview that elicit additional symptom information before providing a diagnosis, perform substantially better than baseline, user-guided conversations (p < 0.001). An auxiliary analysis on 1,509 conversations from a general US population panel validated that these results generalize beyond wearable device users. We used SymptomAI diagnoses as labels for all 13,917 participants to analyze over 500,000 days of wearable metrics across nearly 400 unique conditions. We identified strong associations between acute infections and physiological shifts (e.g., OR > 7 for influenza). While limited by self-reported ground truth, these results demonstrate the benefits of a dedicated and complete symptom interview compared to a user-guided symptom discussion, which is the default of most consumer LLMs.
Dialogue to Question Generation for Evidence-based Medical Guideline Agent Development
Mar 25, 2026Abstract:Evidence-based medicine (EBM) is central to high-quality care, but remains difficult to implement in fast-paced primary care settings. Physicians face short consultations, increasing patient loads, and lengthy guideline documents that are impractical to consult in real time. To address this gap, we investigate the feasibility of using large language models (LLMs) as ambient assistants that surface targeted, evidence-based questions during physician-patient encounters. Our study focuses on question generation rather than question answering, with the aim of scaffolding physician reasoning and integrating guideline-based practice into brief consultations. We implemented two prompting strategies, a zero-shot baseline and a multi-stage reasoning variant, using Gemini 2.5 as the backbone model. We evaluated on a benchmark of 80 de-identified transcripts from real clinical encounters, with six experienced physicians contributing over 90 hours of structured review. Results indicate that while general-purpose LLMs are not yet fully reliable, they can produce clinically meaningful and guideline-relevant questions, suggesting significant potential to reduce cognitive burden and make EBM more actionable at the point of care.
* 9 pages. To appear in Proceedings of Machine Learning Research (PMLR), Machine Learning for Health (ML4H) Symposium 2025
Managing the Stochastic: Foundations of Learning in Neuro-Symbolic Systems for Software Engineering
Dec 18, 2025Abstract:Current approaches to AI coding agents appear to blur the lines between the Large Language Model (LLM) and the agent itself, asking the LLM to make decisions best left to deterministic processes. This leads to systems prone to stochastic failures such as gaming unit tests or hallucinating syntax. Drawing on established software engineering practices that provide deterministic frameworks for managing unpredictable processes, this paper proposes setting the control boundary such that the LLM is treated as a component of the environment environment -- preserving its creative stochasticity -- rather than the decision-making agent. A \textbf{Dual-State Architecture} is formalized, separating workflow state (deterministic control flow) from environment state (stochastic generation). \textbf{Atomic Action Pairs} couple generation with verification as indivisible transactions, where \textbf{Guard Functions} act as sensing actions that project probabilistic outputs onto observable workflow state. The framework is validated on three code generation tasks across 13 LLMs (1.3B--15B parameters). For qualified instruction-following models, task success rates improved by up to 66 percentage points at 1.2--2.1$\times$ baseline computational cost. The results suggest that architectural constraints can substitute for parameter scale in achieving reliable code generation.
Combined fluorescence and photoacoustic imaging of tozuleristide in muscle tissue in vitro -- toward optically-guided solid tumor surgery: feasibility studies
Oct 31, 2025Abstract:Near-infrared fluorescence (NIRF) can deliver high-contrast, video-rate, non-contact imaging of tumor-targeted contrast agents with the potential to guide surgeries excising solid tumors. However, it has been met with skepticism for wide-margin excision due to sensitivity and resolution limitations at depths larger than ~5 mm in tissue. To address this limitation, fast-sweep photoacoustic-ultrasound (PAUS) imaging is proposed to complement NIRF. In an exploratory in vitro feasibility study using dark-red bovine muscle tissue, we observed that PAUS scanning can identify tozuleristide, a clinical stage investigational imaging agent, at a concentration of 20 uM from the background at depths of up to ~34 mm, highly extending the capabilities of NIRF alone. The capability of spectroscopic PAUS imaging was tested by direct injection of 20 uM tozuleristide into bovine muscle tissue at a depth of ~ 8 mm. It is shown that laser-fluence compensation and strong clutter suppression enabled by the unique capabilities of the fast-sweep approach greatly improve spectroscopic accuracy and the PA detection limit, and strongly reduce image artifacts. Thus, the combined NIRF-PAUS approach can be promising for comprehensive pre- (with PA) and intra- (with NIRF) operative solid tumor detection and wide-margin excision in optically guided solid tumor surgery.
kabr-tools: Automated Framework for Multi-Species Behavioral Monitoring
Oct 02, 2025Abstract:A comprehensive understanding of animal behavior ecology depends on scalable approaches to quantify and interpret complex, multidimensional behavioral patterns. Traditional field observations are often limited in scope, time-consuming, and labor-intensive, hindering the assessment of behavioral responses across landscapes. To address this, we present kabr-tools (Kenyan Animal Behavior Recognition Tools), an open-source package for automated multi-species behavioral monitoring. This framework integrates drone-based video with machine learning systems to extract behavioral, social, and spatial metrics from wildlife footage. Our pipeline leverages object detection, tracking, and behavioral classification systems to generate key metrics, including time budgets, behavioral transitions, social interactions, habitat associations, and group composition dynamics. Compared to ground-based methods, drone-based observations significantly improved behavioral granularity, reducing visibility loss by 15% and capturing more transitions with higher accuracy and continuity. We validate kabr-tools through three case studies, analyzing 969 behavioral sequences, surpassing the capacity of traditional methods for data capture and annotation. We found that, like Plains zebras, vigilance in Grevy's zebras decreases with herd size, but, unlike Plains zebras, habitat has a negligible impact. Plains and Grevy's zebras exhibit strong behavioral inertia, with rare transitions to alert behaviors and observed spatial segregation between Grevy's zebras, Plains zebras, and giraffes in mixed-species herds. By enabling automated behavioral monitoring at scale, kabr-tools offers a powerful tool for ecosystem-wide studies, advancing conservation, biodiversity research, and ecological monitoring.
Optimizing Image Capture for Computer Vision-Powered Taxonomic Identification and Trait Recognition of Biodiversity Specimens
May 22, 2025Abstract:Biological collections house millions of specimens documenting Earth's biodiversity, with digital images increasingly available through open-access platforms. Most imaging protocols were developed for human visual interpretation without considering computational analysis requirements. This paper aims to bridge the gap between current imaging practices and the potential for automated analysis by presenting key considerations for creating biological specimen images optimized for computer vision applications. We provide conceptual computer vision topics for context, addressing fundamental concerns including model generalization, data leakage, and comprehensive metadata documentation, and outline practical guidance on specimen imagine, and data storage. These recommendations were synthesized through interdisciplinary collaboration between taxonomists, collection managers, ecologists, and computer scientists. Through this synthesis, we have identified ten interconnected considerations that form a framework for successfully integrating biological specimen images into computer vision pipelines. The key elements include: (1) comprehensive metadata documentation, (2) standardized specimen positioning, (3) consistent size and color calibration, (4) protocols for handling multiple specimens in one image, (5) uniform background selection, (6) controlled lighting, (7) appropriate resolution and magnification, (8) optimal file formats, (9) robust data archiving strategies, and (10) accessible data sharing practices. By implementing these recommendations, collection managers, taxonomists, and biodiversity informaticians can generate images that support automated trait extraction, species identification, and novel ecological and evolutionary analyses at unprecedented scales. Successful implementation lies in thorough documentation of methodological choices.
Extracting COVID-19 Diagnoses and Symptoms From Clinical Text: A New Annotated Corpus and Neural Event Extraction Framework
Dec 02, 2020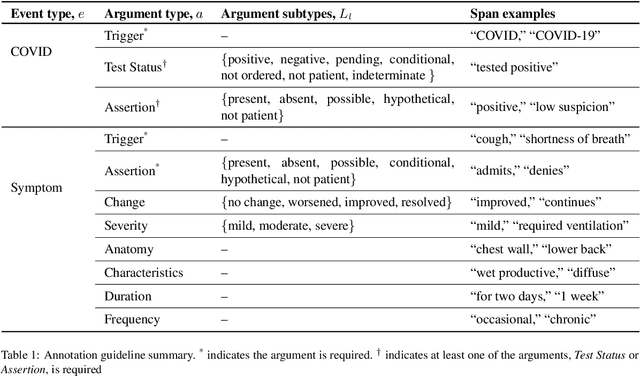
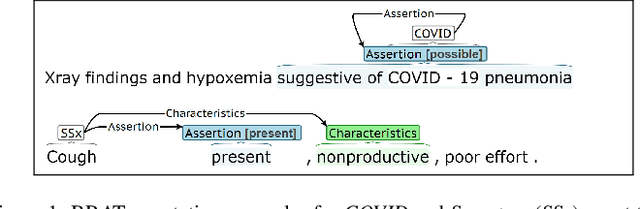
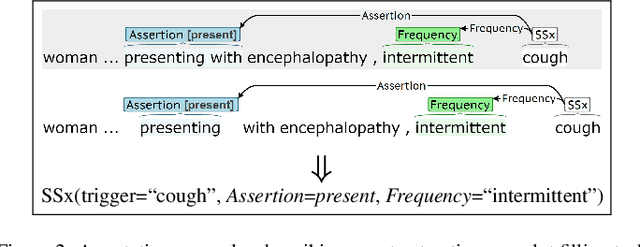
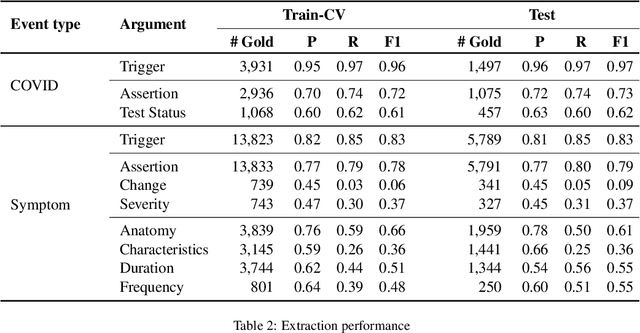
Abstract:Coronavirus disease 2019 (COVID-19) is a global pandemic. Although much has been learned about the novel coronavirus since its emergence, there are many open questions related to tracking its spread, describing symptomology, predicting the severity of infection, and forecasting healthcare utilization. Free-text clinical notes contain critical information for resolving these questions. Data-driven, automatic information extraction models are needed to use this text-encoded information in large-scale studies. This work presents a new clinical corpus, referred to as the COVID-19 Annotated Clinical Text (CACT) Corpus, which comprises 1,472 notes with detailed annotations characterizing COVID-19 diagnoses, testing, and clinical presentation. We introduce a span-based event extraction model that jointly extracts all annotated phenomena, achieving high performance in identifying COVID-19 and symptom events with associated assertion values (0.83-0.97 F1 for events and 0.73-0.79 F1 for assertions). In a secondary use application, we explored the prediction of COVID-19 test results using structured patient data (e.g. vital signs and laboratory results) and automatically extracted symptom information. The automatically extracted symptoms improve prediction performance, beyond structured data alone.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge