Marcel Roed
Synthetic Data for any Differentiable Target
Apr 09, 2026Abstract:What are the limits of controlling language models via synthetic training data? We develop a reinforcement learning (RL) primitive, the Dataset Policy Gradient (DPG), which can precisely optimize synthetic data generators to produce a dataset of targeted examples. When used for supervised fine-tuning (SFT) of a target model, these examples cause the target model to do well on a differentiable metric of our choice. Our approach achieves this by taking exact data attribution via higher-order gradients and using those scores as policy gradient rewards. We prove that this procedure closely approximates the true, intractable gradient for the synthetic data generator. To illustrate the potential of DPG, we show that, using only SFT on generated examples, we can cause the target model's LM head weights to (1) embed a QR code, (2) embed the pattern $\texttt{67}$, and (3) have lower $\ell^2$ norm. We additionally show that we can cause the generator to (4) rephrase inputs in a new language and (5) produce a specific UUID, even though neither of these objectives is conveyed in the generator's input prompts. These findings suggest that DPG is a powerful and flexible technique for shaping model properties using only synthetic training examples.
How to Build the Virtual Cell with Artificial Intelligence: Priorities and Opportunities
Sep 18, 2024

Abstract:The cell is arguably the smallest unit of life and is central to understanding biology. Accurate modeling of cells is important for this understanding as well as for determining the root causes of disease. Recent advances in artificial intelligence (AI), combined with the ability to generate large-scale experimental data, present novel opportunities to model cells. Here we propose a vision of AI-powered Virtual Cells, where robust representations of cells and cellular systems under different conditions are directly learned from growing biological data across measurements and scales. We discuss desired capabilities of AI Virtual Cells, including generating universal representations of biological entities across scales, and facilitating interpretable in silico experiments to predict and understand their behavior using Virtual Instruments. We further address the challenges, opportunities and requirements to realize this vision including data needs, evaluation strategies, and community standards and engagement to ensure biological accuracy and broad utility. We envision a future where AI Virtual Cells help identify new drug targets, predict cellular responses to perturbations, as well as scale hypothesis exploration. With open science collaborations across the biomedical ecosystem that includes academia, philanthropy, and the biopharma and AI industries, a comprehensive predictive understanding of cell mechanisms and interactions is within reach.
Real-time semantic segmentation on FPGAs for autonomous vehicles with hls4ml
May 16, 2022
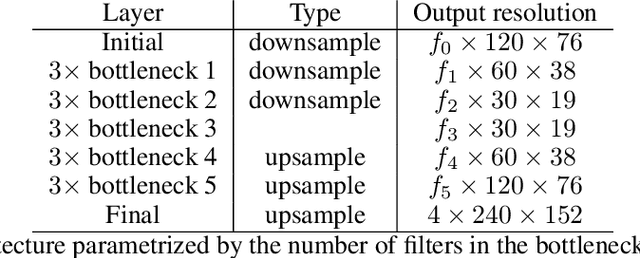

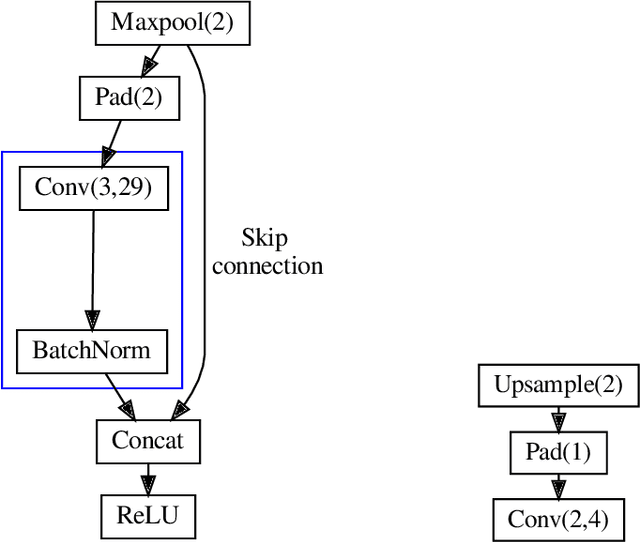
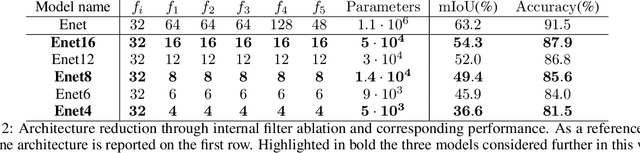
Abstract:In this paper, we investigate how field programmable gate arrays can serve as hardware accelerators for real-time semantic segmentation tasks relevant for autonomous driving. Considering compressed versions of the ENet convolutional neural network architecture, we demonstrate a fully-on-chip deployment with a latency of 4.9 ms per image, using less than 30% of the available resources on a Xilinx ZCU102 evaluation board. The latency is reduced to 3 ms per image when increasing the batch size to ten, corresponding to the use case where the autonomous vehicle receives inputs from multiple cameras simultaneously. We show, through aggressive filter reduction and heterogeneous quantization-aware training, and an optimized implementation of convolutional layers, that the power consumption and resource utilization can be significantly reduced while maintaining accuracy on the Cityscapes dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge