Laura A. Hallock
Characterizing Healthy & Post-Stroke Neuromotor Behavior During 6D Upper-Limb Isometric Gaming: Implications for Design of End-Effector Rehabilitation Robot Interfaces
Mar 10, 2026Abstract:Successful robot-mediated rehabilitation requires designing games and robot interventions that promote healthy motor practice. However, the interplay between a given user's neuromotor behavior, the gaming interface, and the physical robot makes designing system elements -- and even characterizing what behaviors are "healthy" or pathological -- challenging. We leverage our OpenRobotRehab 1.0 open access data set to assess the characteristics of 13 healthy and 2 post-stroke users' force output, muscle activations, and game performance while executing isometric trajectory tracking tasks using an end-effector rehabilitation robot. We present an assessment of how subtle aspects of interface design impact user behavior; an analysis of how pathological neuromotor behaviors are reflected in end-effector force dynamics; and a novel hidden Markov model (HMM)-based neuromotor behavior classification method based on surface electromyography (sEMG) signals during cyclic motions. We demonstrate that task specification (including which axes are constrained and how users interpret tracking instructions) shapes user behavior; that pathology-related features are detectable in 6D end-effector force data during isometric task execution (with significant differences between healthy and post-stroke profiles in force error and average force production at $p=0.05$); and that healthy neuromotor strategies are heterogeneous and inherently difficult to characterize. We also show that our HMM-based models discriminate healthy and post-stroke neuromotor dynamics where synergy-based decompositions reflect no such differentiation. Lastly, we discuss these results' implications for the design of adaptive end-effector rehabilitation robots capable of promoting healthier movement strategies across diverse user populations.
A Computer Vision Pipeline for Automated Determination of Cardiac Structure and Function and Detection of Disease by Two-Dimensional Echocardiography
Jan 12, 2018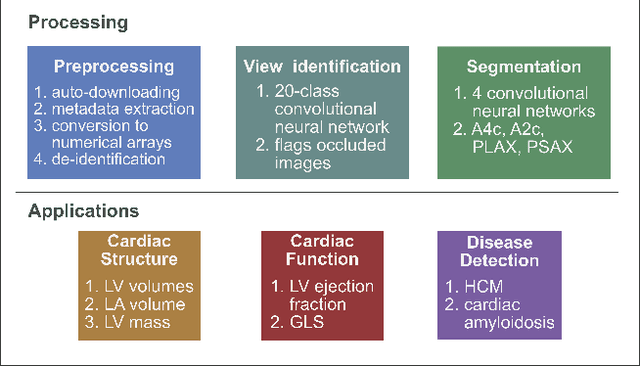
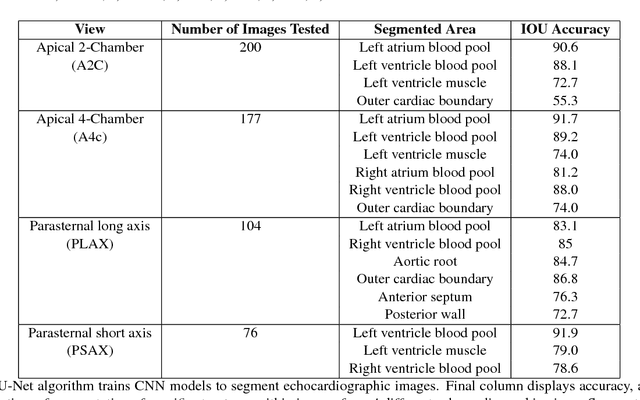
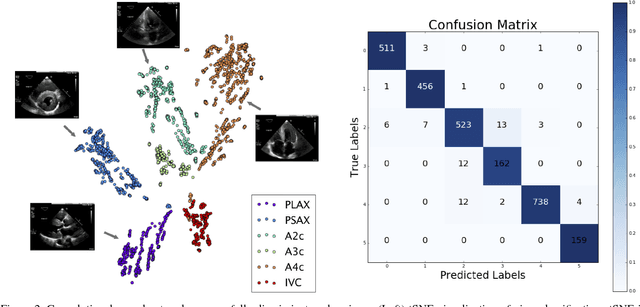
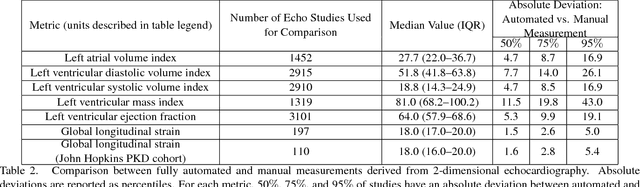
Abstract:Automated cardiac image interpretation has the potential to transform clinical practice in multiple ways including enabling low-cost serial assessment of cardiac function in the primary care and rural setting. We hypothesized that advances in computer vision could enable building a fully automated, scalable analysis pipeline for echocardiogram (echo) interpretation. Our approach entailed: 1) preprocessing; 2) convolutional neural networks (CNN) for view identification, image segmentation, and phasing of the cardiac cycle; 3) quantification of chamber volumes and left ventricular mass; 4) particle tracking to compute longitudinal strain; and 5) targeted disease detection. CNNs accurately identified views (e.g. 99% for apical 4-chamber) and segmented individual cardiac chambers. Cardiac structure measurements agreed with study report values (e.g. mean absolute deviations (MAD) of 7.7 mL/kg/m2 for left ventricular diastolic volume index, 2918 studies). We computed automated ejection fraction and longitudinal strain measurements (within 2 cohorts), which agreed with commercial software-derived values [for ejection fraction, MAD=5.3%, N=3101 studies; for strain, MAD=1.5% (n=197) and 1.6% (n=110)], and demonstrated applicability to serial monitoring of breast cancer patients for trastuzumab cardiotoxicity. Overall, we found that, compared to manual measurements, automated measurements had superior performance across seven internal consistency metrics with an average increase in the Spearman correlation coefficient of 0.05 (p=0.02). Finally, we developed disease detection algorithms for hypertrophic cardiomyopathy and cardiac amyloidosis, with C-statistics of 0.93 and 0.84, respectively. Our pipeline lays the groundwork for using automated interpretation to support point-of-care handheld cardiac ultrasound and large-scale analysis of the millions of echos archived within healthcare systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge