Joseph Jacobson
Accelerating Protein Molecular Dynamics Simulation with DeepJump
Sep 16, 2025



Abstract:Unraveling the dynamical motions of biomolecules is essential for bridging their structure and function, yet it remains a major computational challenge. Molecular dynamics (MD) simulation provides a detailed depiction of biomolecular motion, but its high-resolution temporal evolution comes at significant computational cost, limiting its applicability to timescales of biological relevance. Deep learning approaches have emerged as promising solutions to overcome these computational limitations by learning to predict long-timescale dynamics. However, generalizable kinetics models for proteins remain largely unexplored, and the fundamental limits of achievable acceleration while preserving dynamical accuracy are poorly understood. In this work, we fill this gap with DeepJump, an Euclidean-Equivariant Flow Matching-based model for predicting protein conformational dynamics across multiple temporal scales. We train DeepJump on trajectories of the diverse proteins of mdCATH, systematically studying our model's performance in generalizing to long-term dynamics of fast-folding proteins and characterizing the trade-off between computational acceleration and prediction accuracy. We demonstrate the application of DeepJump to ab initio folding, showcasing prediction of folding pathways and native states. Our results demonstrate that DeepJump achieves significant $\approx$1000$\times$ computational acceleration while effectively recovering long-timescale dynamics, providing a stepping stone for enabling routine simulation of proteins.
RiboGen: RNA Sequence and Structure Co-Generation with Equivariant MultiFlow
Mar 03, 2025



Abstract:Ribonucleic acid (RNA) plays fundamental roles in biological systems, from carrying genetic information to performing enzymatic function. Understanding and designing RNA can enable novel therapeutic application and biotechnological innovation. To enhance RNA design, in this paper we introduce RiboGen, the first deep learning model to simultaneously generate RNA sequence and all-atom 3D structure. RiboGen leverages the standard Flow Matching with Discrete Flow Matching in a multimodal data representation. RiboGen is based on Euclidean Equivariant neural networks for efficiently processing and learning three-dimensional geometry. Our experiments show that RiboGen can efficiently generate chemically plausible and self-consistent RNA samples. Our results suggest that co-generation of sequence and structure is a competitive approach for modeling RNA.
EquiJump: Protein Dynamics Simulation via SO(3)-Equivariant Stochastic Interpolants
Oct 12, 2024



Abstract:Mapping the conformational dynamics of proteins is crucial for elucidating their functional mechanisms. While Molecular Dynamics (MD) simulation enables detailed time evolution of protein motion, its computational toll hinders its use in practice. To address this challenge, multiple deep learning models for reproducing and accelerating MD have been proposed drawing on transport-based generative methods. However, existing work focuses on generation through transport of samples from prior distributions, that can often be distant from the data manifold. The recently proposed framework of stochastic interpolants, instead, enables transport between arbitrary distribution endpoints. Building upon this work, we introduce EquiJump, a transferable SO(3)-equivariant model that bridges all-atom protein dynamics simulation time steps directly. Our approach unifies diverse sampling methods and is benchmarked against existing models on trajectory data of fast folding proteins. EquiJump achieves state-of-the-art results on dynamics simulation with a transferable model on all of the fast folding proteins.
E3STO: Orbital Inspired SE(3)-Equivariant Molecular Representation for Electron Density Prediction
Oct 08, 2024



Abstract:Electron density prediction stands as a cornerstone challenge in molecular systems, pivotal for various applications such as understanding molecular interactions and conducting precise quantum mechanical calculations. However, the scaling of density functional theory (DFT) calculations is prohibitively expensive. Machine learning methods provide an alternative, offering efficiency and accuracy. We introduce a novel SE(3)-equivariant architecture, drawing inspiration from Slater-Type Orbitals (STO), to learn representations of molecular electronic structures. Our approach offers an alternative functional form for learned orbital-like molecular representation. We showcase the effectiveness of our method by achieving SOTA prediction accuracy of molecular electron density with 30-70\% improvement over other work on Molecular Dynamics data.
Ophiuchus: Scalable Modeling of Protein Structures through Hierarchical Coarse-graining SO(3)-Equivariant Autoencoders
Oct 04, 2023Abstract:Three-dimensional native states of natural proteins display recurring and hierarchical patterns. Yet, traditional graph-based modeling of protein structures is often limited to operate within a single fine-grained resolution, and lacks hourglass neural architectures to learn those high-level building blocks. We narrow this gap by introducing Ophiuchus, an SO(3)-equivariant coarse-graining model that efficiently operates on all heavy atoms of standard protein residues, while respecting their relevant symmetries. Our model departs from current approaches that employ graph modeling, instead focusing on local convolutional coarsening to model sequence-motif interactions in log-linear length complexity. We train Ophiuchus on contiguous fragments of PDB monomers, investigating its reconstruction capabilities across different compression rates. We examine the learned latent space and demonstrate its prompt usage in conformational interpolation, comparing interpolated trajectories to structure snapshots from the PDBFlex dataset. Finally, we leverage denoising diffusion probabilistic models (DDPM) to efficiently sample readily-decodable latent embeddings of diverse miniproteins. Our experiments demonstrate Ophiuchus to be a scalable basis for efficient protein modeling and generation.
Interpretable Neuroevolutionary Models for Learning Non-Differentiable Functions and Programs
Jul 16, 2020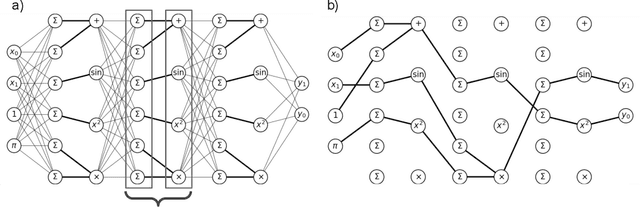
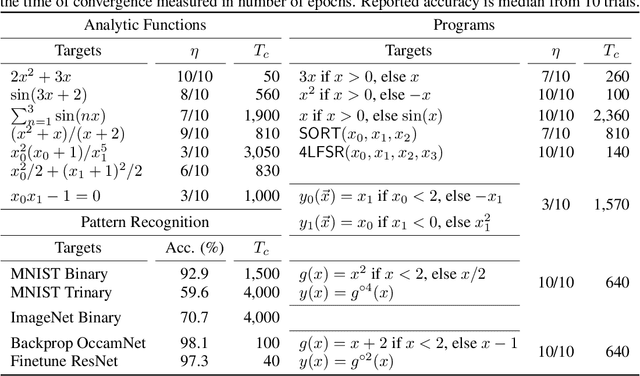
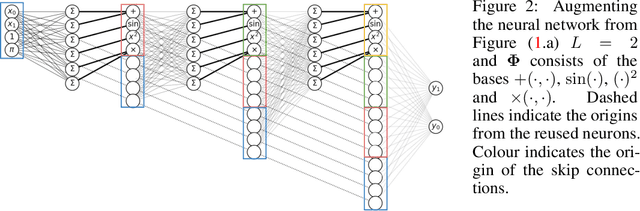
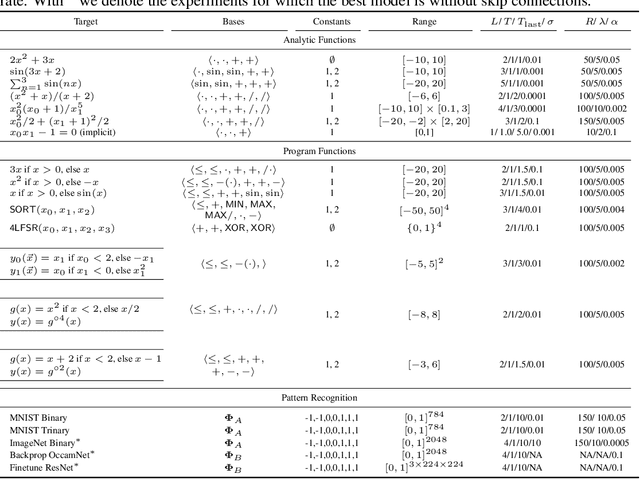
Abstract:A key factor in the modern success of deep learning is the astonishing expressive power of neural networks. However, this comes at the cost of complex, black-boxed models that are unable to extrapolate beyond the domain of the training dataset, conflicting with goals of expressing physical laws or building human-readable programs. In this paper, we introduce OccamNet, a neural network model that can find interpretable, compact and sparse solutions for fitting data, \`{a} la Occam's razor. Our model defines a probability distribution over a non-differentiable function space, and we introduce an optimization method that samples functions and updates the weights based on cross-entropy matching in an evolutionary strategy: we train by biasing the probability mass towards better fitting solutions. We demonstrate that we can fit a variety of algorithms, ranging from simple analytic functions through recursive programs to even simple image classification. Our method takes minimal memory footprint, does not require AI accelerators for efficient training, fits complicated functions in minutes of training on a single CPU, and demonstrates significant performance gains when scaled on GPU. Our implementation, demonstrations and instructions for reproducing the experiments are available at https://github.com/AllanSCosta/occam-net.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge