John Guttag
ScribblePrompt: Fast and Flexible Interactive Segmentation for Any Medical Image
Dec 12, 2023Abstract:Semantic medical image segmentation is a crucial part of both scientific research and clinical care. With enough labelled data, deep learning models can be trained to accurately automate specific medical image segmentation tasks. However, manually segmenting images to create training data is highly labor intensive. In this paper, we present ScribblePrompt, an interactive segmentation framework for medical imaging that enables human annotators to segment unseen structures using scribbles, clicks, and bounding boxes. Scribbles are an intuitive and effective form of user interaction for complex tasks, however most existing methods focus on click-based interactions. We introduce algorithms for simulating realistic scribbles that enable training models that are amenable to multiple types of interaction. To achieve generalization to new tasks, we train on a diverse collection of 65 open-access biomedical datasets -- using both real and synthetic labels. We test ScribblePrompt on multiple network architectures and unseen datasets, and demonstrate that it can be used in real-time on a single CPU. We evaluate ScribblePrompt using manually-collected scribbles, simulated interactions, and a user study. ScribblePrompt outperforms existing methods in all our evaluations. In the user study, ScribblePrompt reduced annotation time by 28% while improving Dice by 15% compared to existing methods. We showcase ScribblePrompt in an online demo and provide code at https://scribbleprompt.csail.mit.edu
GIST: Generating Image-Specific Text for Fine-grained Object Classification
Aug 04, 2023



Abstract:Recent vision-language models outperform vision-only models on many image classification tasks. However, because of the absence of paired text/image descriptions, it remains difficult to fine-tune these models for fine-grained image classification. In this work, we propose a method, GIST, for generating image-specific fine-grained text descriptions from image-only datasets, and show that these text descriptions can be used to improve classification. Key parts of our method include 1. prompting a pretrained large language model with domain-specific prompts to generate diverse fine-grained text descriptions for each class and 2. using a pretrained vision-language model to match each image to label-preserving text descriptions that capture relevant visual features in the image. We demonstrate the utility of GIST by fine-tuning vision-language models on the image-and-generated-text pairs to learn an aligned vision-language representation space for improved classification. We evaluate our learned representation space in full-shot and few-shot scenarios across four diverse fine-grained classification datasets, each from a different domain. Our method achieves an average improvement of $4.1\%$ in accuracy over CLIP linear probes and an average of $1.1\%$ improvement in accuracy over the previous state-of-the-art image-text classification method on the full-shot datasets. Our method achieves similar improvements across few-shot regimes. Code is available at https://github.com/emu1729/GIST.
Sequential Multi-Dimensional Self-Supervised Learning for Clinical Time Series
Jul 20, 2023



Abstract:Self-supervised learning (SSL) for clinical time series data has received significant attention in recent literature, since these data are highly rich and provide important information about a patient's physiological state. However, most existing SSL methods for clinical time series are limited in that they are designed for unimodal time series, such as a sequence of structured features (e.g., lab values and vitals signs) or an individual high-dimensional physiological signal (e.g., an electrocardiogram). These existing methods cannot be readily extended to model time series that exhibit multimodality, with structured features and high-dimensional data being recorded at each timestep in the sequence. In this work, we address this gap and propose a new SSL method -- Sequential Multi-Dimensional SSL -- where a SSL loss is applied both at the level of the entire sequence and at the level of the individual high-dimensional data points in the sequence in order to better capture information at both scales. Our strategy is agnostic to the specific form of loss function used at each level -- it can be contrastive, as in SimCLR, or non-contrastive, as in VICReg. We evaluate our method on two real-world clinical datasets, where the time series contains sequences of (1) high-frequency electrocardiograms and (2) structured data from lab values and vitals signs. Our experimental results indicate that pre-training with our method and then fine-tuning on downstream tasks improves performance over baselines on both datasets, and in several settings, can lead to improvements across different self-supervised loss functions.
Multi-Similarity Contrastive Learning
Jul 06, 2023Abstract:Given a similarity metric, contrastive methods learn a representation in which examples that are similar are pushed together and examples that are dissimilar are pulled apart. Contrastive learning techniques have been utilized extensively to learn representations for tasks ranging from image classification to caption generation. However, existing contrastive learning approaches can fail to generalize because they do not take into account the possibility of different similarity relations. In this paper, we propose a novel multi-similarity contrastive loss (MSCon), that learns generalizable embeddings by jointly utilizing supervision from multiple metrics of similarity. Our method automatically learns contrastive similarity weightings based on the uncertainty in the corresponding similarity, down-weighting uncertain tasks and leading to better out-of-domain generalization to new tasks. We show empirically that networks trained with MSCon outperform state-of-the-art baselines on in-domain and out-of-domain settings.
Coarse race data conceals disparities in clinical risk score performance
Apr 18, 2023



Abstract:Healthcare data in the United States often records only a patient's coarse race group: for example, both Indian and Chinese patients are typically coded as ``Asian.'' It is unknown, however, whether this coarse coding conceals meaningful disparities in the performance of clinical risk scores across granular race groups. Here we show that it does. Using data from 418K emergency department visits, we assess clinical risk score performance disparities across granular race groups for three outcomes, five risk scores, and four performance metrics. Across outcomes and metrics, we show that there are significant granular disparities in performance within coarse race categories. In fact, variation in performance metrics within coarse groups often exceeds the variation between coarse groups. We explore why these disparities arise, finding that outcome rates, feature distributions, and the relationships between features and outcomes all vary significantly across granular race categories. Our results suggest that healthcare providers, hospital systems, and machine learning researchers should strive to collect, release, and use granular race data in place of coarse race data, and that existing analyses may significantly underestimate racial disparities in performance.
Non-Proportional Parametrizations for Stable Hypernetwork Learning
Apr 15, 2023



Abstract:Hypernetworks are neural networks that generate the parameters of another neural network. In many scenarios, current hypernetwork training strategies are unstable, and convergence is often far slower than for non-hypernetwork models. We show that this problem is linked to an issue that arises when using common choices of hypernetwork architecture and initialization. We demonstrate analytically and experimentally how this numerical issue can lead to an instability during training that slows, and sometimes even prevents, convergence. We also demonstrate that popular deep learning normalization strategies fail to address these issues. We then propose a solution to the problem based on a revised hypernetwork formulation that uses non-proportional additive parametrizations. We test the proposed reparametrization on several tasks, and demonstrate that it consistently leads to more stable training, achieving faster convergence.
UniverSeg: Universal Medical Image Segmentation
Apr 12, 2023



Abstract:While deep learning models have become the predominant method for medical image segmentation, they are typically not capable of generalizing to unseen segmentation tasks involving new anatomies, image modalities, or labels. Given a new segmentation task, researchers generally have to train or fine-tune models, which is time-consuming and poses a substantial barrier for clinical researchers, who often lack the resources and expertise to train neural networks. We present UniverSeg, a method for solving unseen medical segmentation tasks without additional training. Given a query image and example set of image-label pairs that define a new segmentation task, UniverSeg employs a new Cross-Block mechanism to produce accurate segmentation maps without the need for additional training. To achieve generalization to new tasks, we have gathered and standardized a collection of 53 open-access medical segmentation datasets with over 22,000 scans, which we refer to as MegaMedical. We used this collection to train UniverSeg on a diverse set of anatomies and imaging modalities. We demonstrate that UniverSeg substantially outperforms several related methods on unseen tasks, and thoroughly analyze and draw insights about important aspects of the proposed system. The UniverSeg source code and model weights are freely available at https://universeg.csail.mit.edu
Amortized Learning of Dynamic Feature Scaling for Image Segmentation
Apr 11, 2023



Abstract:Convolutional neural networks (CNN) have become the predominant model for image segmentation tasks. Most CNN segmentation architectures resize spatial dimensions by a fixed factor of two to aggregate spatial context. Recent work has explored using other resizing factors to improve model accuracy for specific applications. However, finding the appropriate rescaling factor most often involves training a separate network for many different factors and comparing the performance of each model. The computational burden of these models means that in practice it is rarely done, and when done only a few different scaling factors are considered. In this work, we present a hypernetwork strategy that can be used to easily and rapidly generate the Pareto frontier for the trade-off between accuracy and efficiency as the rescaling factor varies. We show how to train a single hypernetwork that generates CNN parameters conditioned on a rescaling factor. This enables a user to quickly choose a rescaling factor that appropriately balances accuracy and computational efficiency for their particular needs. We focus on image segmentation tasks, and demonstrate the value of this approach across various domains. We also find that, for a given rescaling factor, our single hypernetwork outperforms CNNs trained with fixed rescaling factors.
SizeGAN: Improving Size Representation in Clothing Catalogs
Nov 05, 2022



Abstract:Online clothing catalogs lack diversity in body shape and garment size. Brands commonly display their garments on models of one or two sizes, rarely including plus-size models. In this work, we propose a new method, SizeGAN, for generating images of garments on different-sized models. To change the garment and model size while maintaining a photorealistic image, we incorporate image alignment ideas from the medical imaging literature into the StyleGAN2-ADA architecture. Our method learns deformation fields at multiple resolutions and uses a spatial transformer to modify the garment and model size. We evaluate our approach along three dimensions: realism, garment faithfulness, and size. To our knowledge, SizeGAN is the first method to focus on this size under-representation problem for modeling clothing. We provide an analysis comparing SizeGAN to other plausible approaches and additionally provide the first clothing dataset with size labels. In a user study comparing SizeGAN and two recent virtual try-on methods, we show that our method ranks first in each dimension, and was vastly preferred for realism and garment faithfulness. In comparison to most previous work, which has focused on generating photorealistic images of garments, our work shows that it is possible to generate images that are both photorealistic and cover diverse garment sizes.
Improved Text Classification via Test-Time Augmentation
Jun 27, 2022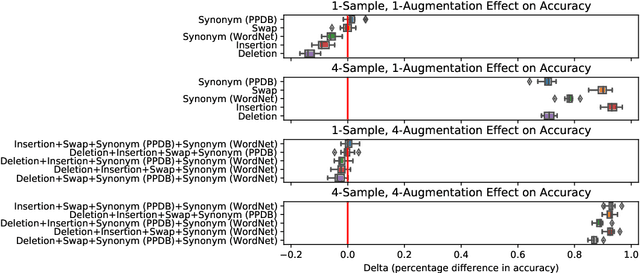

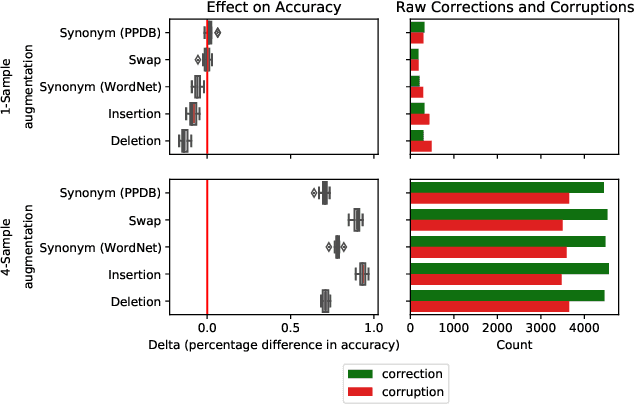
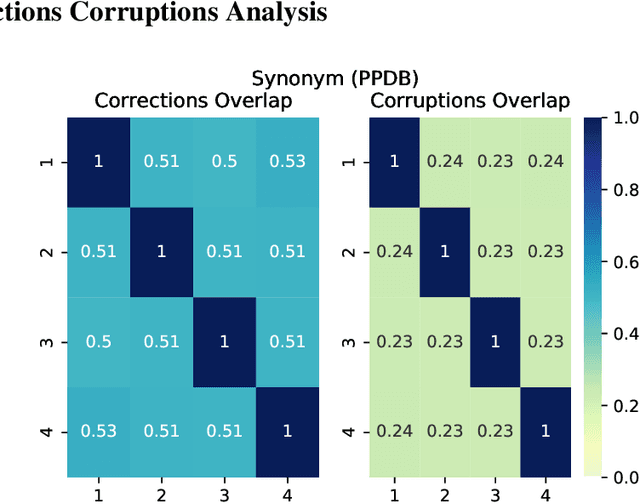
Abstract:Test-time augmentation -- the aggregation of predictions across transformed examples of test inputs -- is an established technique to improve the performance of image classification models. Importantly, TTA can be used to improve model performance post-hoc, without additional training. Although test-time augmentation (TTA) can be applied to any data modality, it has seen limited adoption in NLP due in part to the difficulty of identifying label-preserving transformations. In this paper, we present augmentation policies that yield significant accuracy improvements with language models. A key finding is that augmentation policy design -- for instance, the number of samples generated from a single, non-deterministic augmentation -- has a considerable impact on the benefit of TTA. Experiments across a binary classification task and dataset show that test-time augmentation can deliver consistent improvements over current state-of-the-art approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge